T5: Exploring Transfer Learning Limits
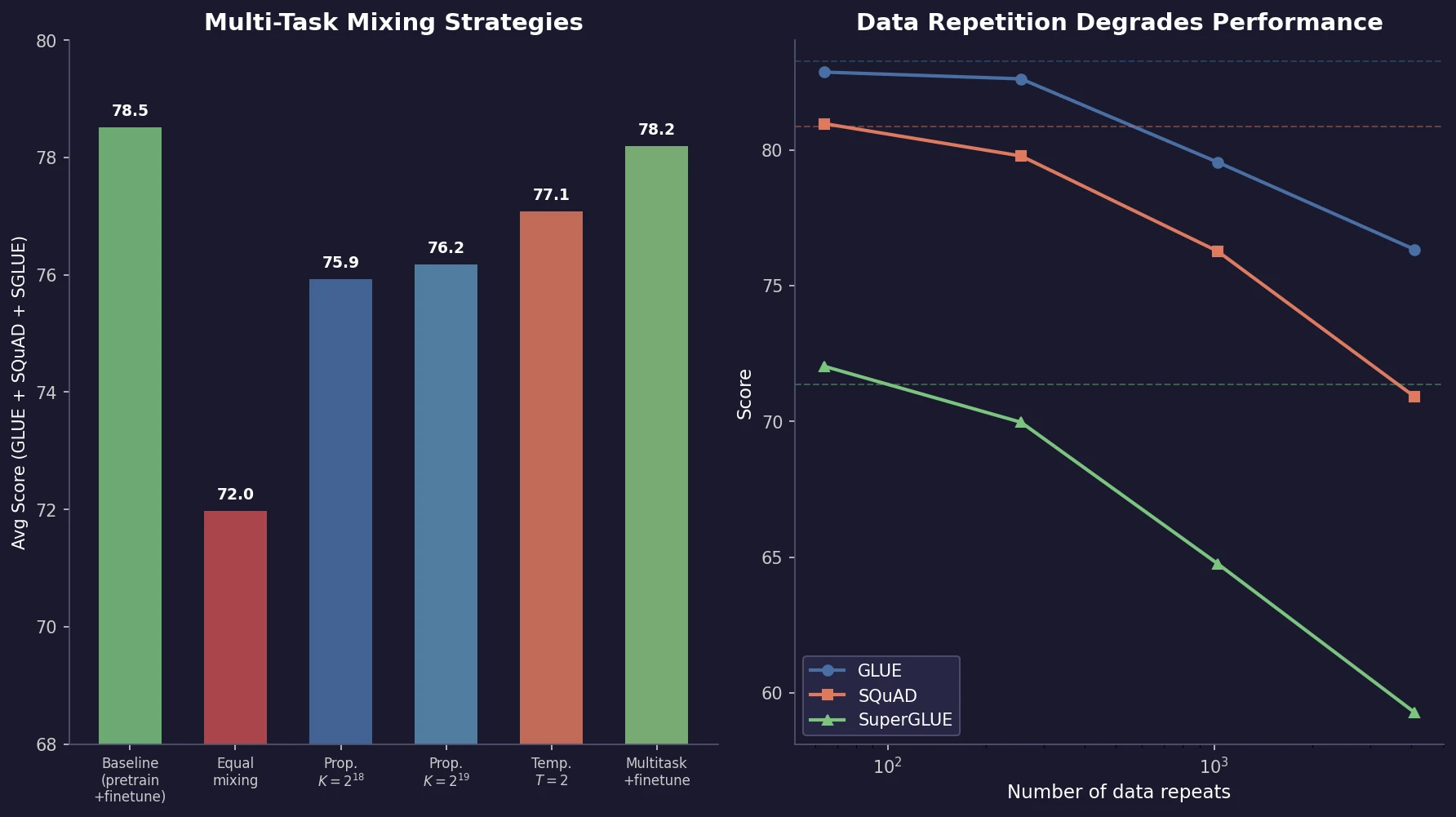
Raffel et al. introduce T5, a unified text-to-text framework for NLP transfer learning. Through systematic ablation of architectures, pre-training objectives, datasets, and multi-task mixing strategies, they identify best practices and scale to 11B parameters, achieving state-of-the-art results across multiple benchmarks.