CLEF-IP 2012: Patent and Chemical Structure Benchmark
A resource paper detailing the CLEF-IP 2012 benchmarking lab. It introduces specific IR tasks for patent processing along with ground-truth datasets.

A resource paper detailing the CLEF-IP 2012 benchmarking lab. It introduces specific IR tasks for patent processing along with ground-truth datasets.
This resource paper details the third TREC Chemical IR campaign, introducing a novel Image-to-Structure task and analyzing 36 runs from 9 groups to benchmark chemical information retrieval.
Imago is an open-source, cross-platform C++ toolkit designed to recognize 2D chemical structure images from scientific papers and convert them into machine-readable molecule formats using a rule-based pipeline.
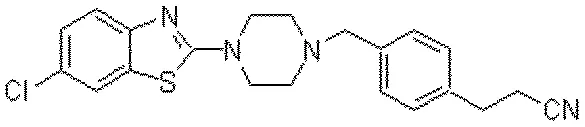
This paper presents OSRA, the first open-source utility for converting graphical chemical structures from documents into machine-readable formats (SMILES/SD). It outlines a pipeline combining existing image processing tools with custom heuristics for bond and atom detection, establishing a foundation for accessible chemical information extraction.
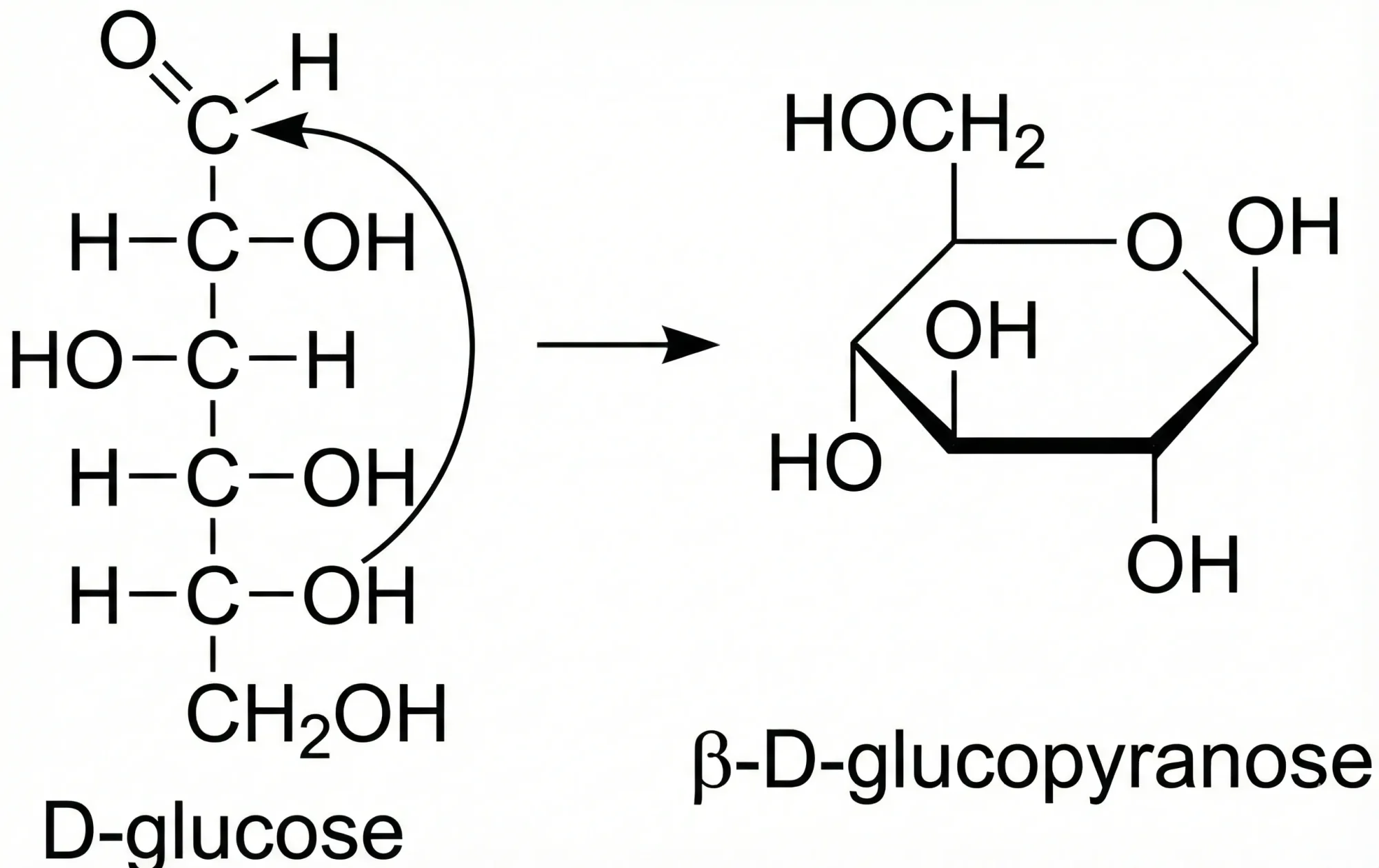
A comprehensive 2020 analysis of the tautomerism problem in chemical databases, compiling 86 tautomeric transformation rules (20 existing, 66 new) and validating them across 400M+ structures to inform algorithmic improvements for InChI V2.
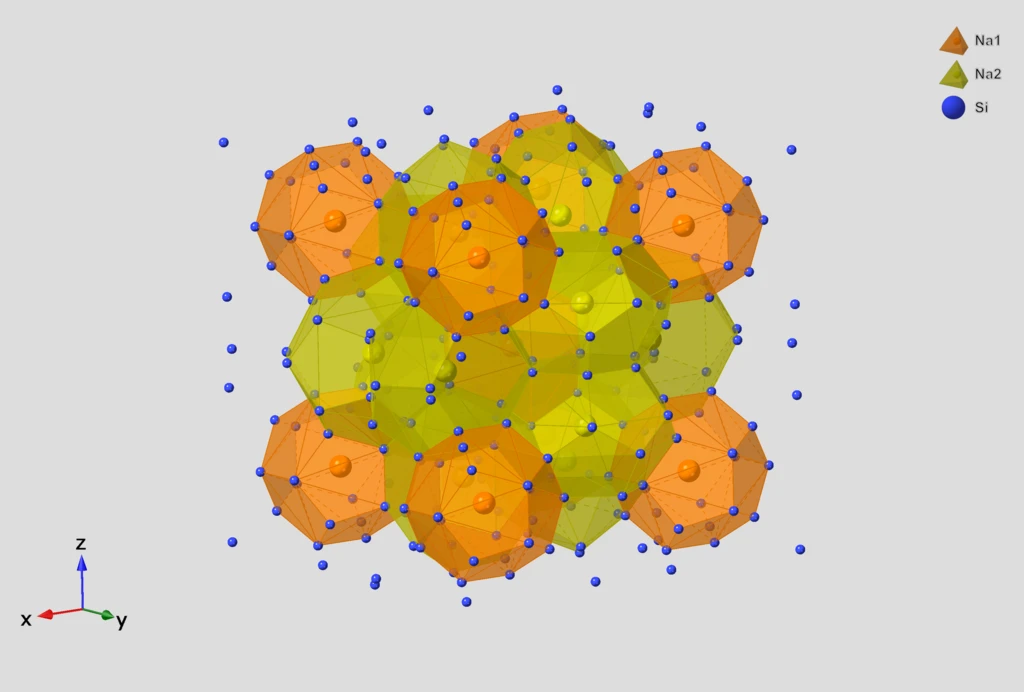
A 2025 Faraday Discussions paper describing the major overhaul of InChI v1.07 that fixed more than 3000 bugs, added support for inorganic and organometallic compounds, and modernized the system to align with FAIR data principles for chemistry databases.

A 2019 format specification introducing two complementary standards for chemical mixtures. Mixfile provides comprehensive mixture descriptions and MInChI provides compact canonical identifiers. This addresses the long-standing lack of standardized machine-readable formats for multi-component chemical systems.
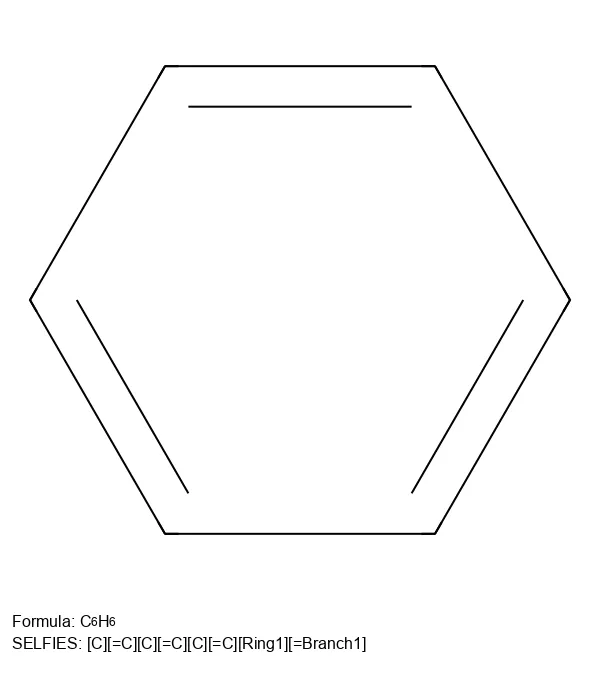
A 2023 software update paper documenting improvements to the SELFIES Python library (v2.1.1), including a streamlined context-free grammar, expanded support for aromatic systems and stereochemistry, customizable semantic constraints, ML utility functions, and performance benchmarks on 300K+ molecules.
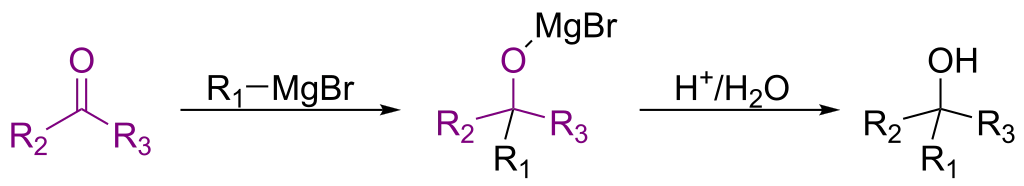
A 2018 infrastructure paper introducing RInChI (Reaction InChI), the first standardized format for uniquely identifying chemical reactions through algorithmic hashing and layering, enabling reaction database searching and duplicate detection analogous to how InChI works for individual molecules.
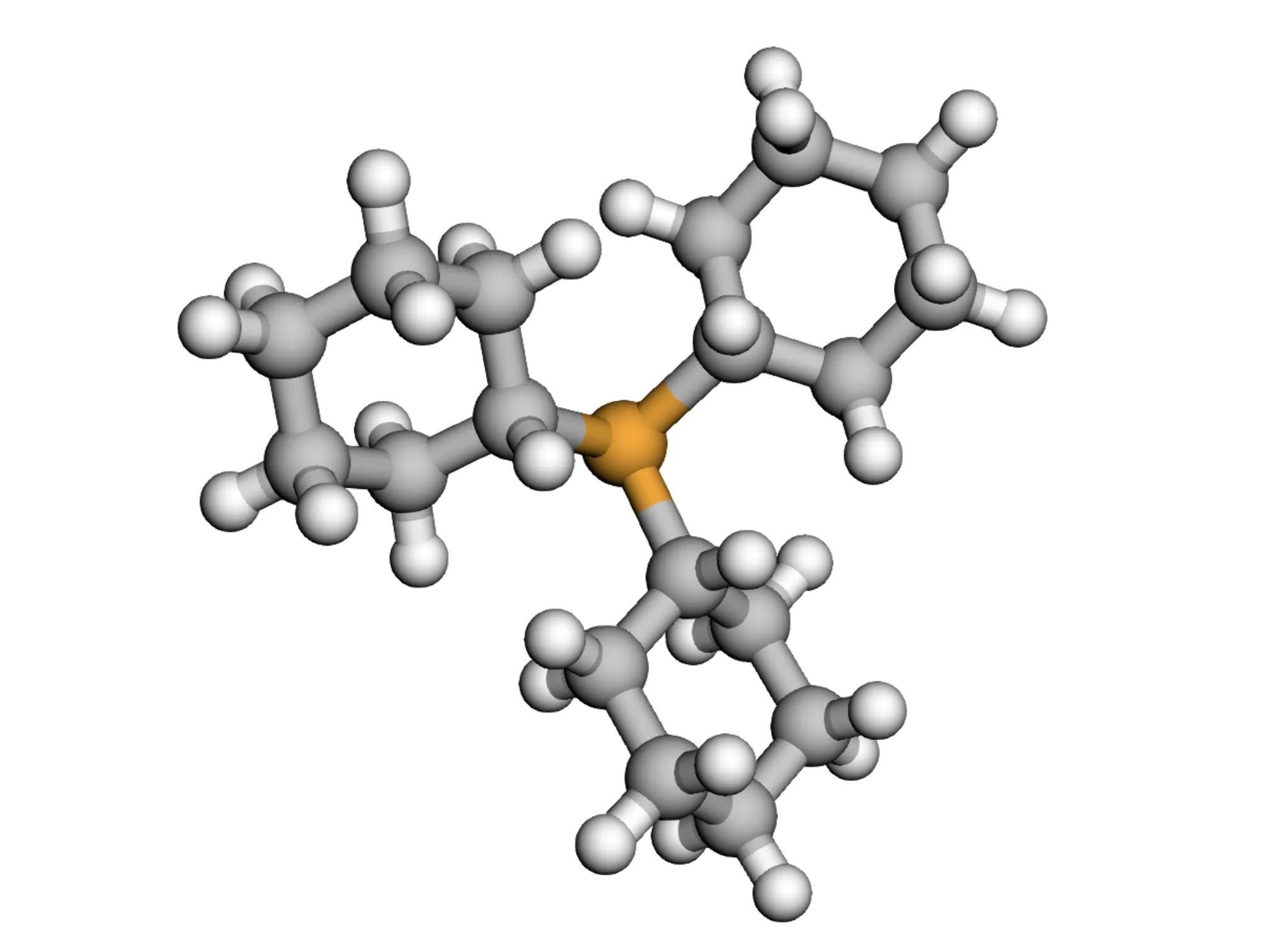
MARCEL provides a comprehensive benchmark for molecular representation learning with 722K+ conformers across four diverse subsets (Drugs-75K, Kraken, EE, BDE), enabling evaluation of conformer ensemble methods for property prediction in drug discovery and catalysis.