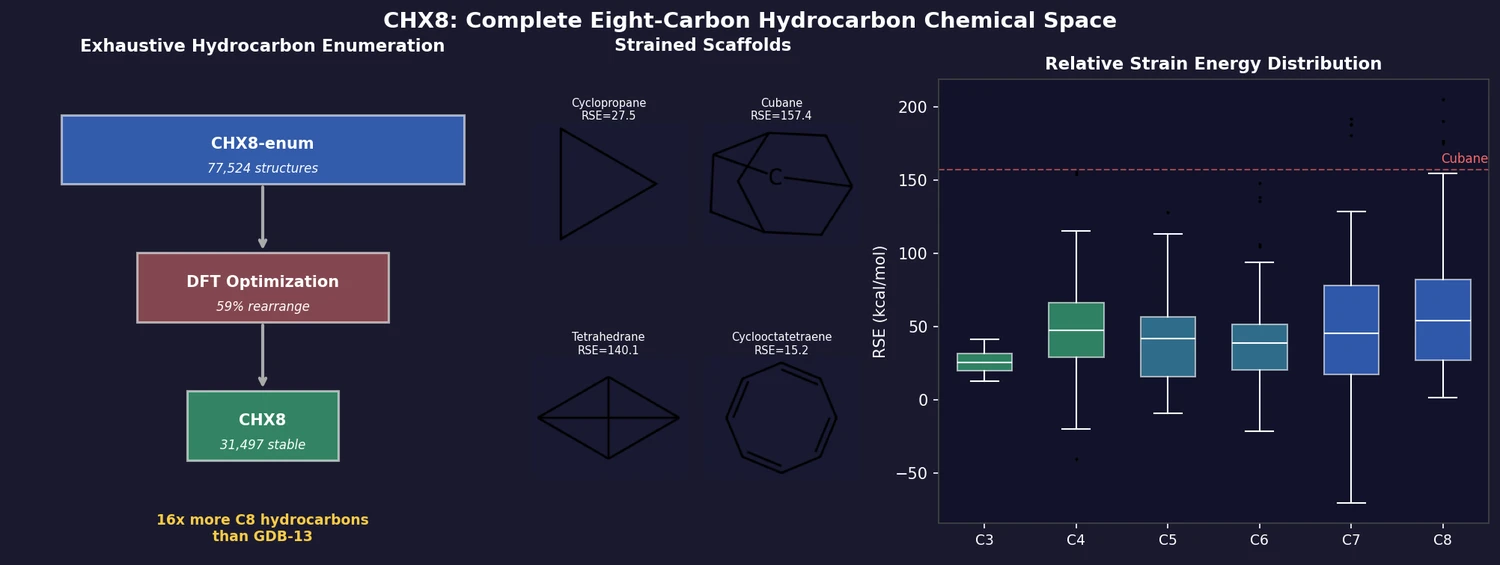
CHX8: Complete Eight-Carbon Hydrocarbon Space
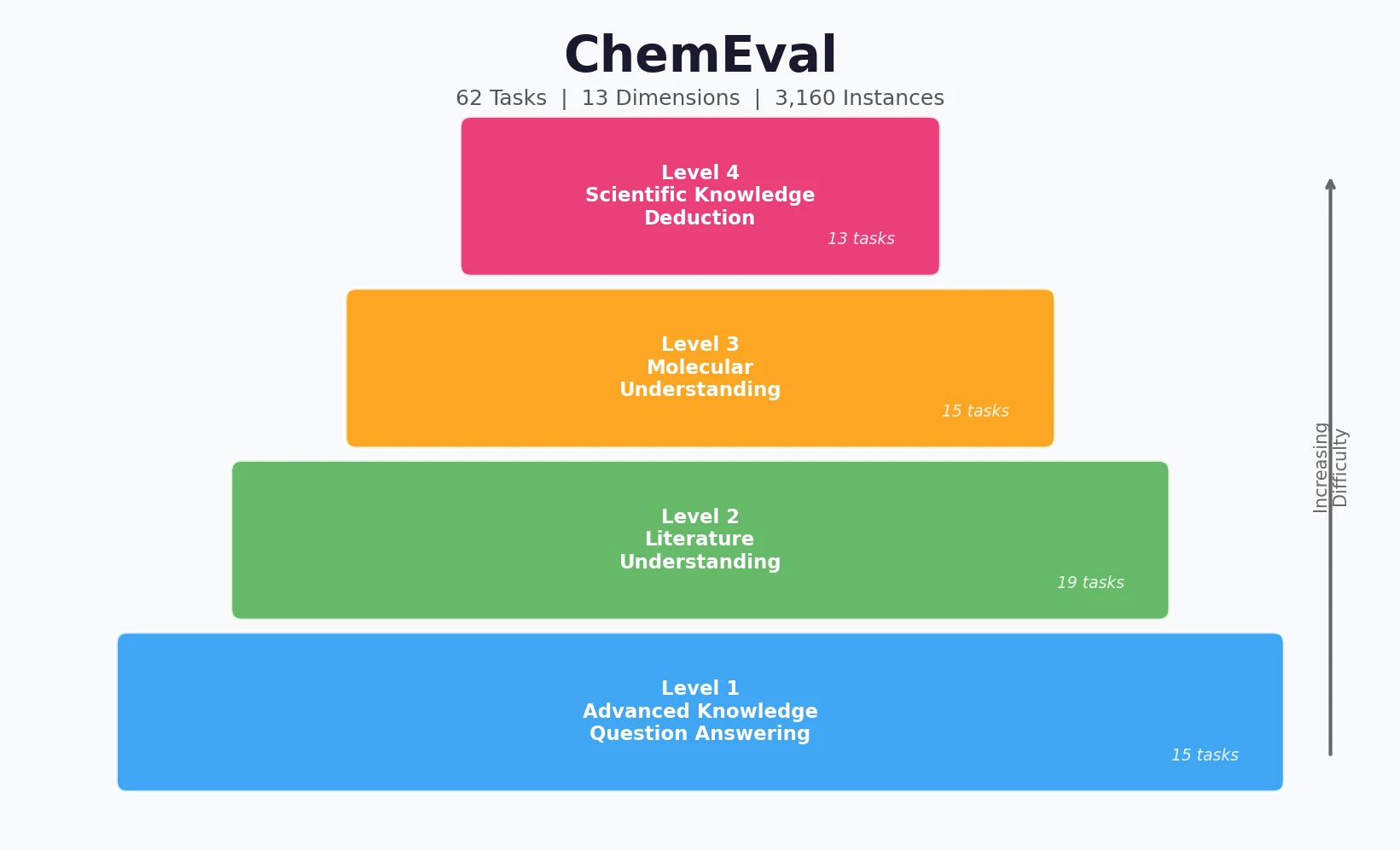
CHX8 exhaustively enumerates all mathematically feasible hydrocarbons with up to eight carbon atoms (77,524 structures), then DFT-optimizes them to identify 31,497 stable molecules. A universal relative strain energy (RSE) metric referenced to cyclohexane serves as a synthesizability proxy. CHX8 covers 16x more C8 hydrocarbons than GDB-13 and reveals that over 90% of novel structures should be synthetically accessible.