X-MOL: Pre-training on 1.1B Molecules for SMILES
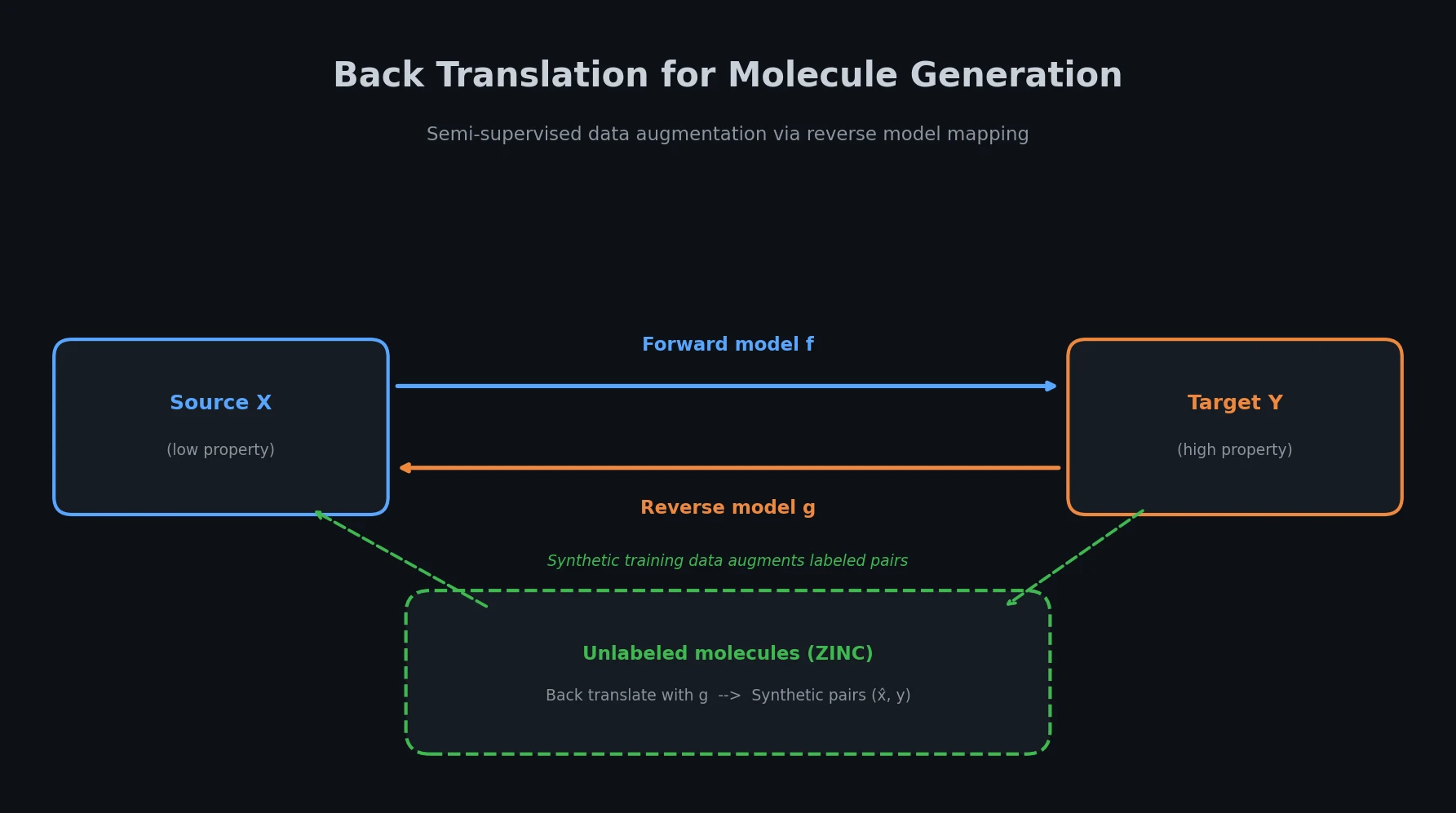
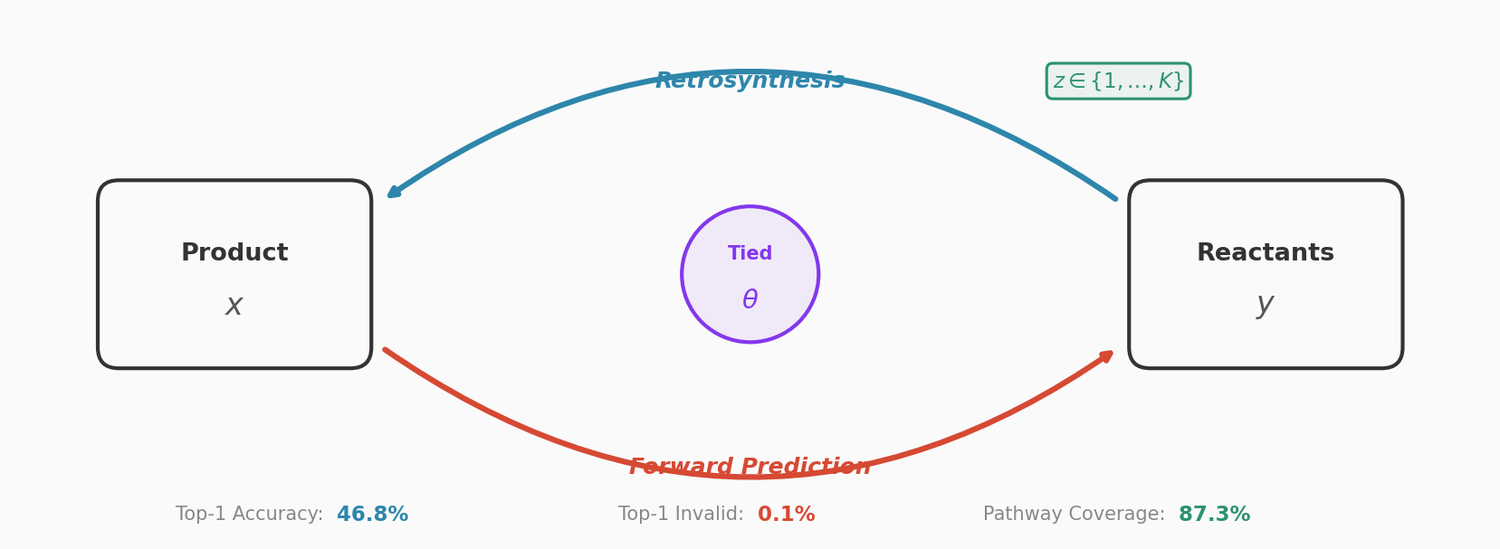
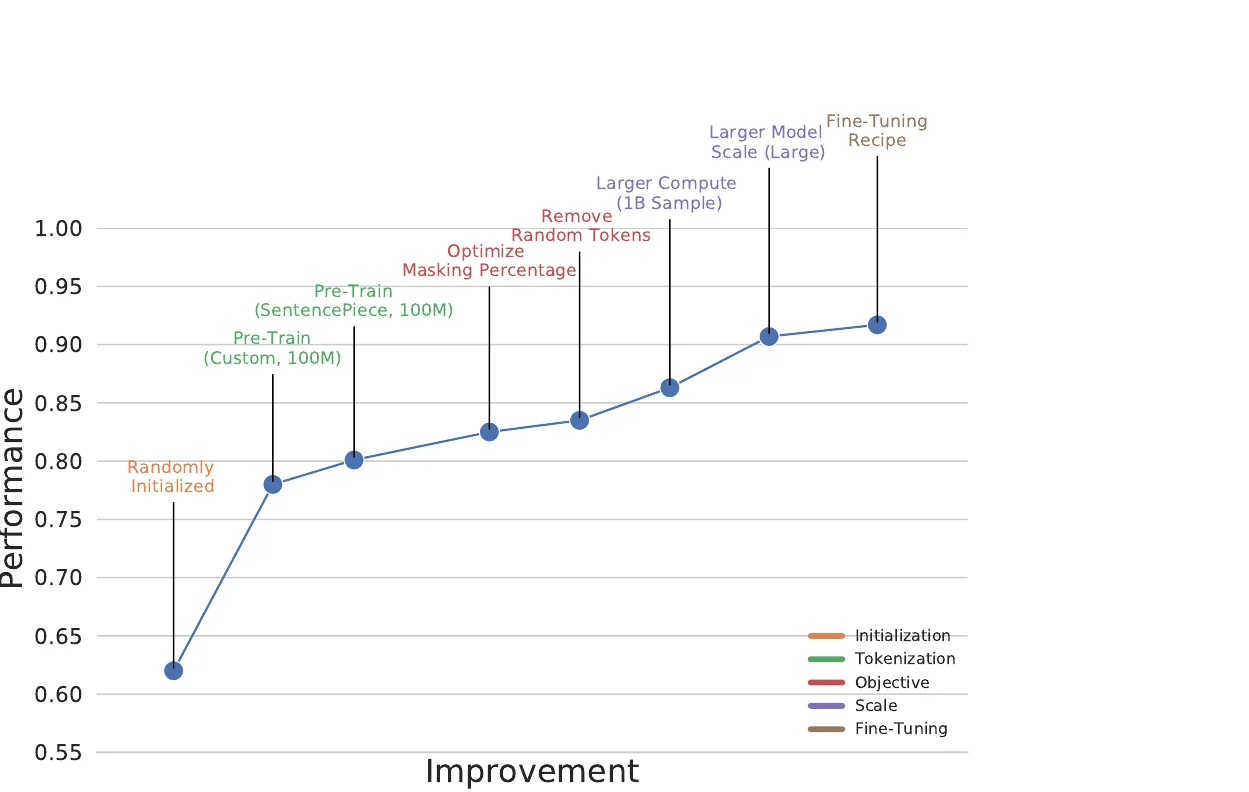
X-MOL applies large-scale Transformer pre-training on 1.1 billion molecules with a generative SMILES-to-SMILES strategy, then fine-tunes for five molecular analysis tasks including property prediction, reaction analysis, and de novo generation.