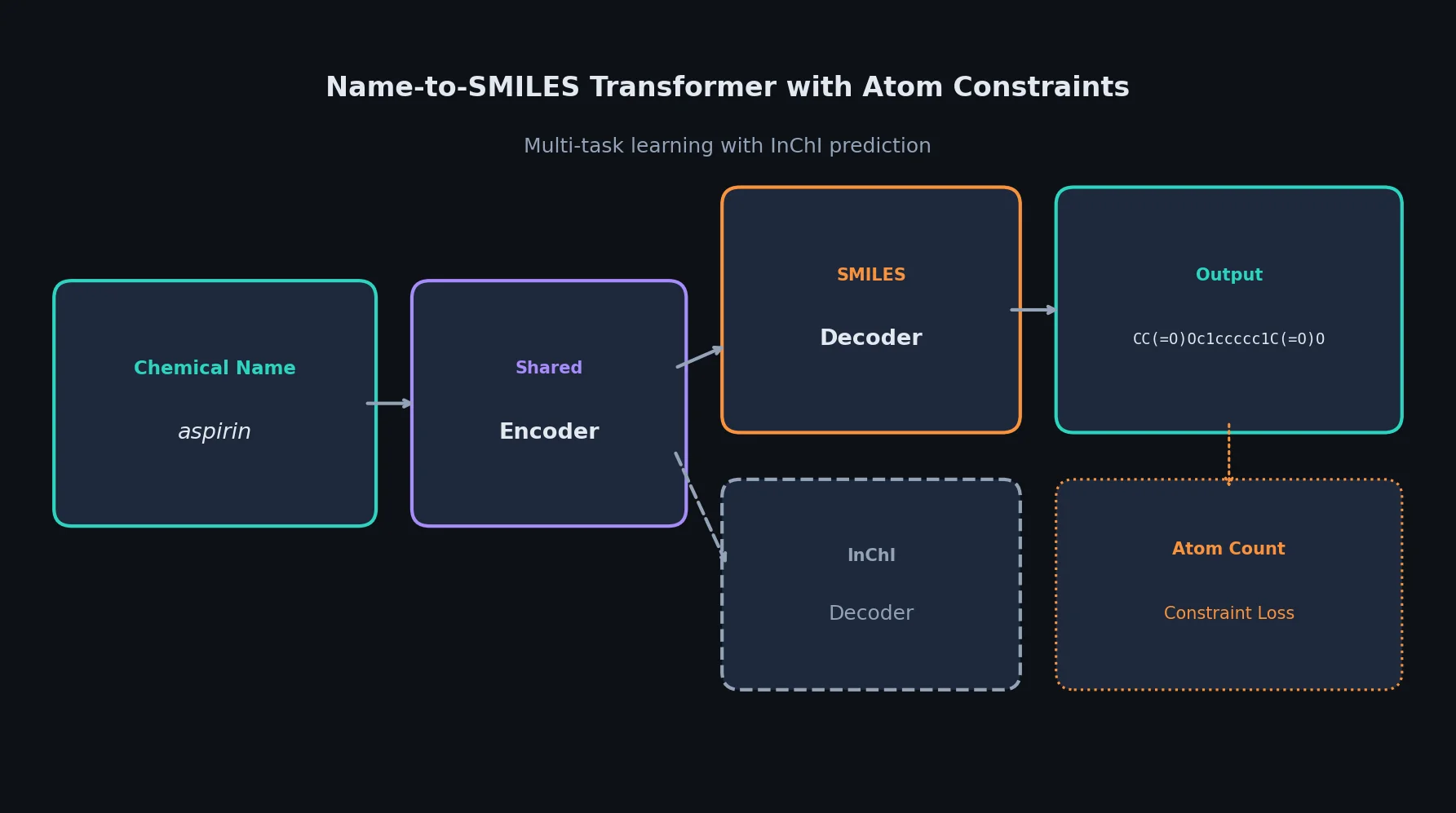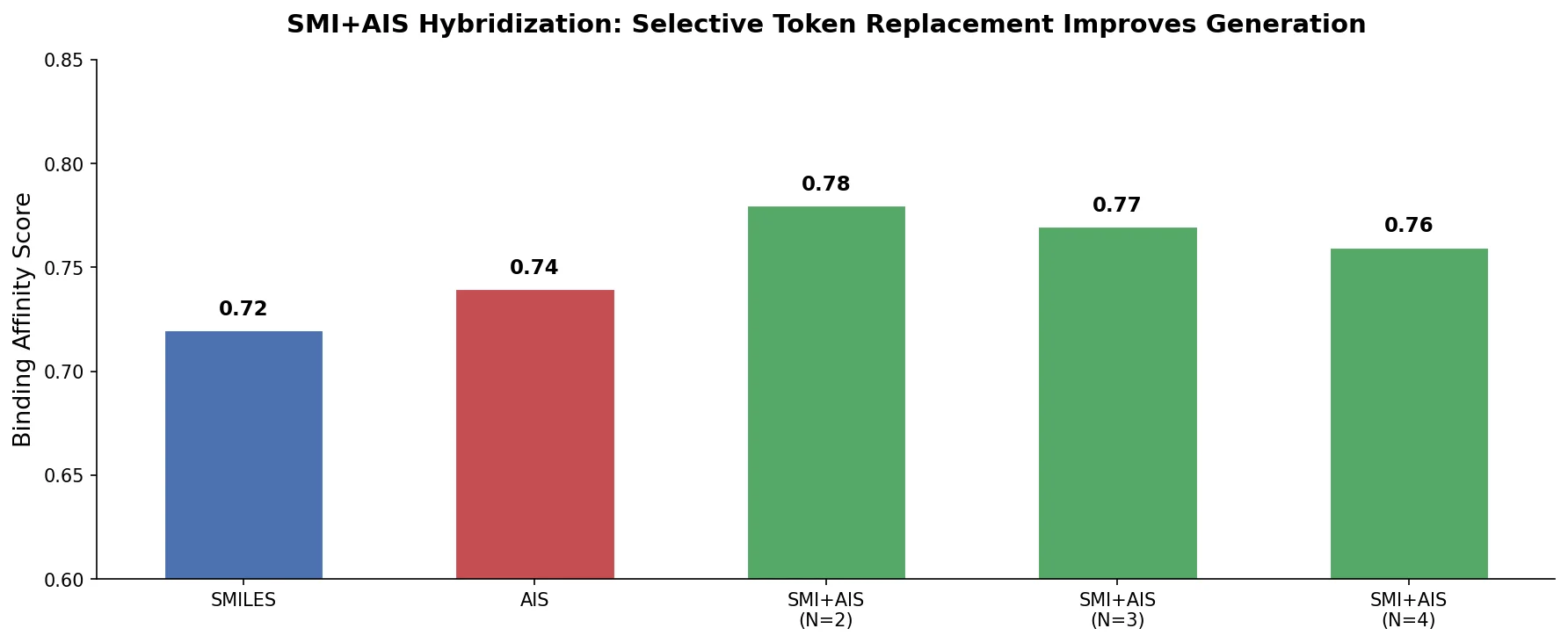
SMI+AIS: Hybridizing SMILES with Environment Tokens
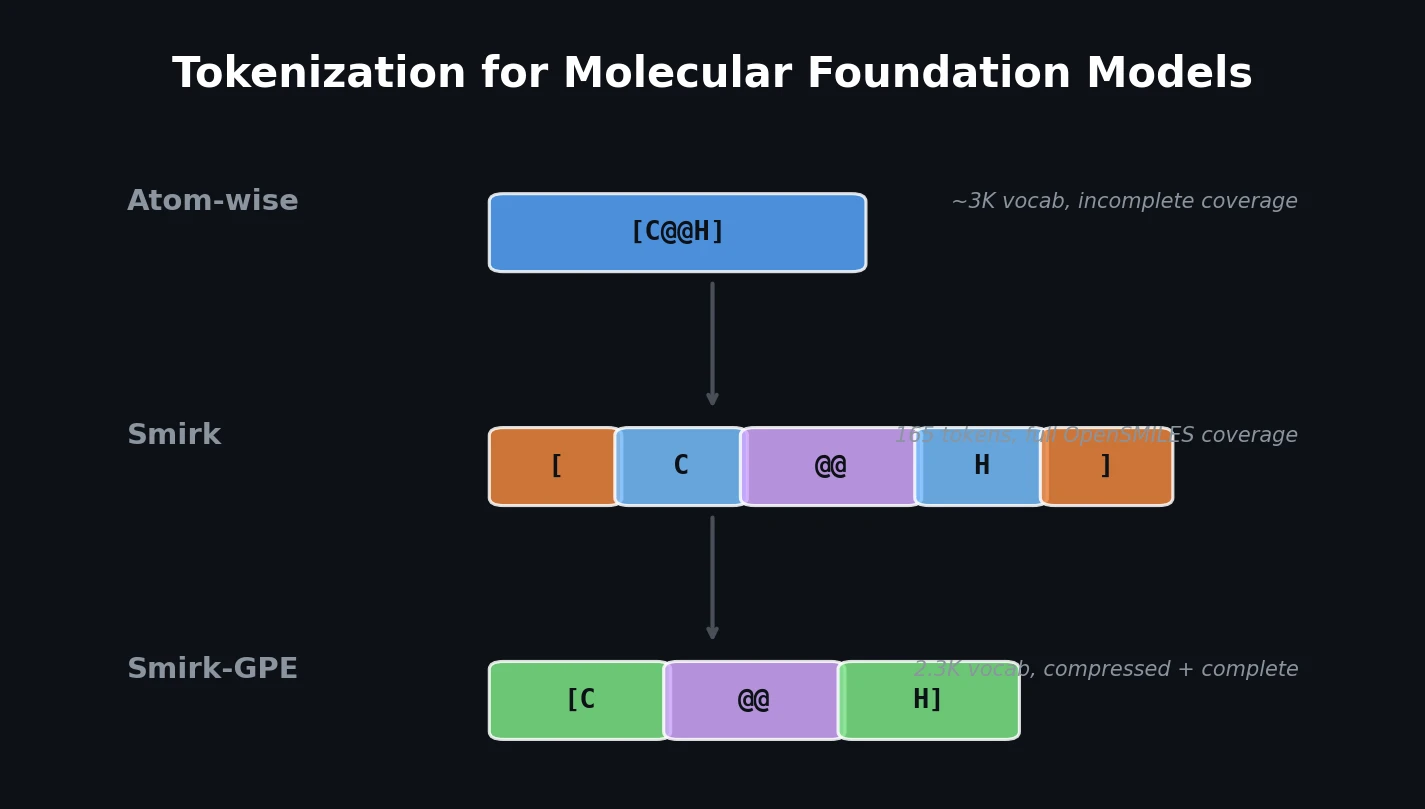
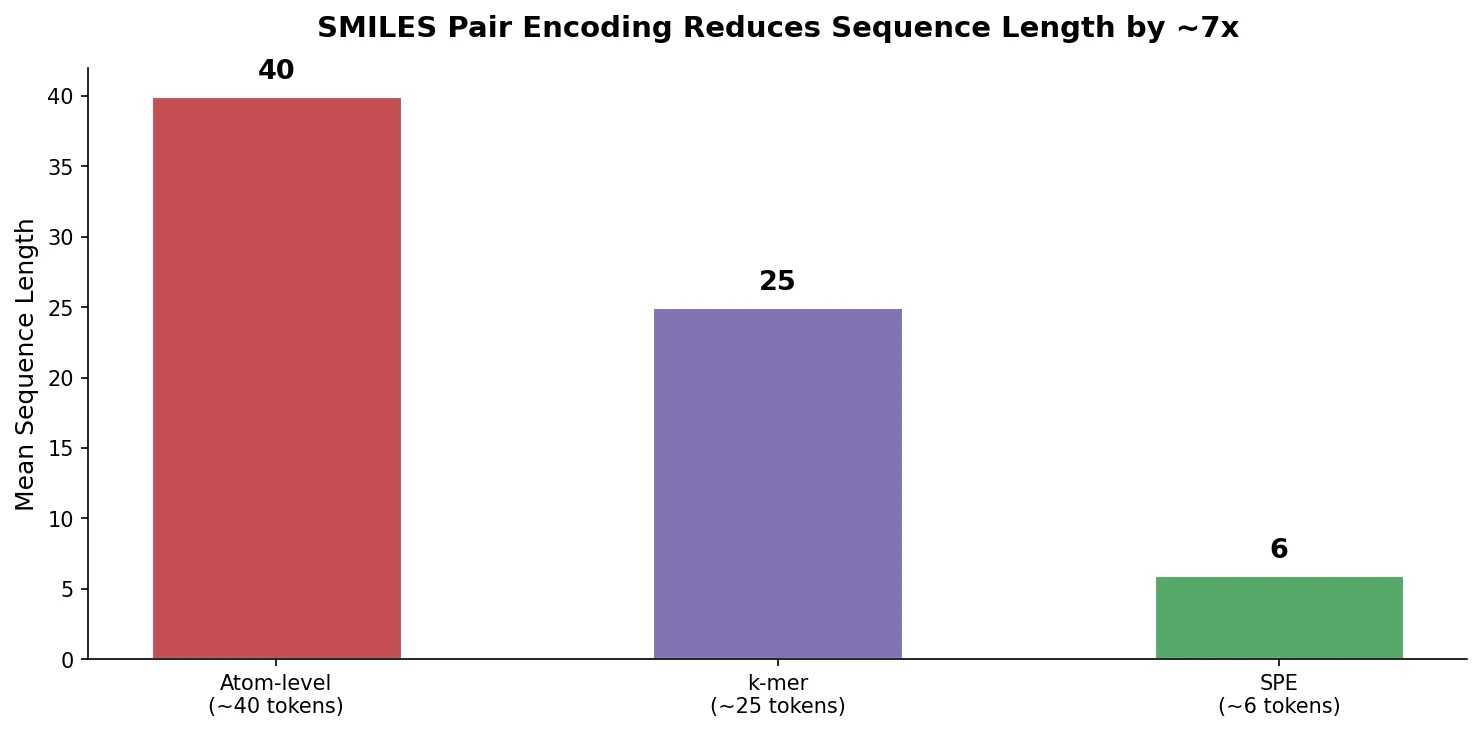
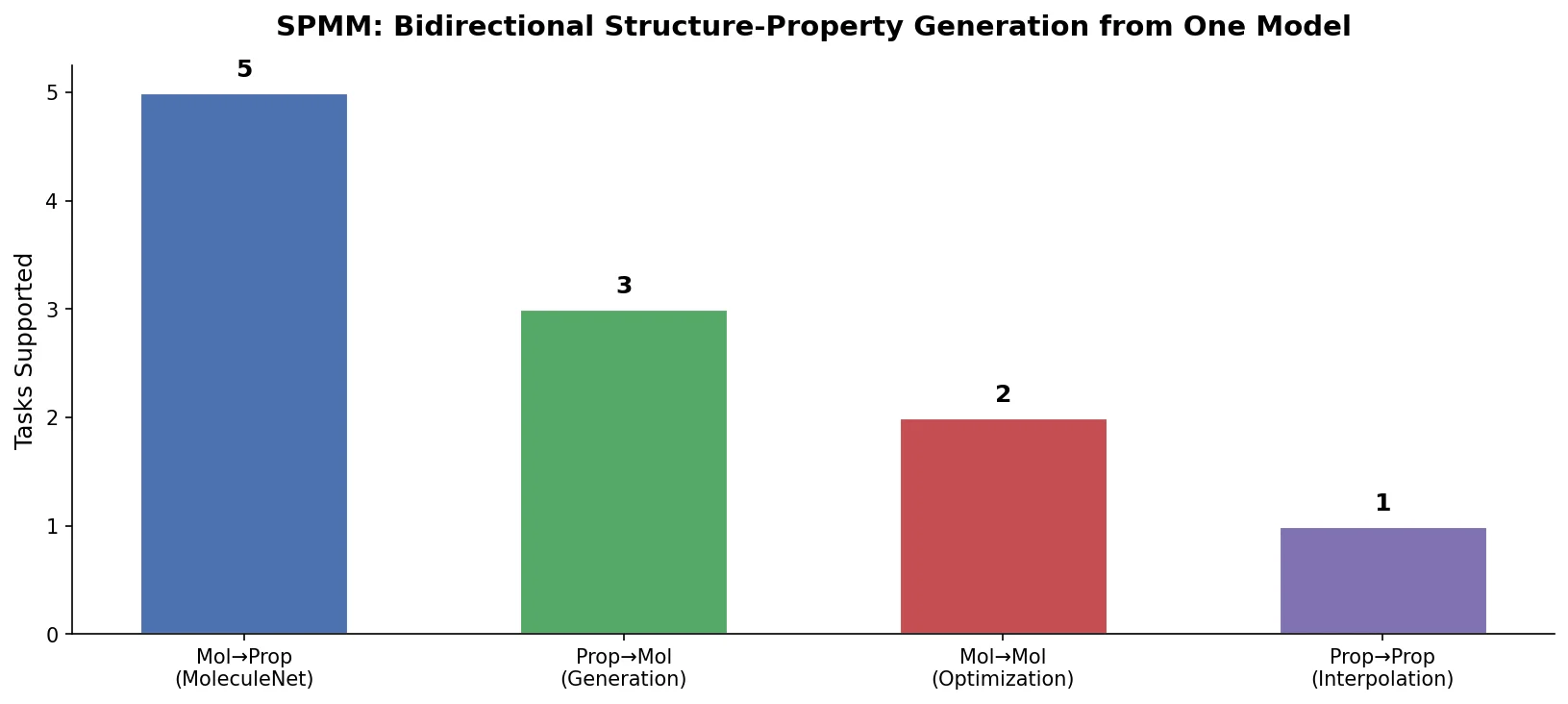
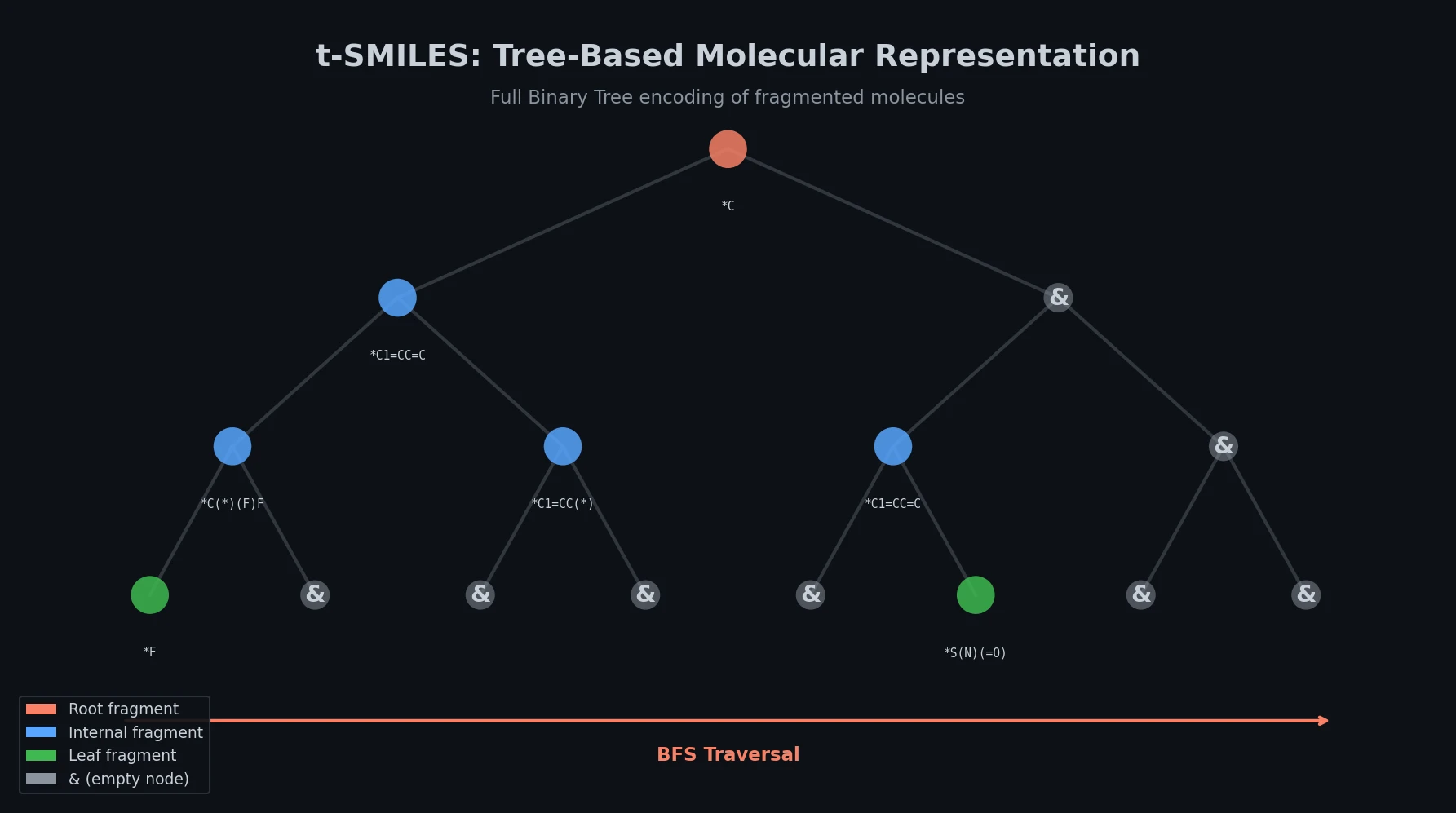
Proposes SMI+AIS, a hybrid molecular representation combining standard SMILES tokens with chemical-environment-aware Atom-In-SMILES tokens, demonstrating improved molecular generation for drug design targets.