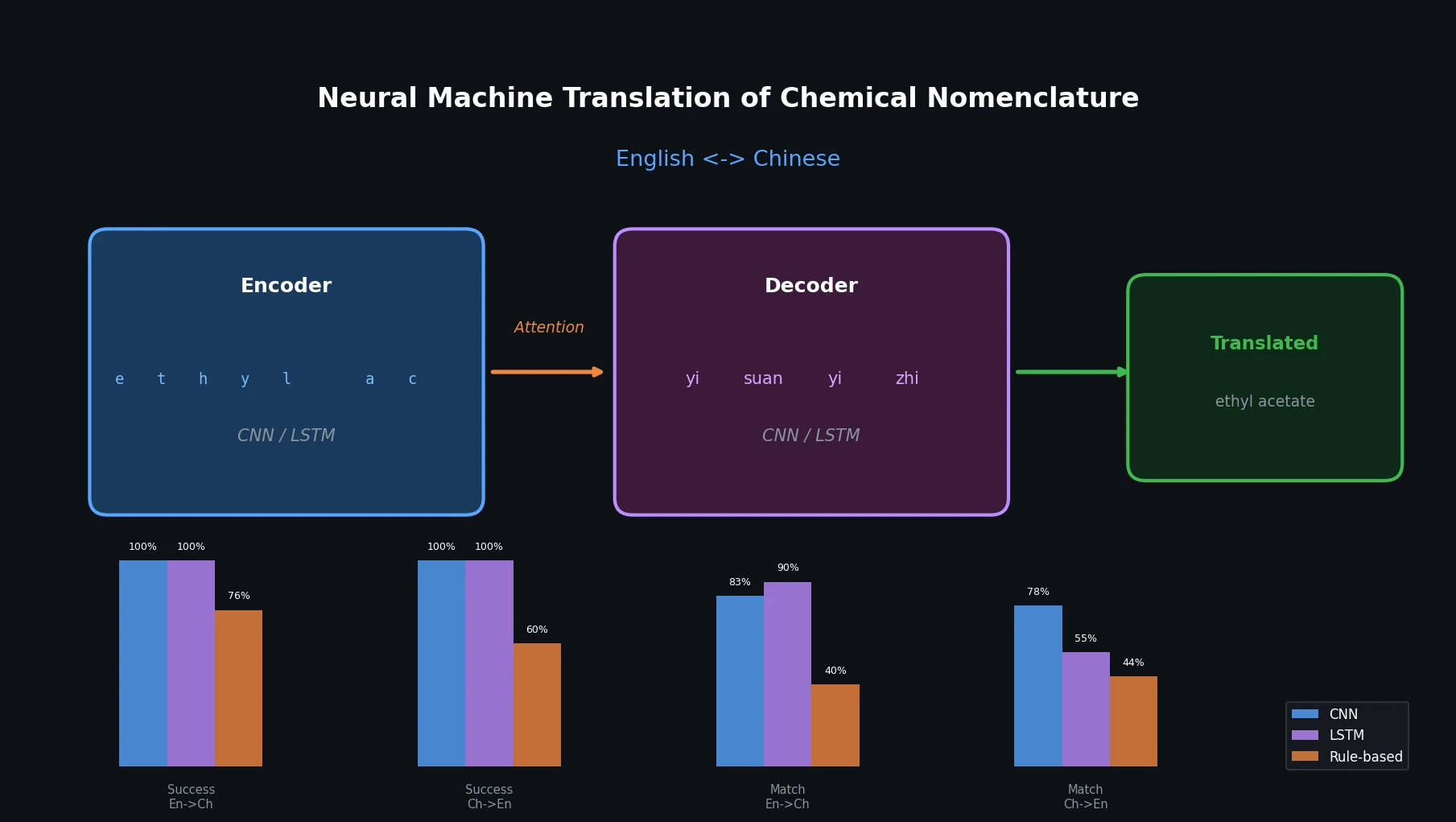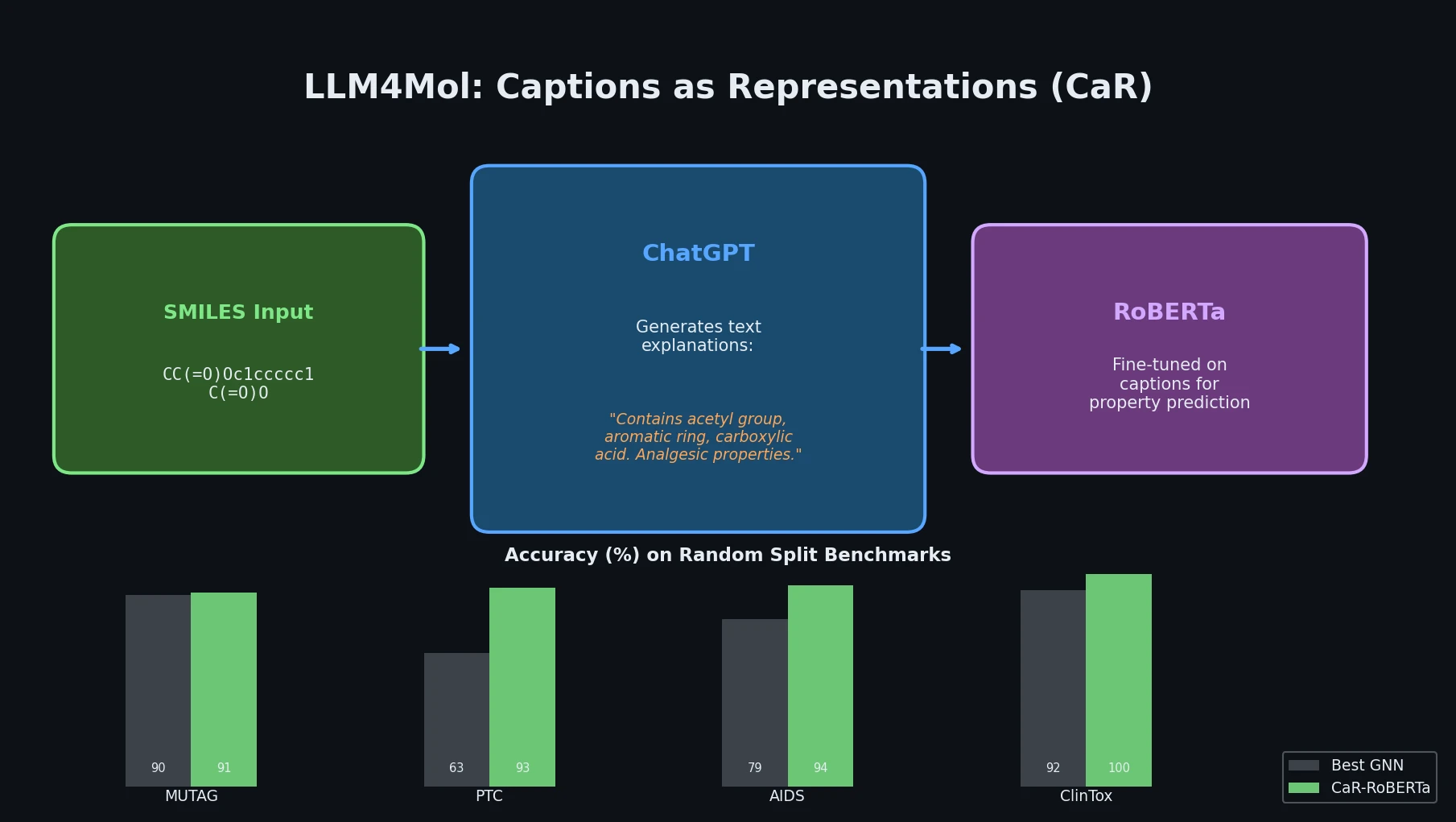
LLM4Mol: ChatGPT Captions as Molecular Representations
Proposes Captions as Representations (CaR), where ChatGPT generates textual explanations for SMILES strings that are then used to fine-tune small language models for molecular property prediction.