Augmented Hill-Climb for RL-Based Molecule Design
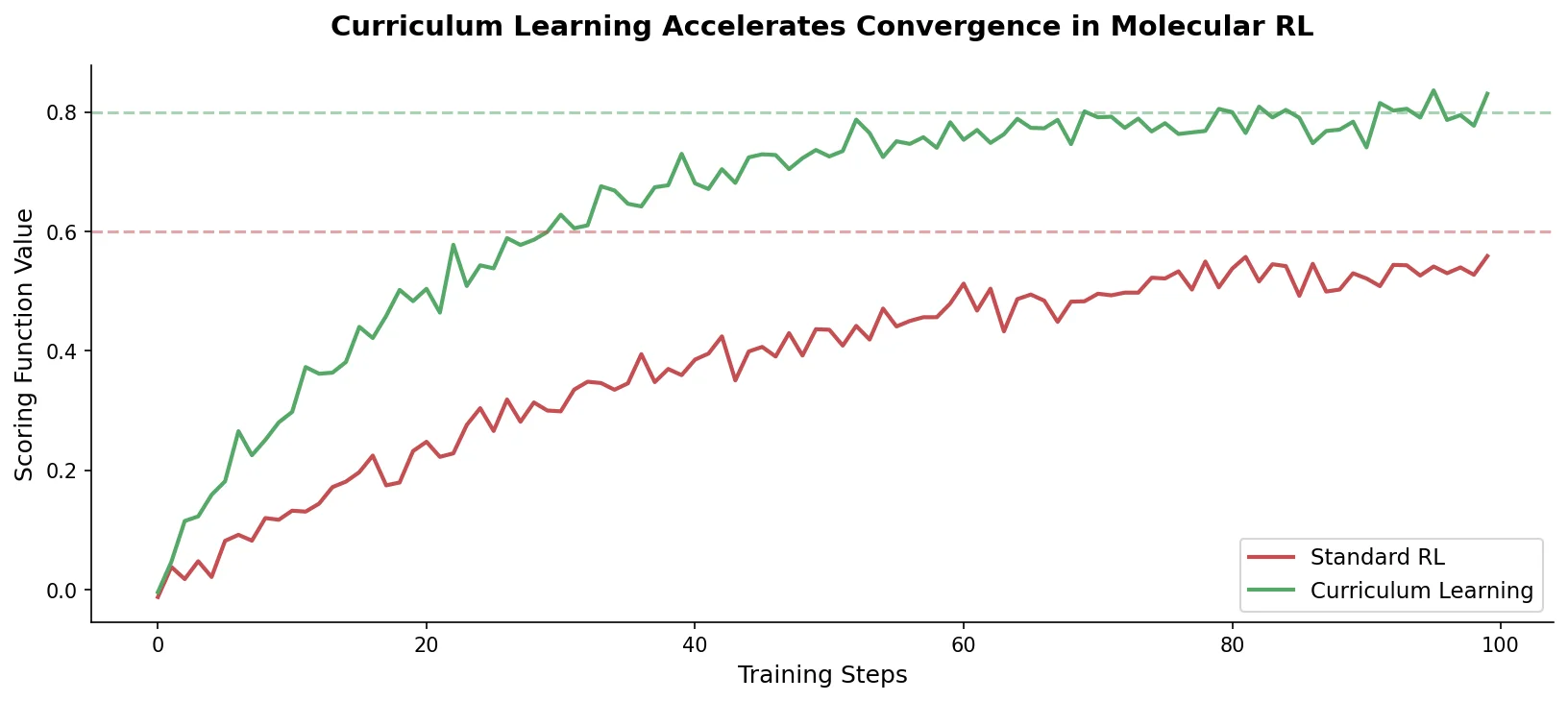
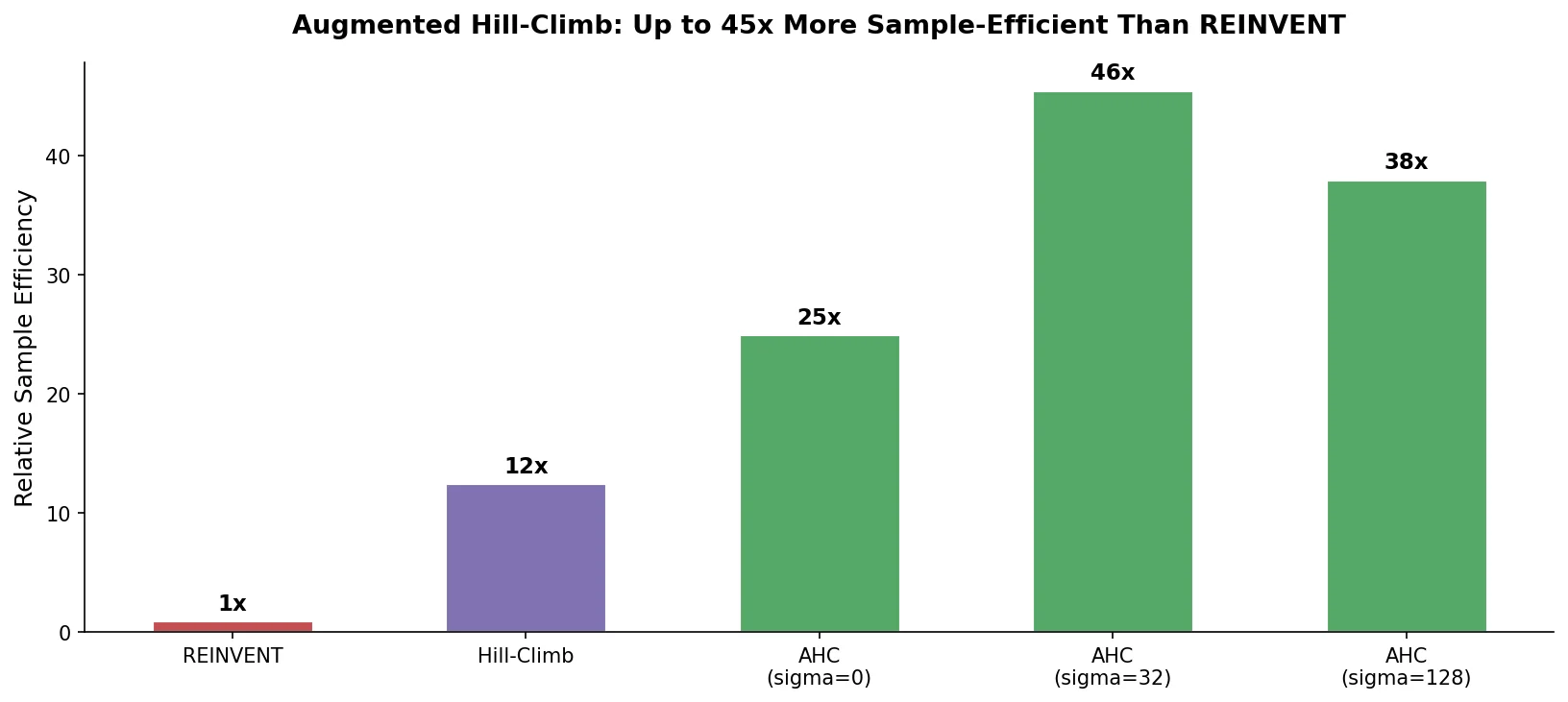
Proposes Augmented Hill-Climb, a hybrid RL strategy for SMILES-based generative models that improves sample efficiency ~45-fold over REINVENT by filtering low-scoring molecules from the loss computation, with diversity filters to prevent mode collapse.