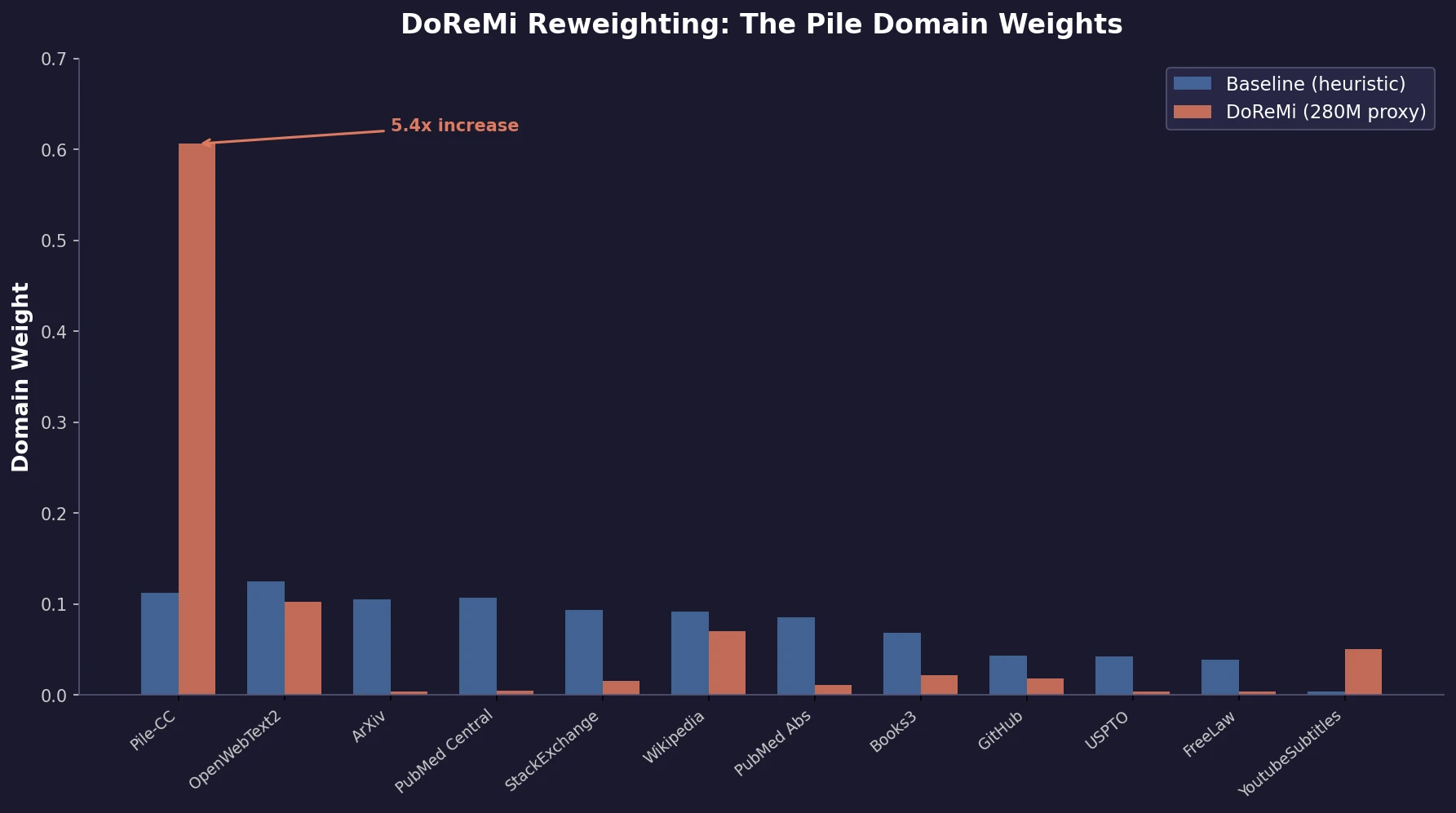
DoReMi: Optimizing Data Mixtures for LM Pretraining
Xie et al. propose DoReMi, which trains a 280M proxy model using Group DRO to find optimal domain mixture weights, then uses those weights to train an 8B model 2.6x faster with 6.5% better downstream accuracy.