GraphReco: Probabilistic Structure Recognition (2026)
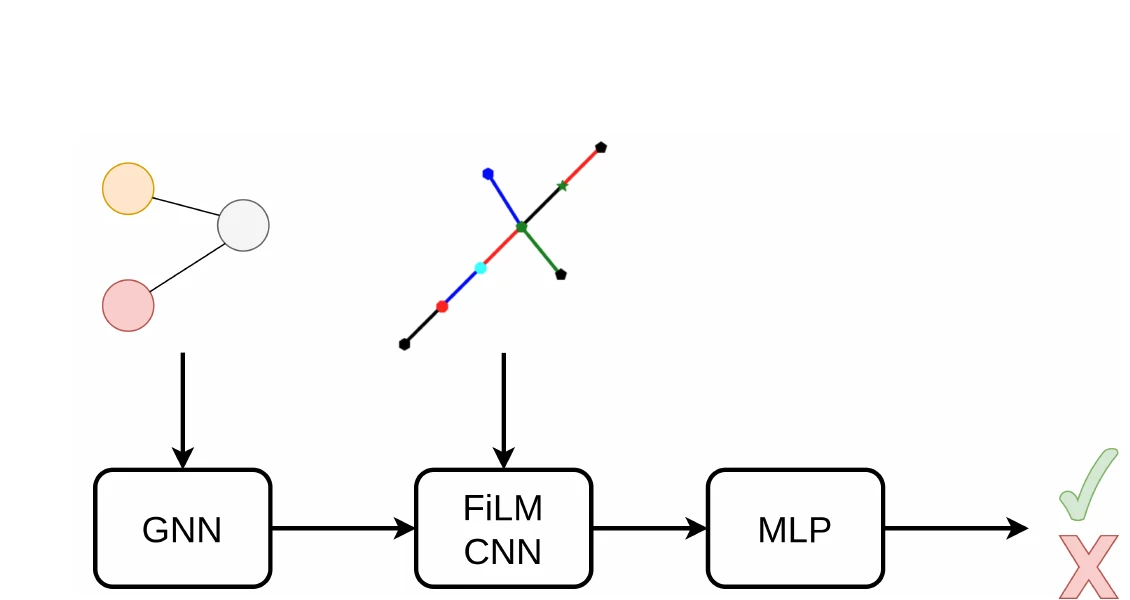
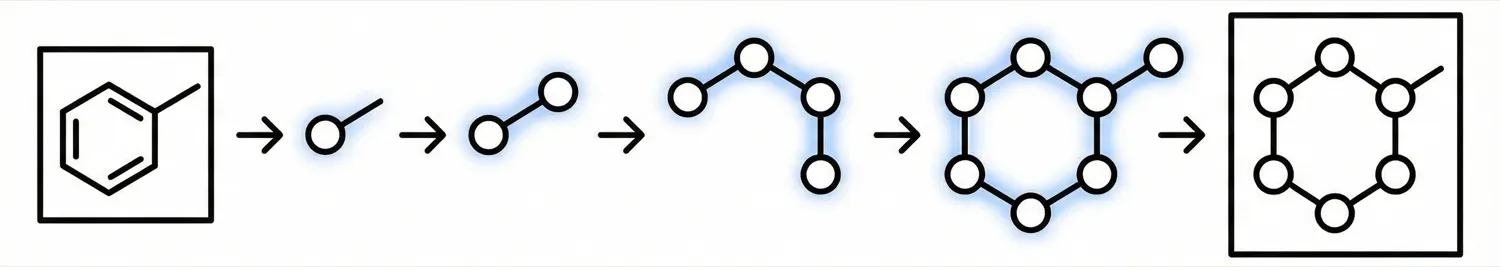
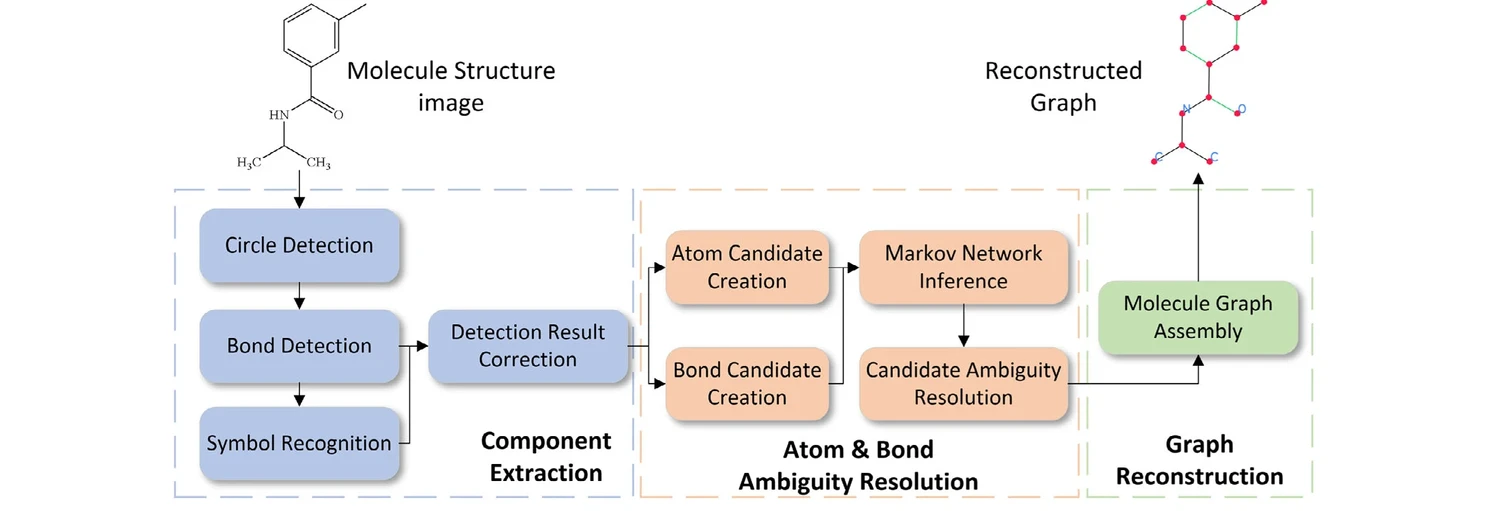
GraphReco presents a rule-based OCSR system with two key innovations: a Fragment Merging line detection algorithm for precise bond identification and a Markov network for probabilistic resolution of atom/bond ambiguity during graph assembly. Achieves 94.2% accuracy on USPTO-10K, outperforming both traditional rule-based and some ML-based methods.