Data Mixing Laws for LM Pretraining Optimization
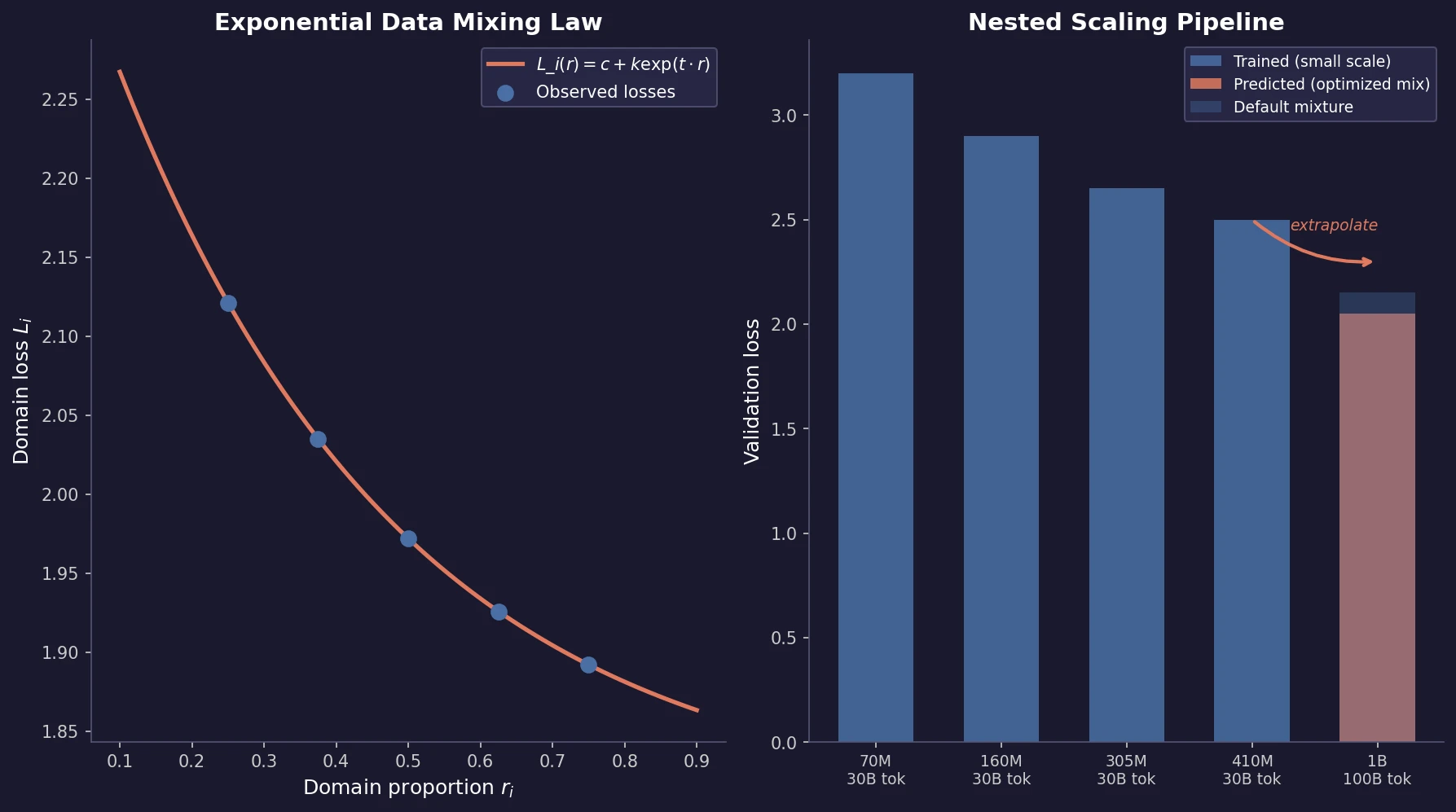
Ye et al. find that language model loss on each domain follows an exponential function of training mixture proportions. By nesting data mixing laws with scaling laws for steps and model size, small-scale experiments can predict and optimize mixtures for large models, achieving 48% training efficiency gains.