A Review of Optical Chemical Structure Recognition Tools
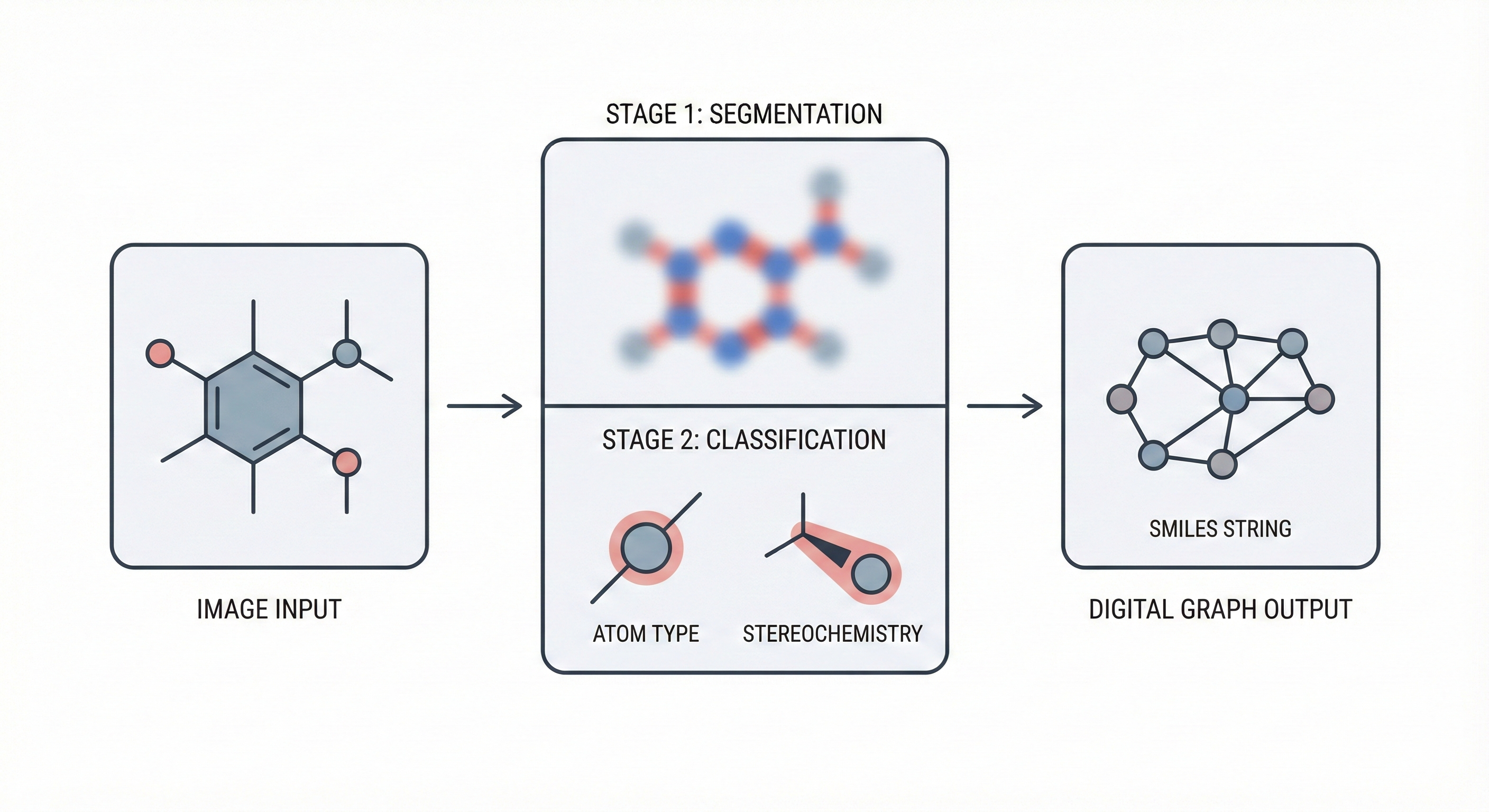
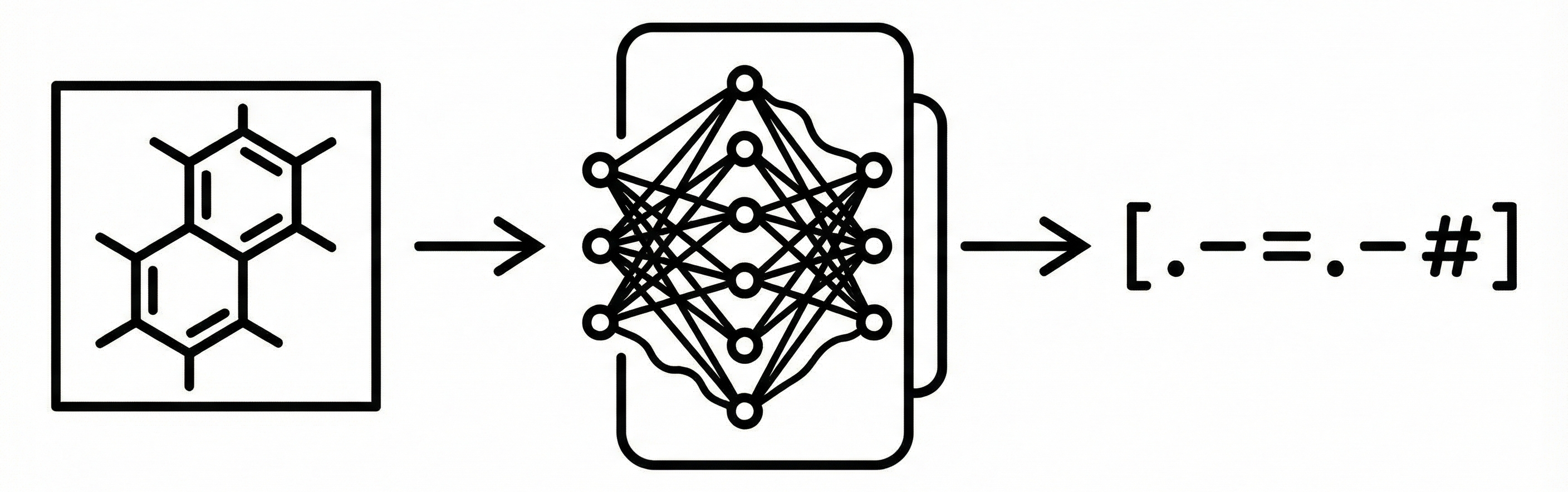
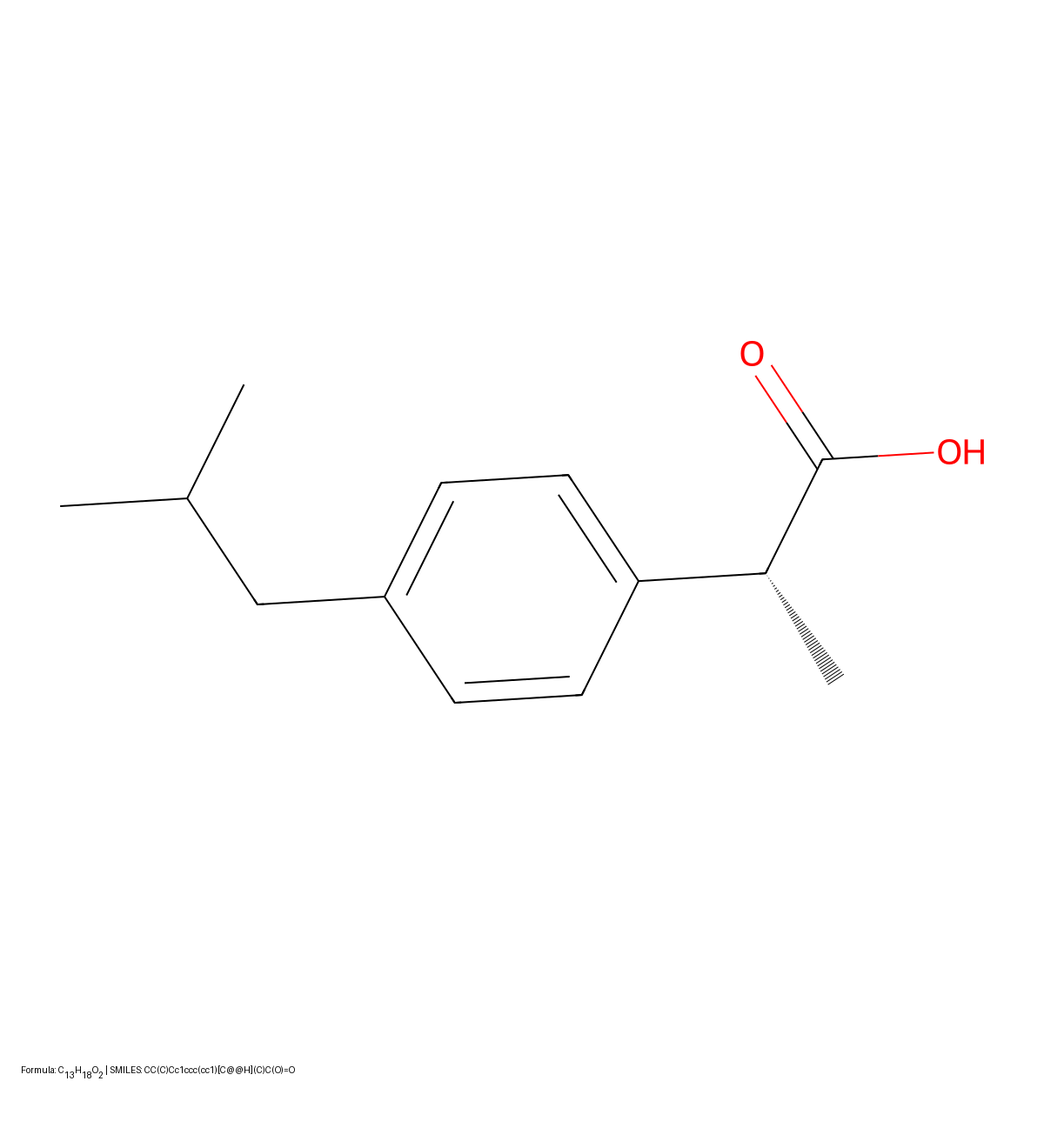
This paper reviews three decades of OCSR development, transitioning from rule-based heuristics to early deep learning approaches. It includes a benchmark study comparing the performance of three open-source tools (OSRA, Imago, MolVec) on four diverse datasets.