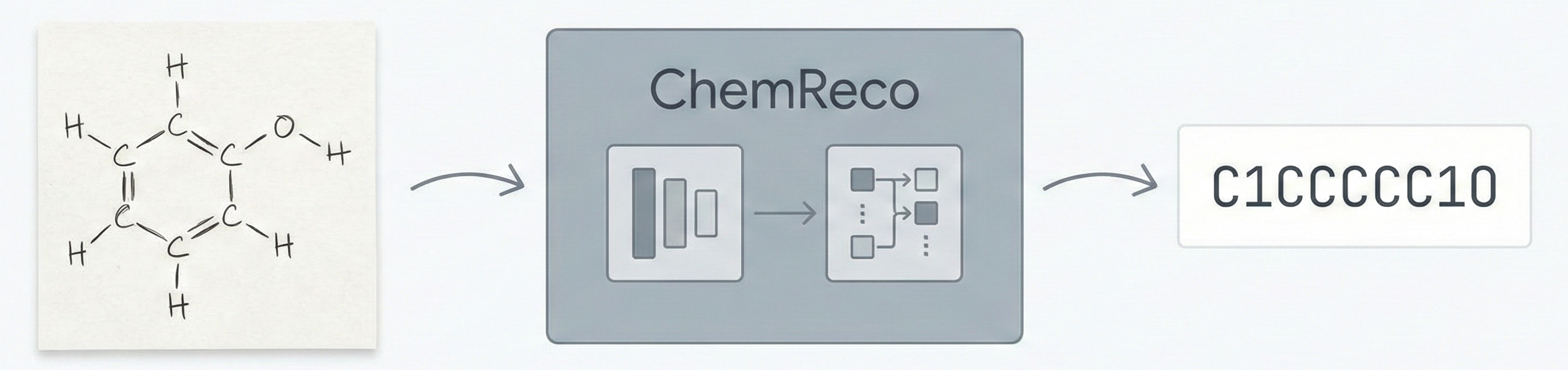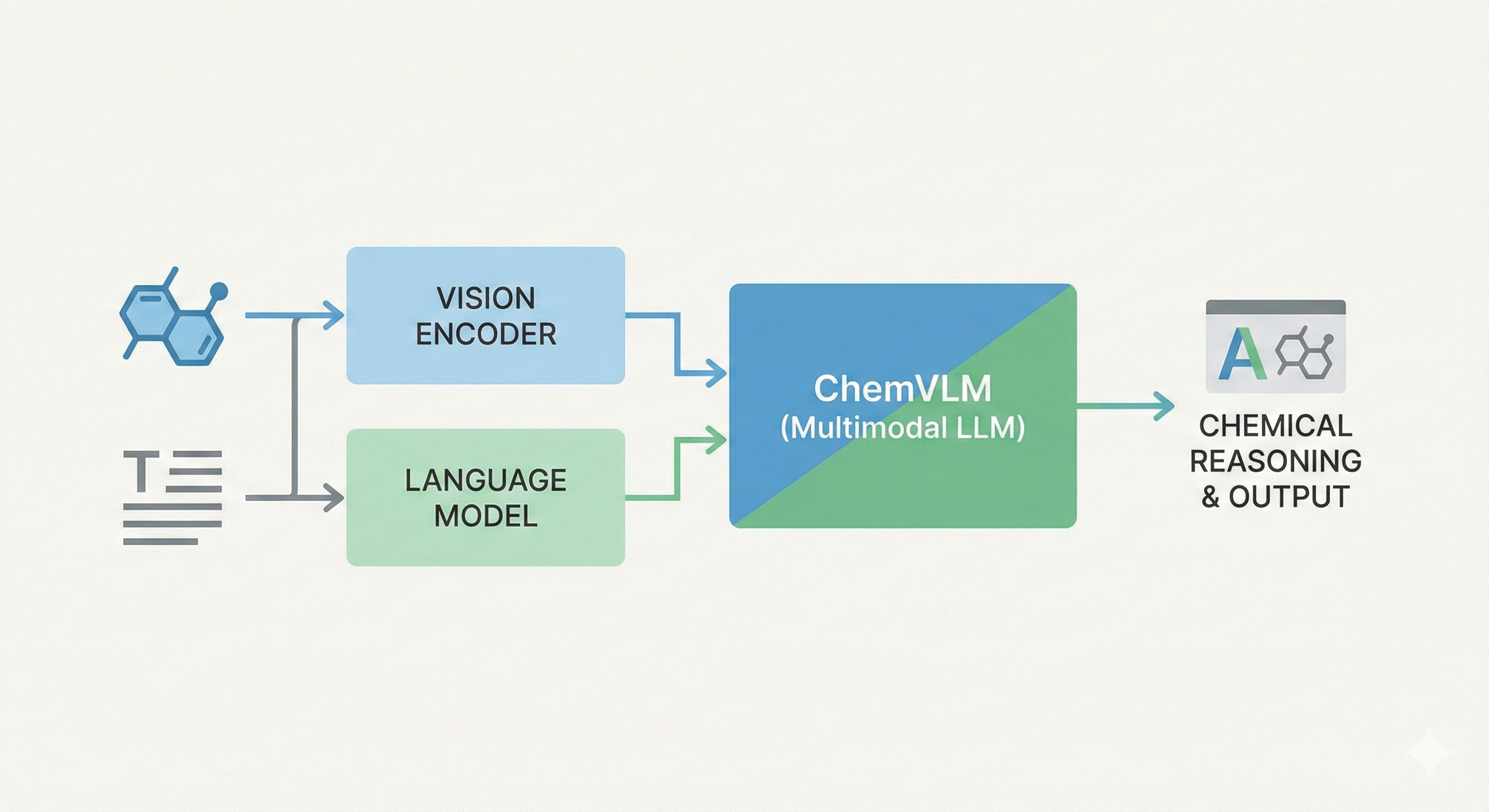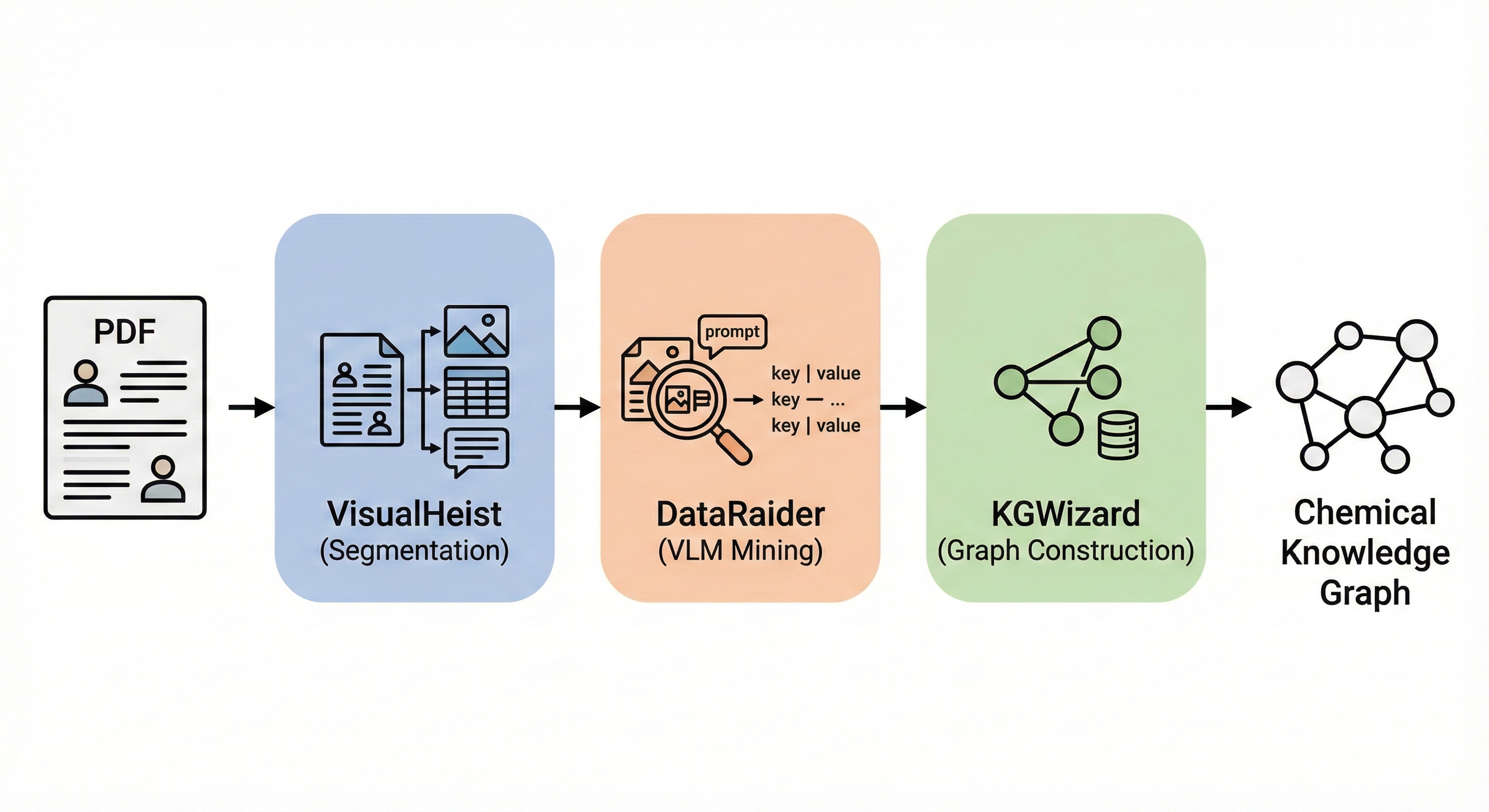
MERMaid: Multimodal Chemical Reaction Mining from PDFs
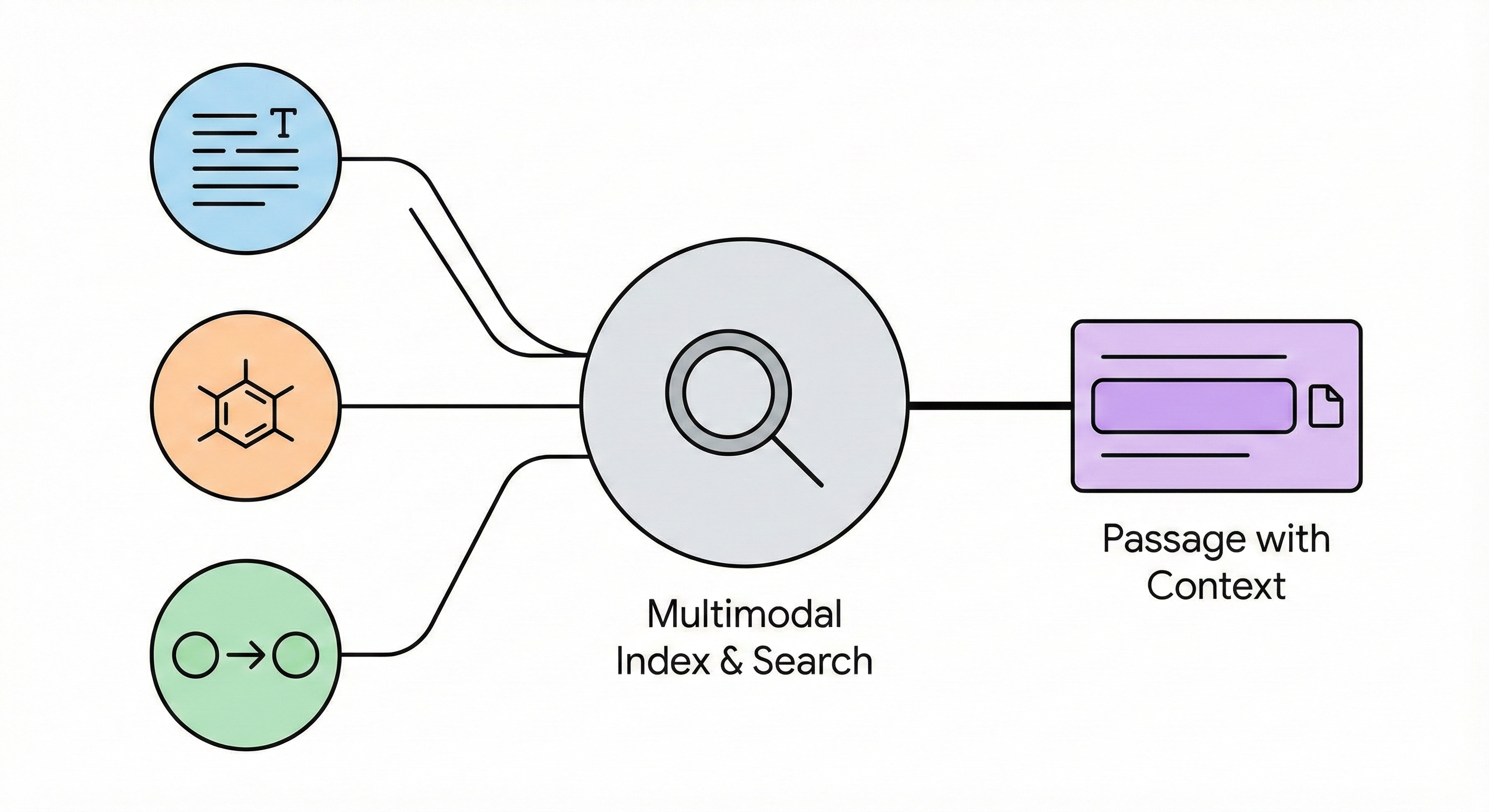
MERMaid leverages fine-tuned vision models and VLM reasoning to mine chemical reaction data directly from PDF figures and tables. By handling context inference and coreference resolution, it builds high-fidelity knowledge graphs with 87% end-to-end accuracy.