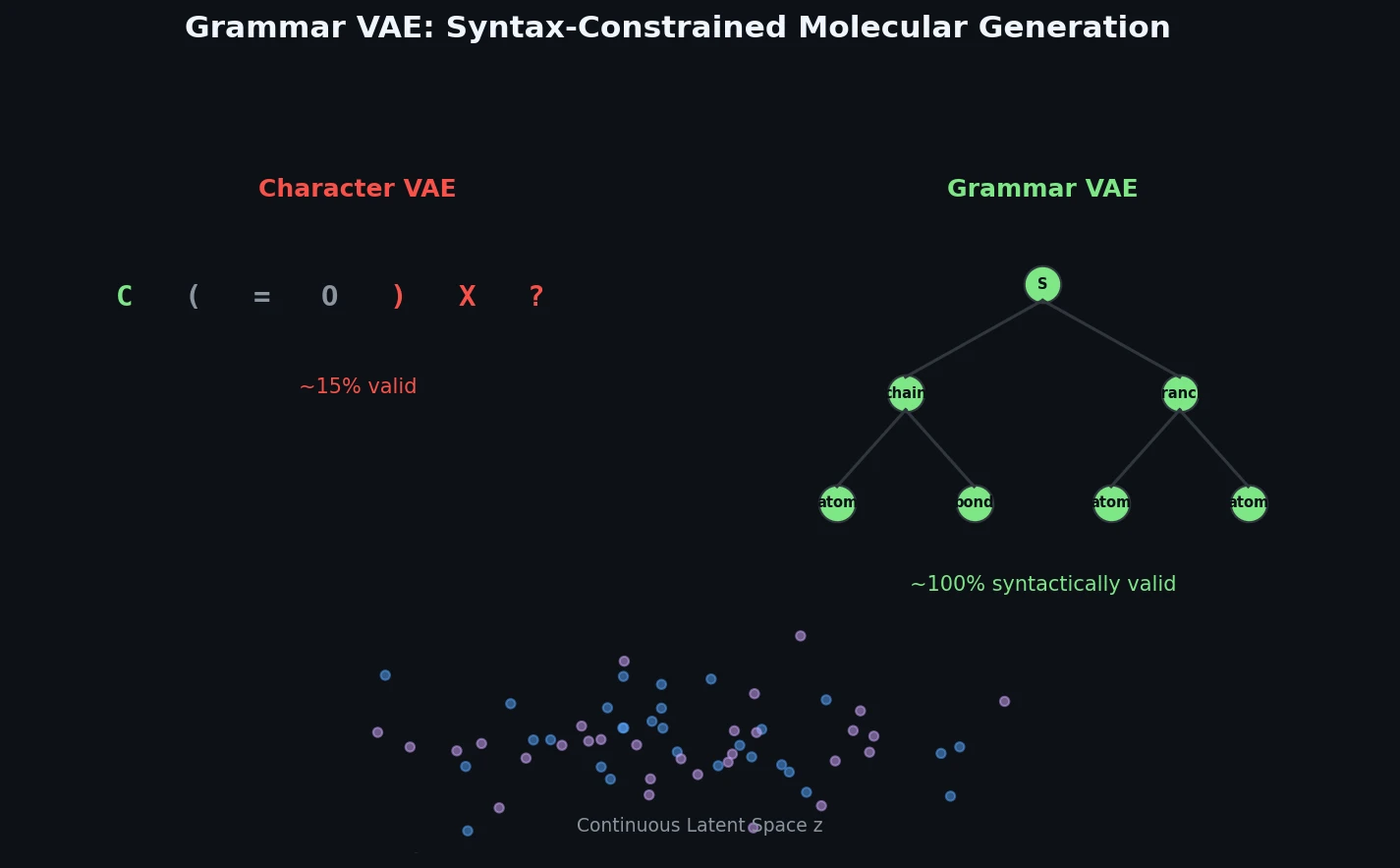
Grammar VAE: Generating Valid Molecules via CFGs
The Grammar VAE replaces character-level decoding with context-free grammar production rules, using a stack-based masking mechanism to guarantee that all generated SMILES strings are syntactically valid. Applied to molecular optimization and symbolic regression, it learns smoother latent spaces and finds better molecules than character-level baselines.