A Convexity Principle for Interacting Gases (McCann 1997)
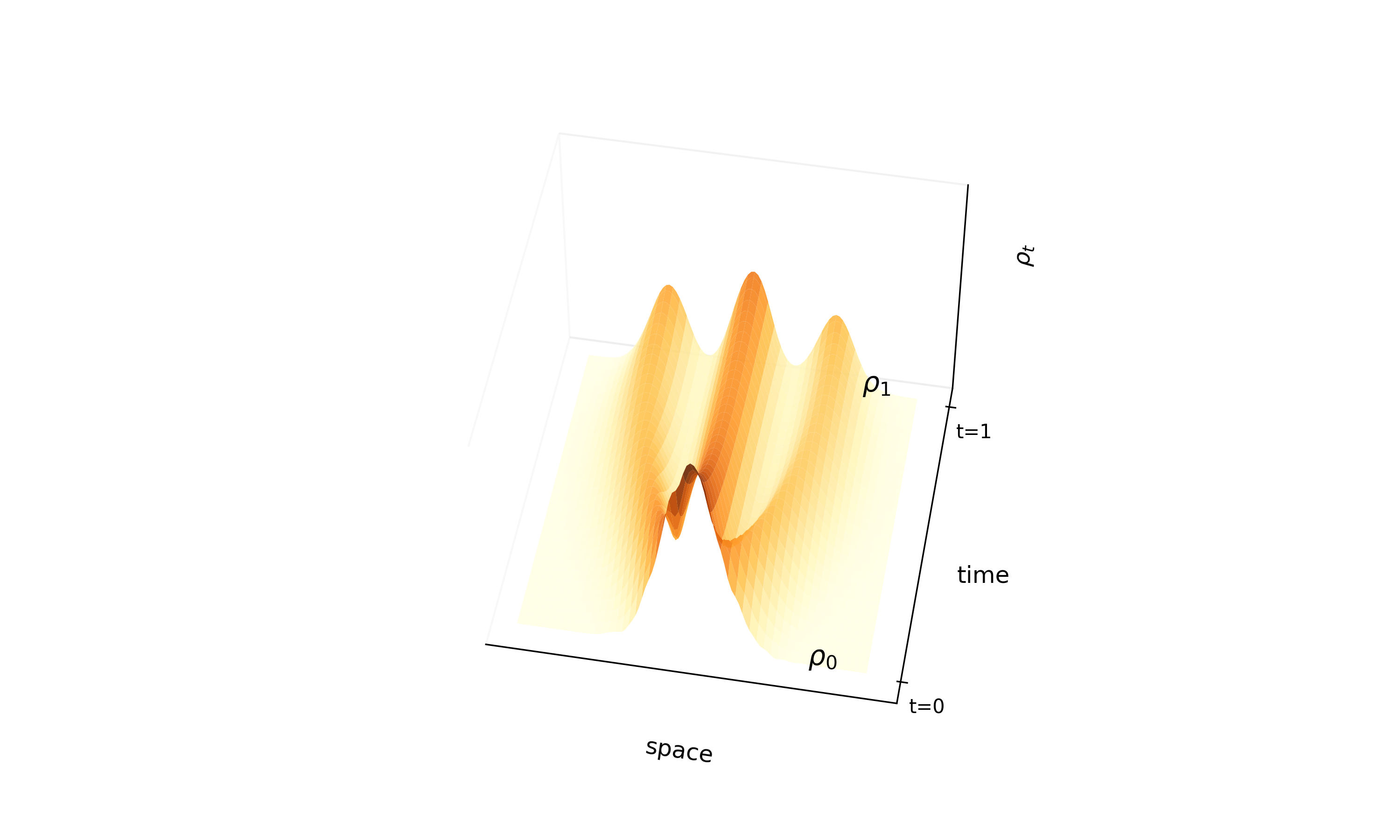
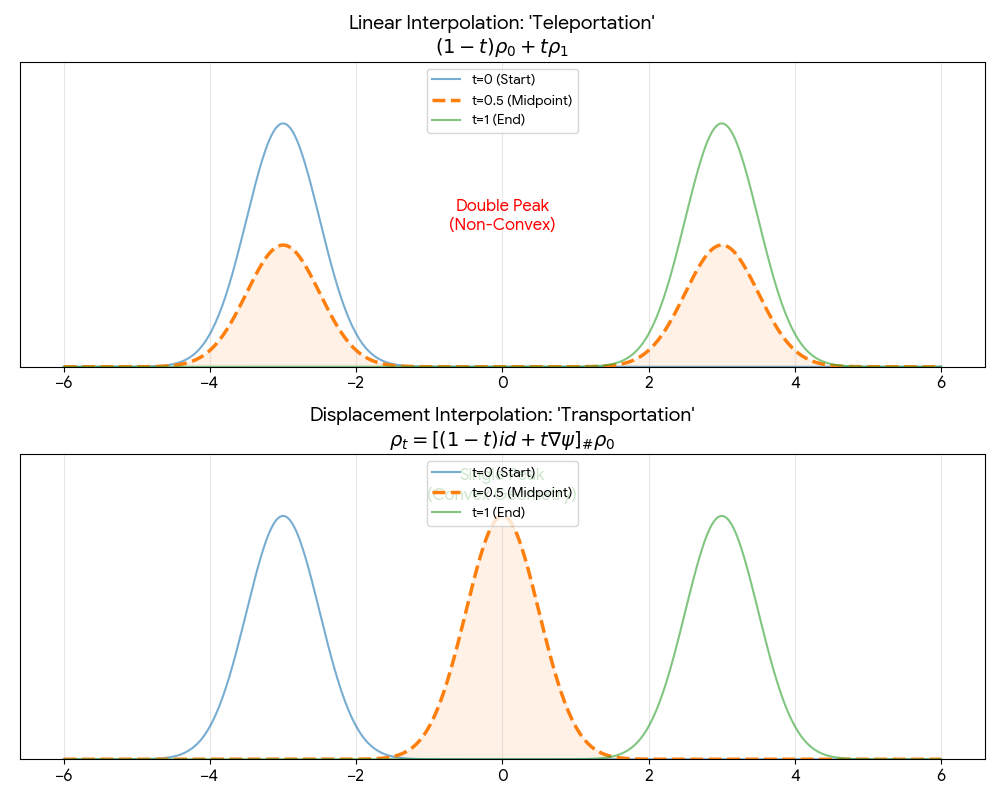
A theoretical paper that introduces displacement interpolation (optimal transport) to establish a new convexity principle for energy functionals. It proves the uniqueness of ground states for interacting gases and generalizes the Brunn-Minkowski inequality, providing mathematical tools later used in flow matching and optimal transport-based generative models.