Lennard-Jones on Adsorption and Diffusion on Surfaces
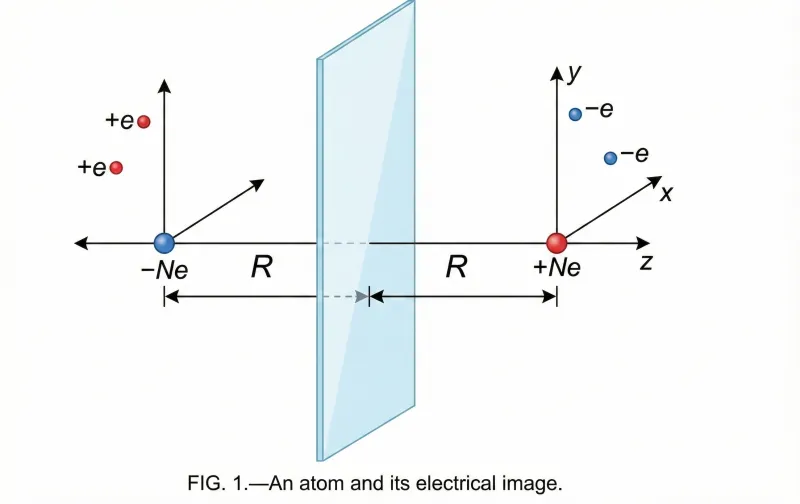
Lennard-Jones’s 1932 theoretical paper applying quantum mechanical potential energy surfaces to gas-solid interactions, providing the first unified framework explaining both physisorption and chemisorption as different regions of the same energy landscape.