Scaling Data-Constrained Language Models
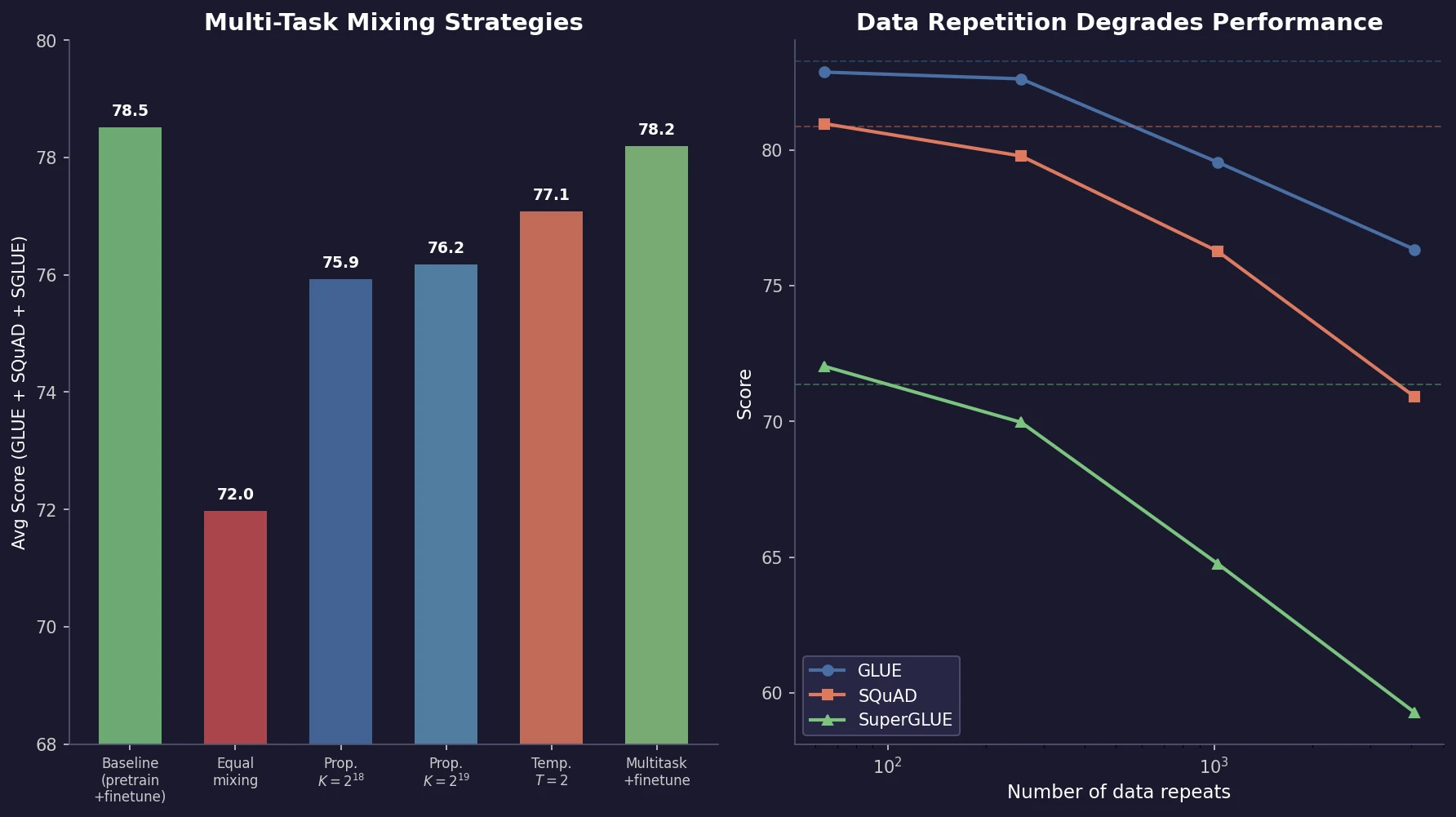
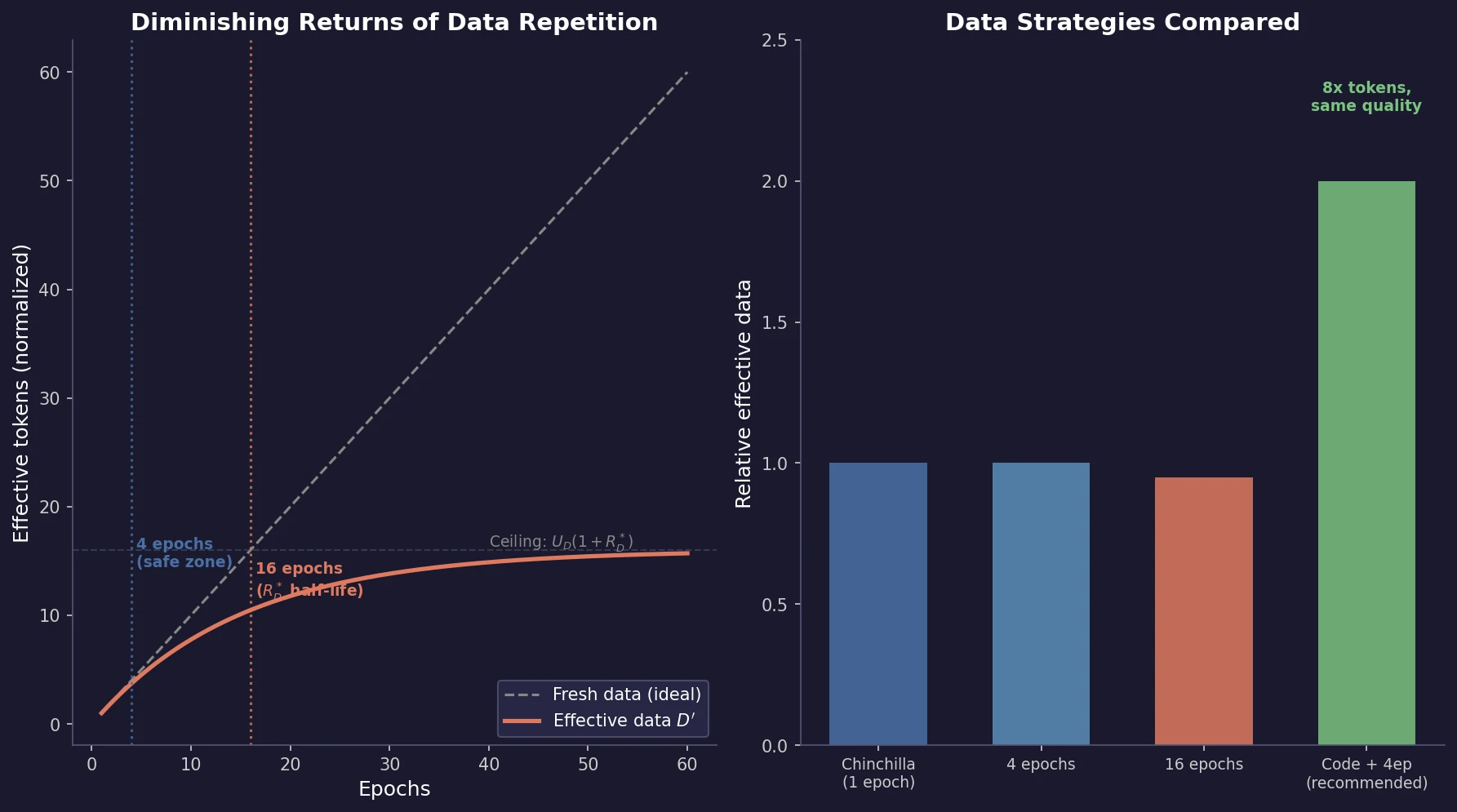
Muennighoff et al. train 400+ models to study how data repetition affects scaling. They propose a data-constrained scaling law with exponential decay for repeated tokens, finding that up to 4 epochs have negligible impact on loss, returns diminish around 16 epochs, and code augmentation provides a 2x effective data boost.