Consistency Models: Fast One-Step Diffusion Generation
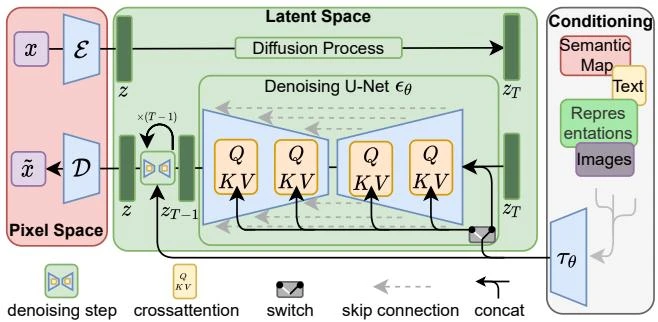
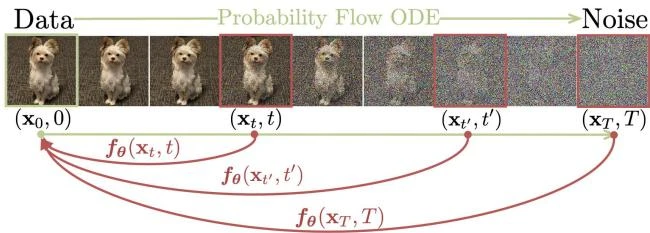
This paper introduces consistency models, a new family of generative models that map any point on a Probability Flow ODE trajectory to its origin. They support fast one-step generation by design, while allowing multi-step sampling for improved quality and zero-shot editing tasks like inpainting and colorization.