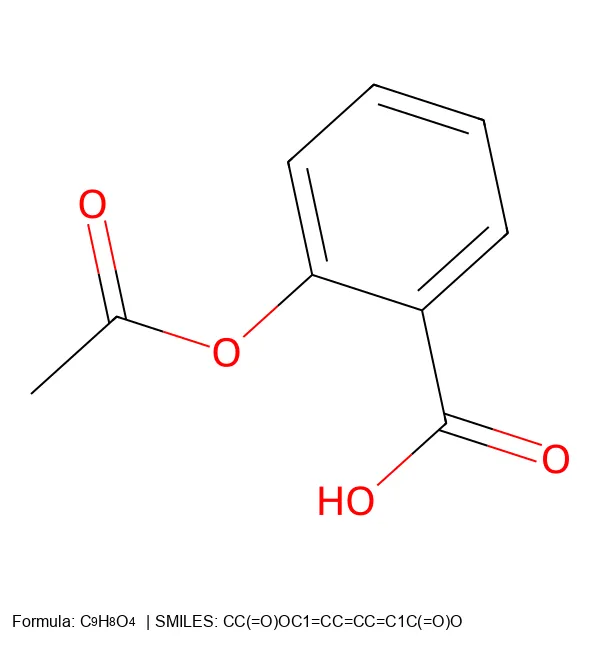
Converting SMILES and SELFIES to 2D Molecular Images
Build a robust Python CLI tool that converts both SMILES and SELFIES notation into publication-quality 2D molecular images, complete with formulas and legends.

Build a robust Python CLI tool that converts both SMILES and SELFIES notation into publication-quality 2D molecular images, complete with formulas and legends.
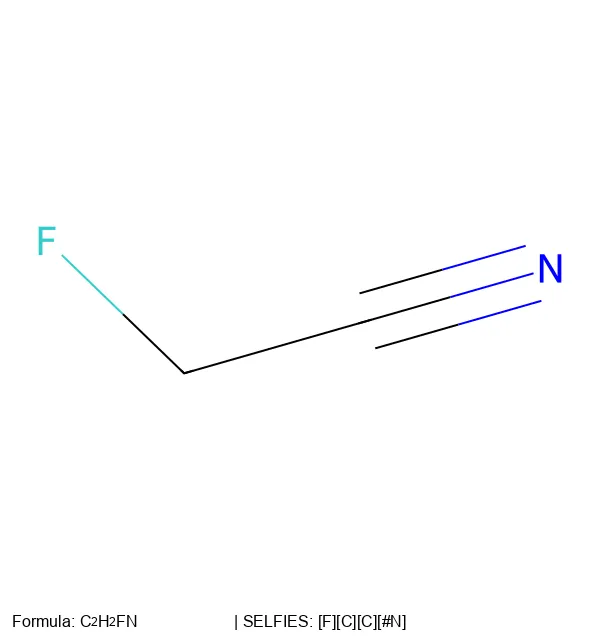
SELFIES is a molecular string representation where every possible string decodes to a valid molecule, solving the invalid-output problem that limits SMILES in generative machine learning.
Shannon’s foundational 1949 paper establishing the mathematical framework for modern information theory, defining channel capacity as the fundamental limit for reliable communication over noisy channels and introducing the sampling theorem (Nyquist-Shannon) that underpins all digital signal processing.
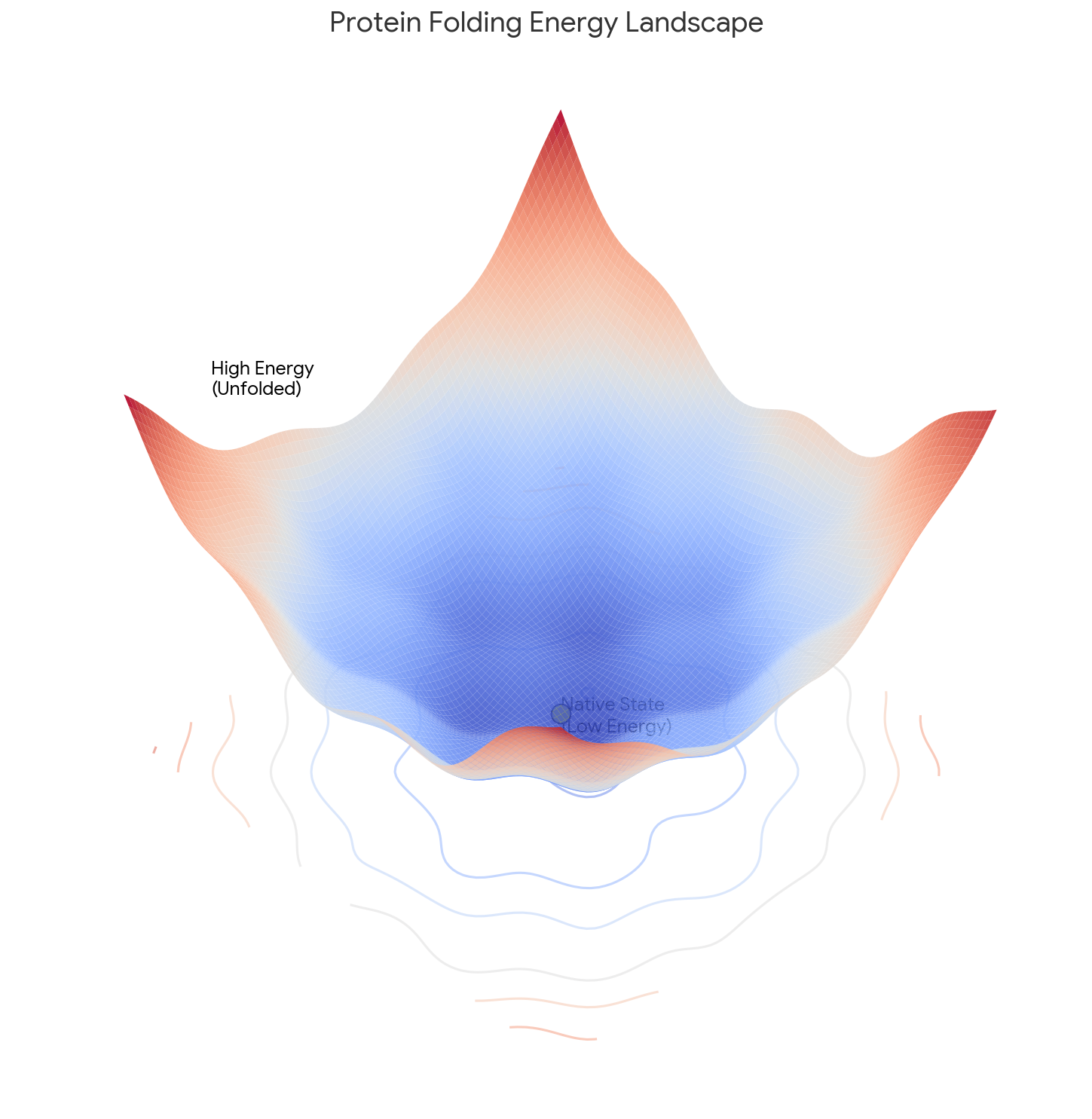
Levinthal’s 1969 perspective paper defined the protein folding paradox by demonstrating the impossibility of random search, establishing the need for kinetic pathways that guide folding faster than thermodynamic equilibration allows.
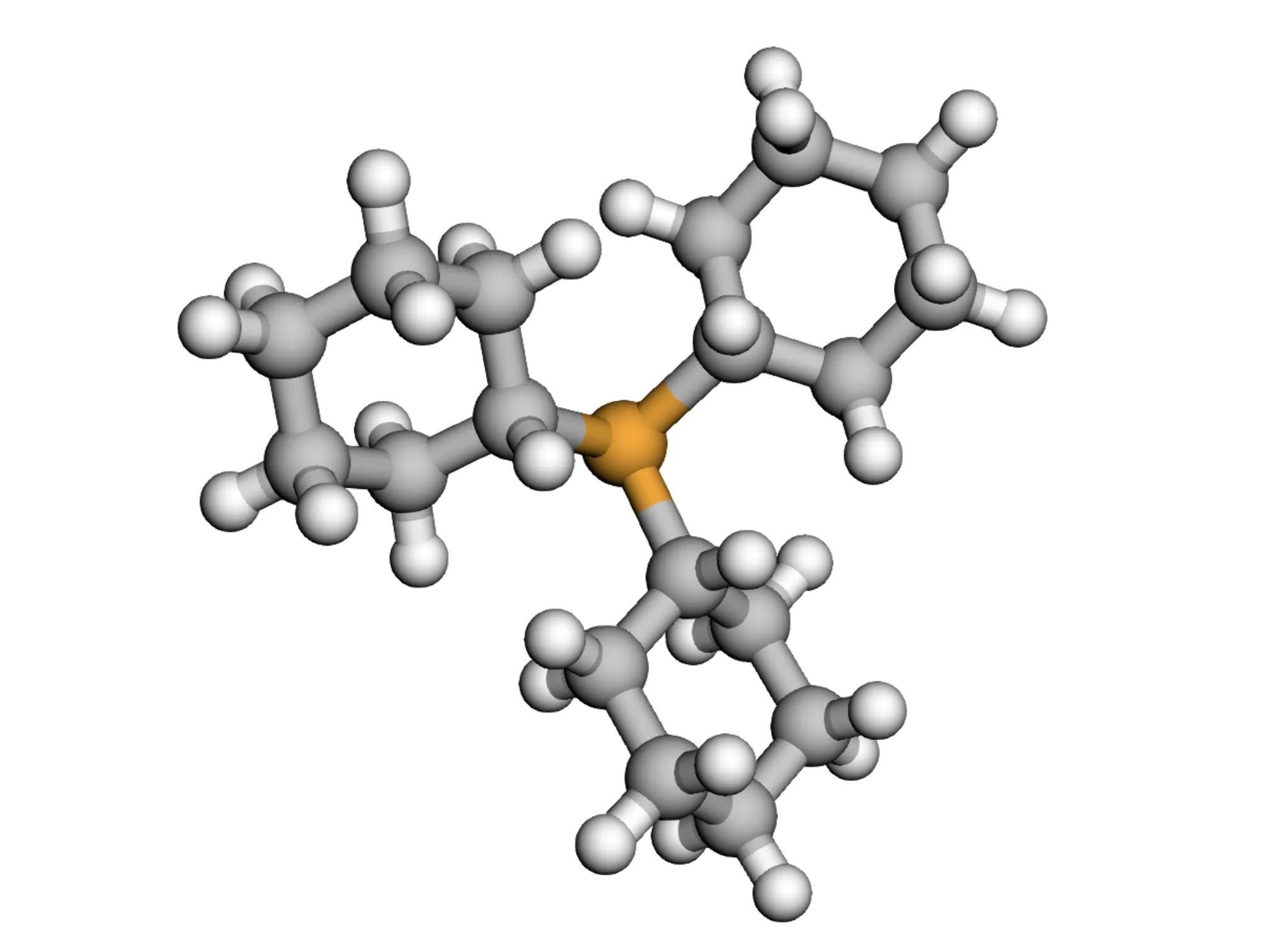
MARCEL provides a comprehensive benchmark for molecular representation learning with 722K+ conformers across four diverse subsets (Drugs-75K, Kraken, EE, BDE), enabling evaluation of conformer ensemble methods for property prediction in drug discovery and catalysis.
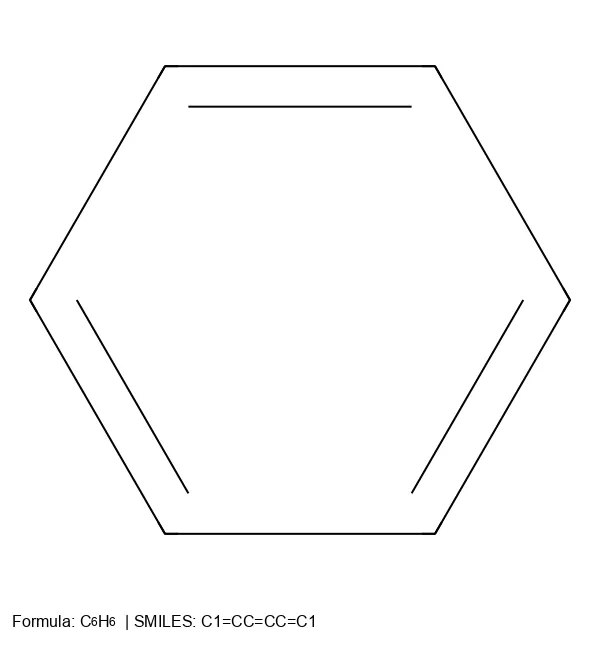
Comprehensive overview of SMILES notation for chemical structures, covering syntax for atoms, bonds, branches, rings, and stereochemistry, plus its key limitations for machine learning.
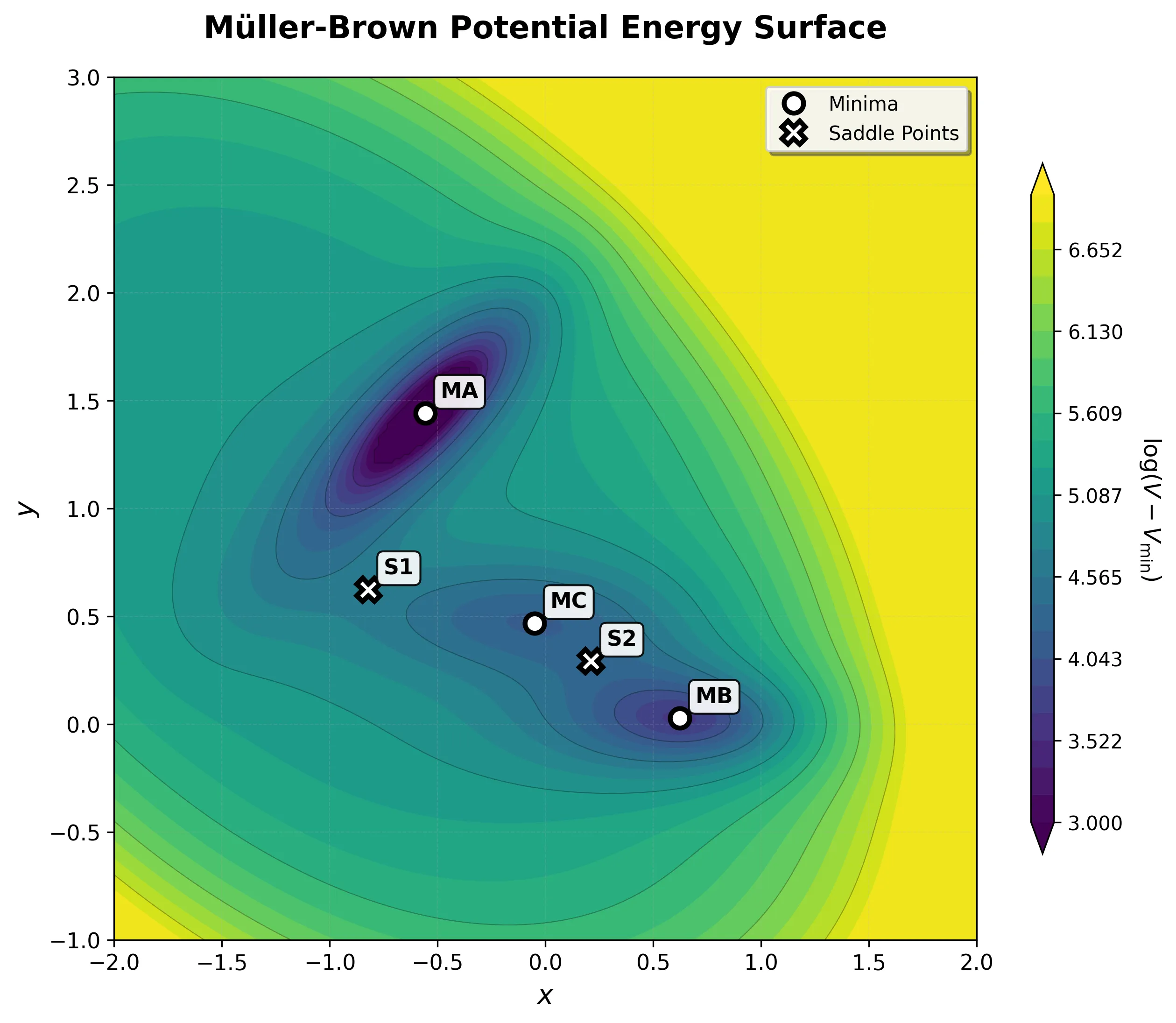
A two-dimensional analytical potential energy surface introduced in 1979 for testing optimization algorithms. It features three minima and curved transition pathways that evaluate an algorithm’s ability to navigate non-trivial topologies.
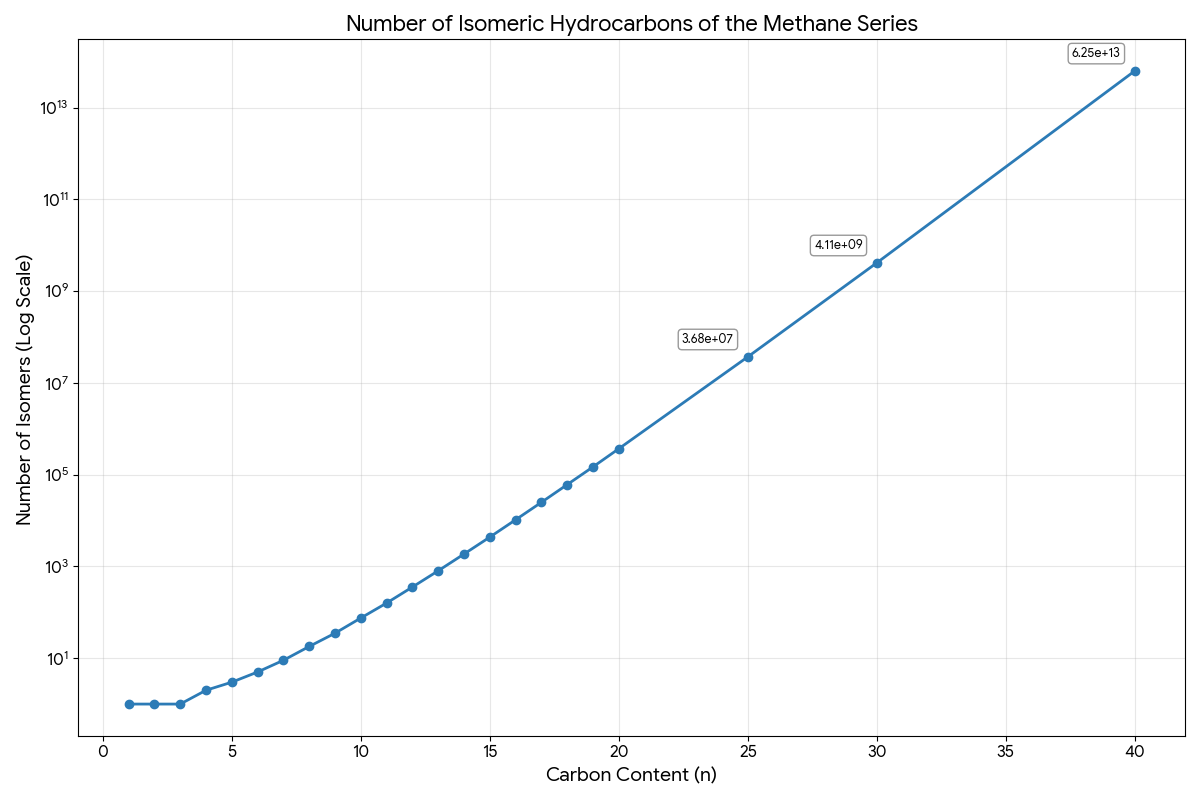
A foundational 1931 paper that derives exact recursive formulas for counting alkane structural isomers, correcting historical errors and establishing the first systematic enumeration up to C₄₀.
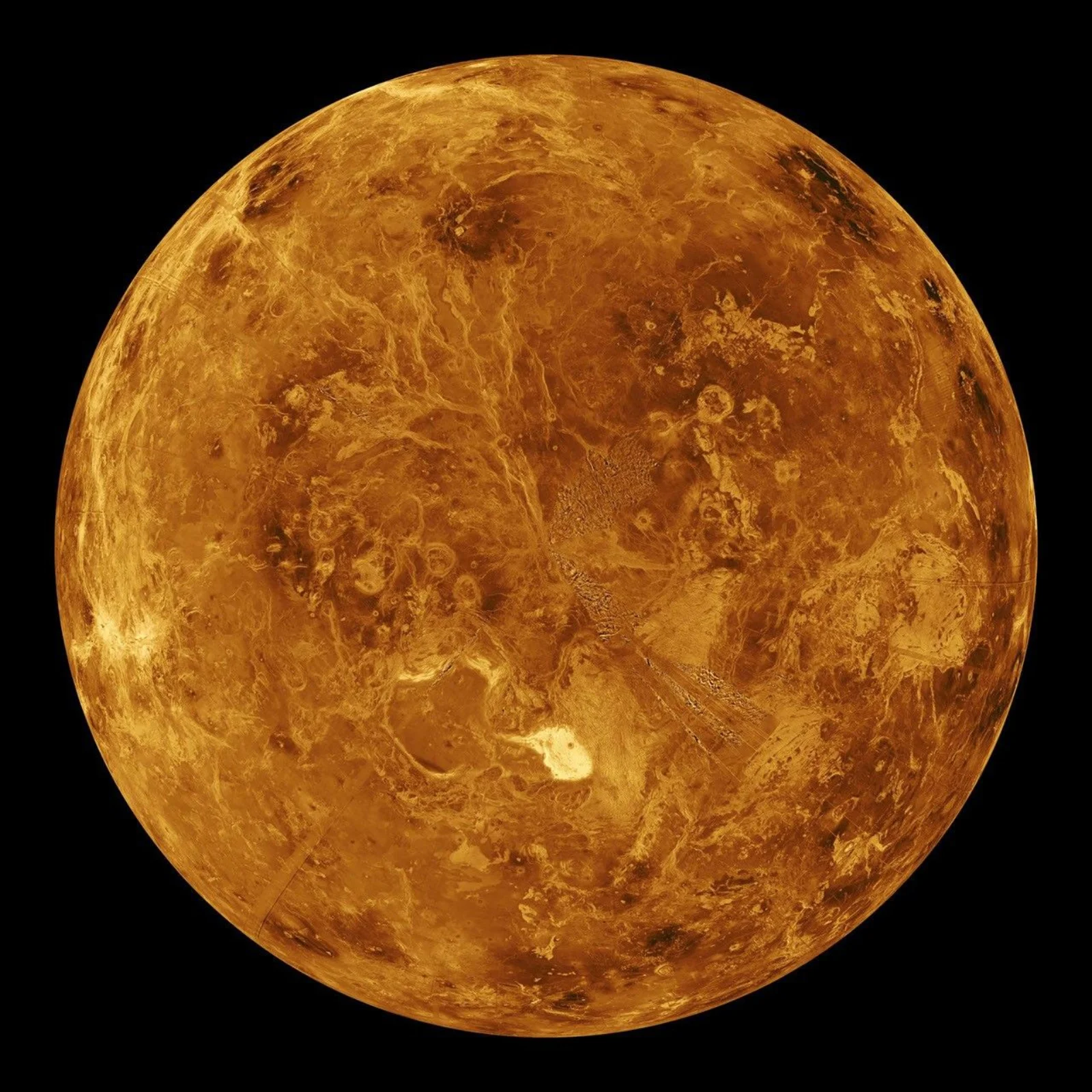

Basilevsky and Head’s comprehensive synthesis reveals a planet that undergoes catastrophic global resurfacing events. We explore the “stagnant lid” model, the synchronous stratigraphy, coronae, and the divergence of Venus’s geological history from Earth’s.
GEOM contains 450k+ molecules with 37M+ conformations, featuring energy annotations from semi-empirical (GFN2-xTB) and DFT methods for property prediction and molecular generation research.

Explore two fundamental approaches to generating exponentially distributed random numbers: the modern inverse transform method using logarithms and von Neumann’s ingenious 1951 comparison-based algorithm that avoids transcendental functions entirely.
GDB-11 contains 26.4 million systematically generated small organic molecules with up to 11 atoms, establishing the methodology for exploring drug-like chemical space computationally.