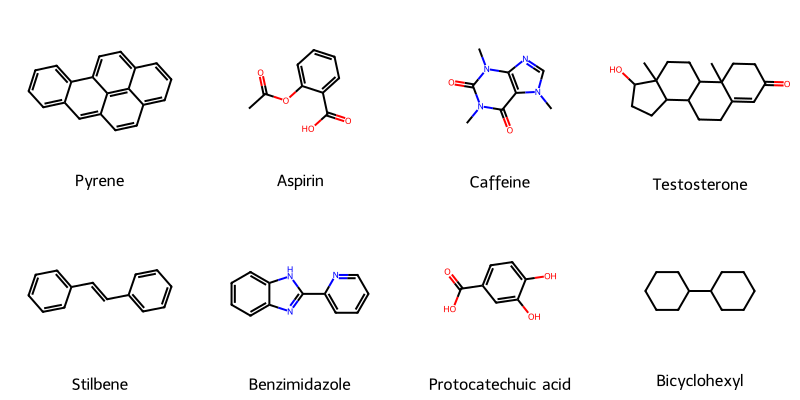
Invalid SMILES Benefit Chemical Language Models: A Study
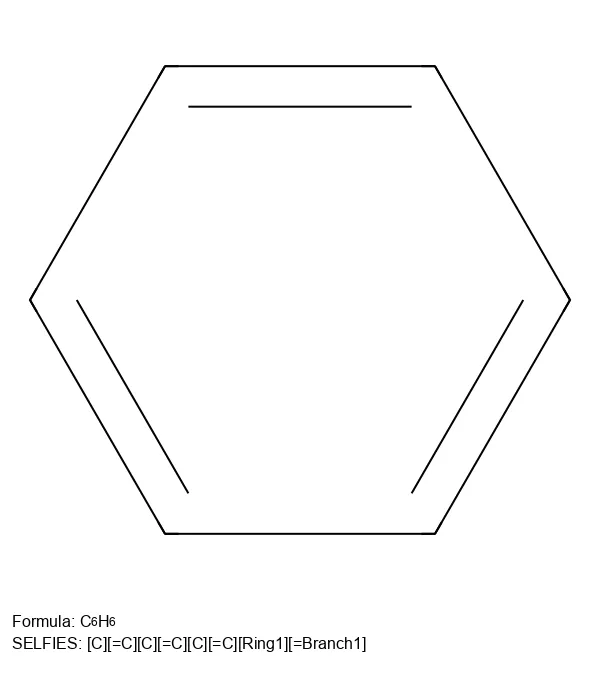
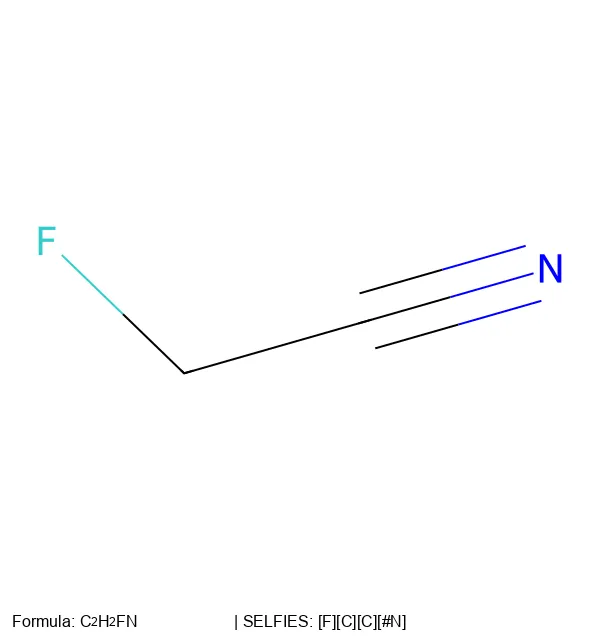
A 2024 Nature Machine Intelligence paper providing causal evidence that invalid SMILES generation improves chemical language model performance by filtering low-likelihood samples, while validity constraints (as in SELFIES) introduce structural biases that impair distribution learning.