PubMed-OCR: PMC Open Access OCR Annotations
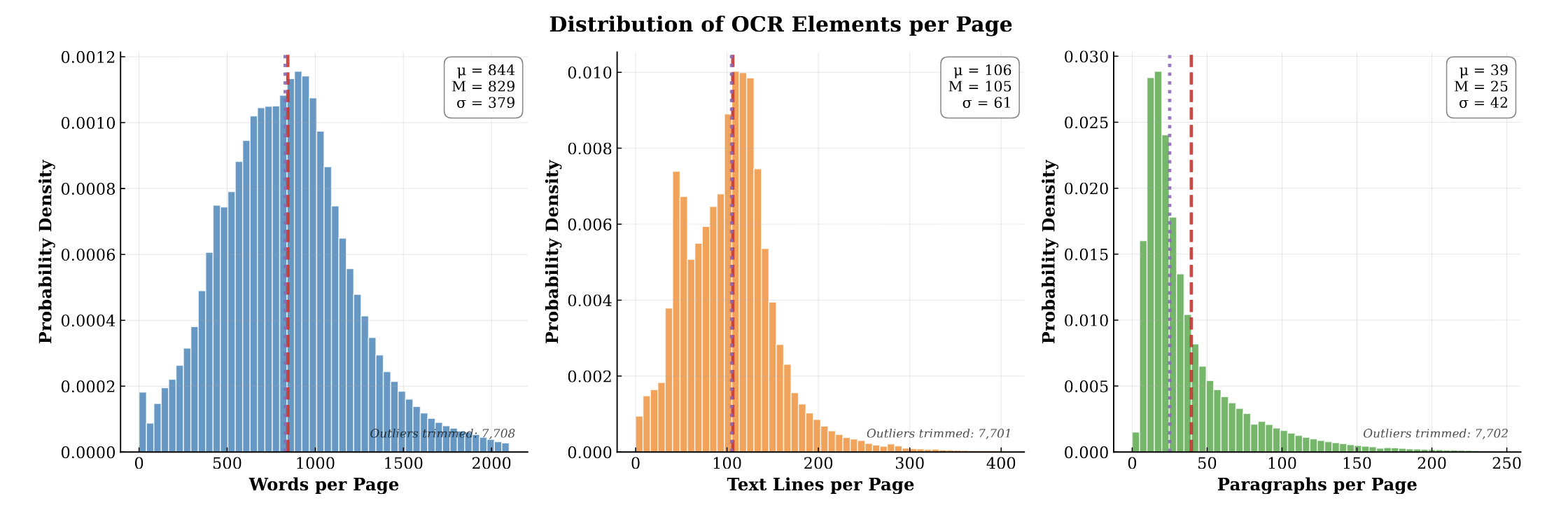
PubMed-OCR provides 1.5M pages of scientific articles with comprehensive OCR annotations and bounding boxes to support layout-aware modeling and document analysis.

PubMed-OCR provides 1.5M pages of scientific articles with comprehensive OCR annotations and bounding boxes to support layout-aware modeling and document analysis.

ChemBERTa-3 provides a unified, scalable infrastructure for pretraining and benchmarking chemical foundation models. It addresses reproducibility gaps in previous studies like MoLFormer through standardized scaffold splitting and open-source tooling.
ChemDFM-R is a 14B-parameter chemical reasoning model that integrates a 101B-token dataset of atomized chemical knowledge. Using a mix-sourced distillation strategy and domain-specific reinforcement learning, it outperforms similarly sized models and DeepSeek-R1 on ChemEval.

This work investigates the scaling hypothesis for molecular transformers, training RoBERTa models on 77M SMILES from PubChem. It compares Masked Language Modeling (MLM) against Multi-Task Regression (MTR) pretraining, finding that MTR yields better downstream performance but is computationally heavier.
This methodological paper proposes a linear-attention transformer decoder trained on 1.1 billion molecules. It introduces pair-tuning for efficient property optimization and establishes empirical scaling laws relating inference compute to generation novelty.

This paper introduces ChemBERTa, a RoBERTa-based model pretrained on 77M SMILES strings. It systematically evaluates the impact of pretraining dataset size, tokenization strategies, and input representations (SMILES vs. SELFIES) on downstream MoleculeNet tasks, finding that performance scales positively with data size.
This paper introduces Chemformer, a BART-based sequence-to-sequence model pre-trained on 100M molecules using a ‘combined’ masking and augmentation task. It achieves top-1 accuracy on reaction prediction benchmarks while significantly reducing training time through transfer learning.
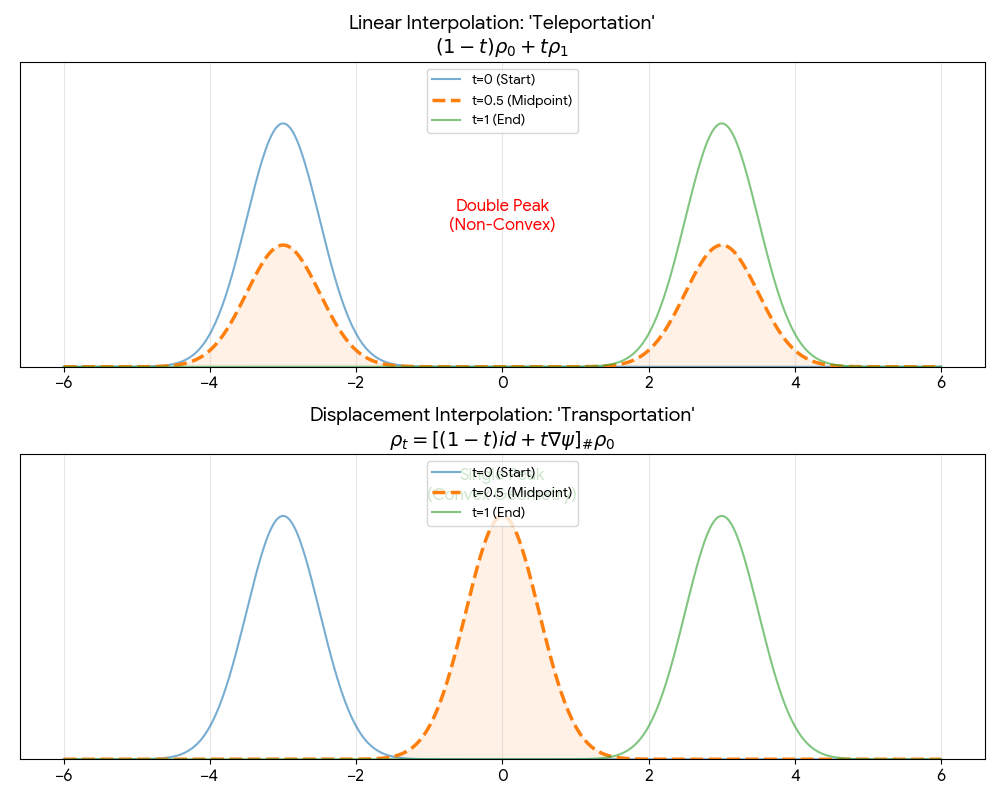
A theoretical paper that introduces displacement interpolation (optimal transport) to establish a new convexity principle for energy functionals. It proves the uniqueness of ground states for interacting gases and generalizes the Brunn-Minkowski inequality, providing mathematical tools later used in flow matching and optimal transport-based generative models.
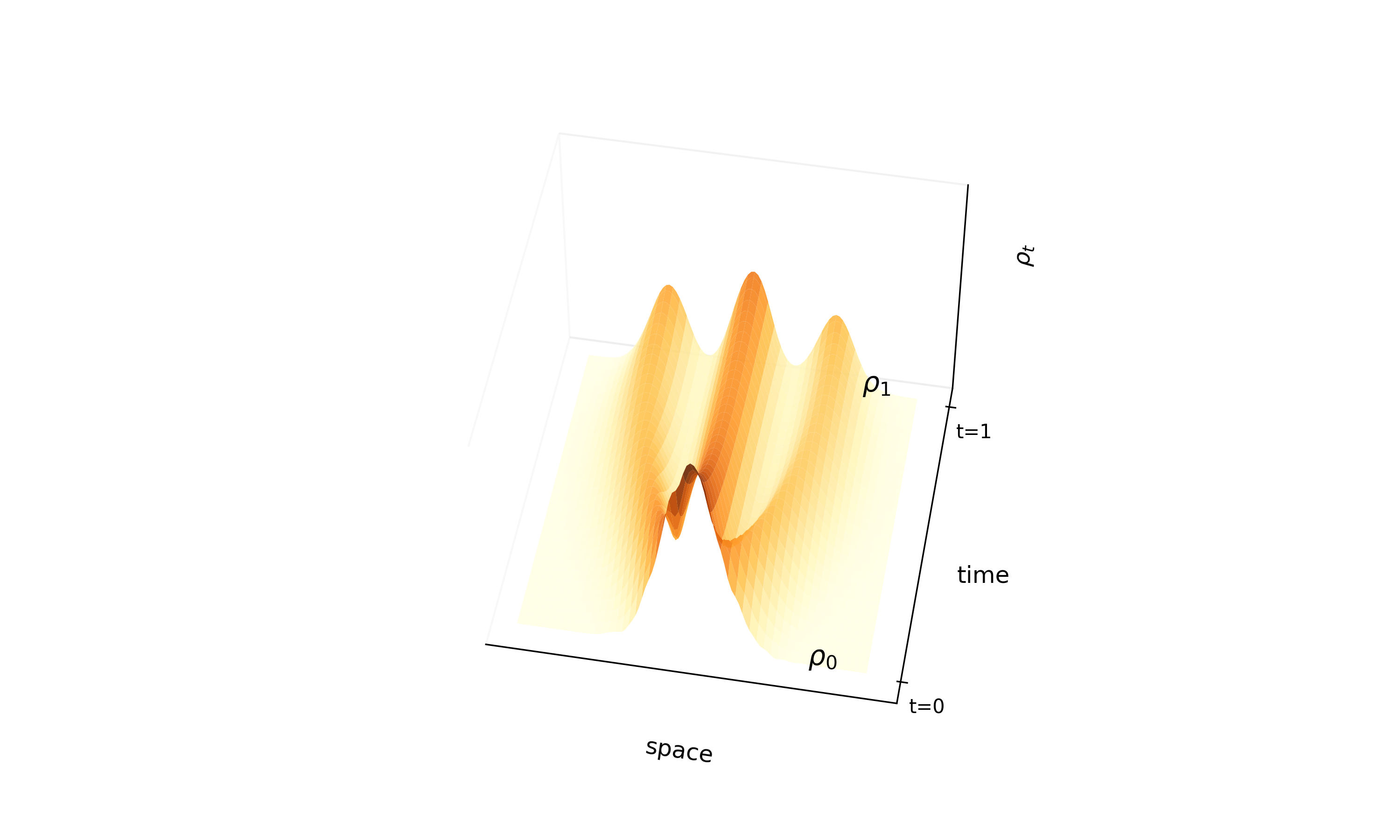
Proposes ‘InterFlow’, a method to learn continuous normalizing flows between arbitrary densities using stochastic interpolants. It avoids ODE backpropagation by minimizing a quadratic objective on the velocity field, enabling scalable ODE-based generation. On CIFAR-10, NLL matches ScoreSDE (2.99 bits per dim) with simulation-free training, though FID (10.27) trails dedicated image models (ScoreSDE: 2.92); the primary strength is tractable likelihood with efficient training cost.
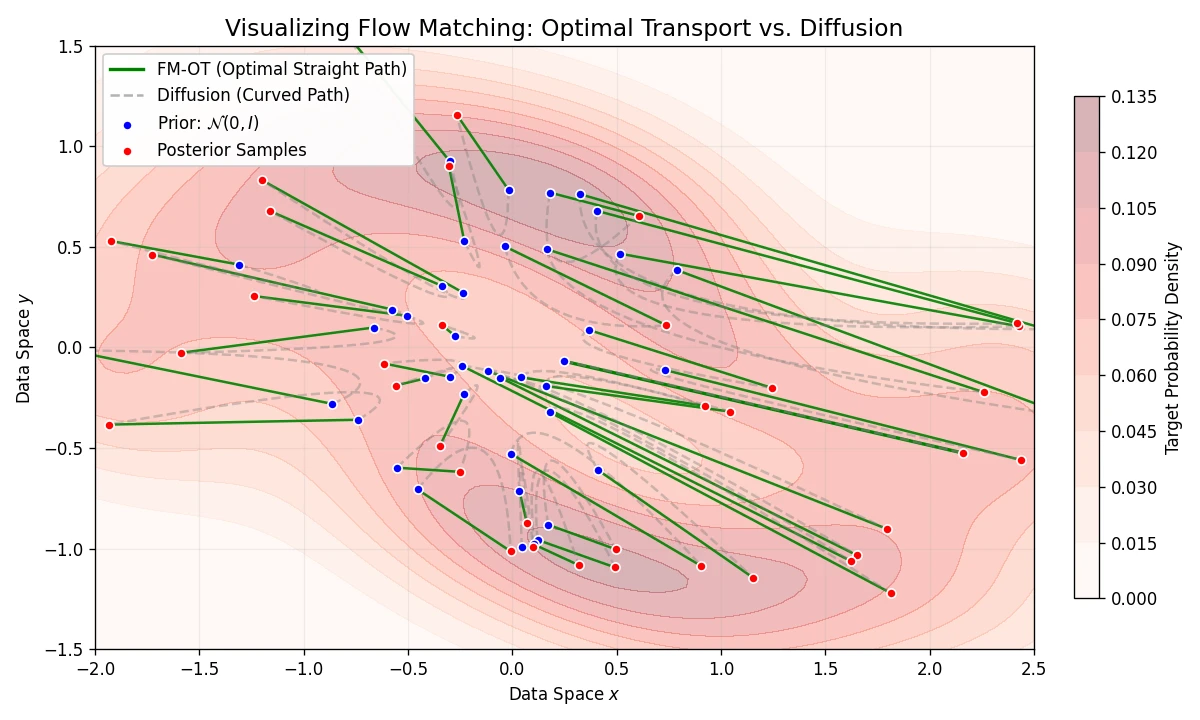
Introduces Flow Matching, a scalable method for training CNFs by regressing vector fields of conditional probability paths. It generalizes diffusion and enables Optimal Transport paths for straighter, more efficient sampling.
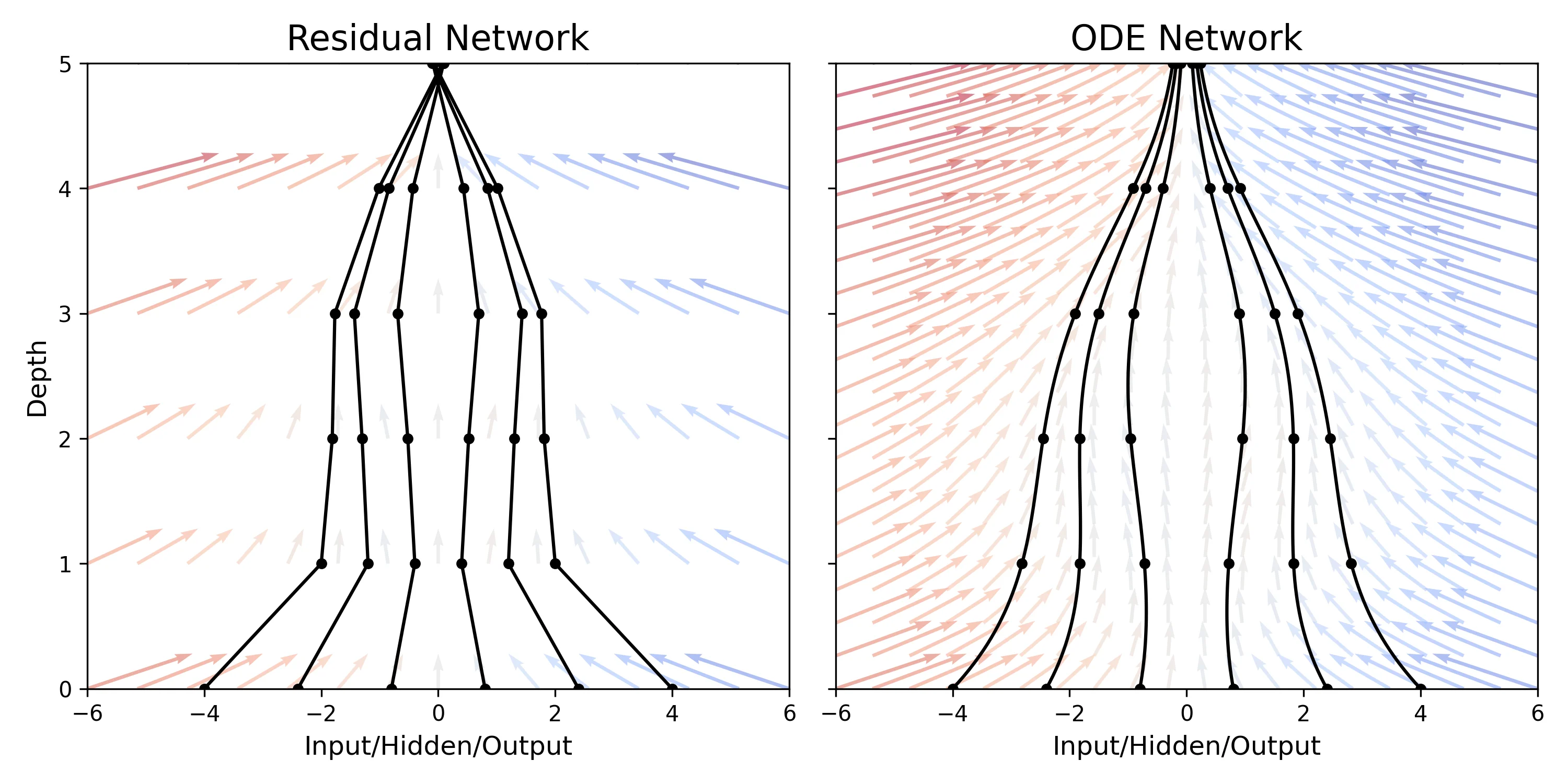
This paper replaces discrete network layers with continuous ordinary differential equations (ODEs), allowing for adaptive computation depth and constant memory cost during training via the adjoint sensitivity method. It introduces Continuous Normalizing Flows and latent ODEs for time-series.
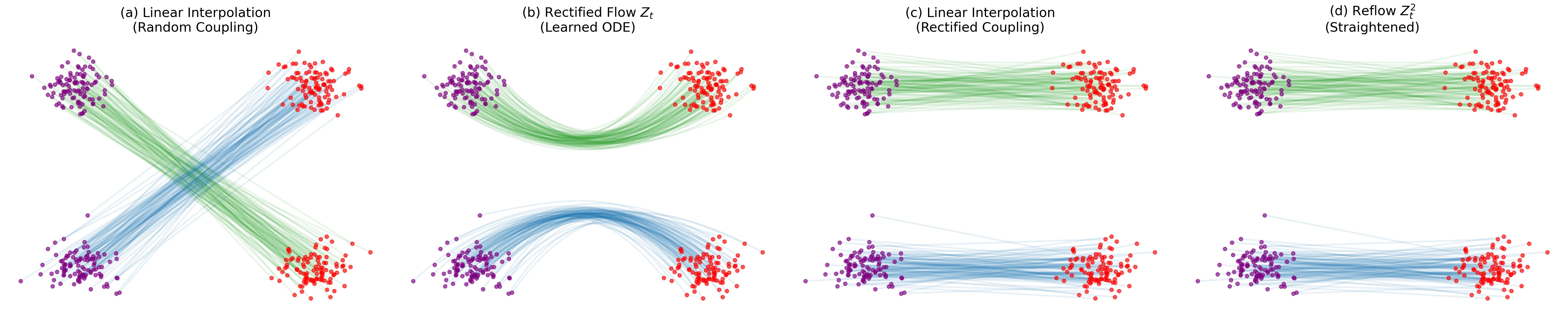
Introduces ‘Rectified Flow,’ a method to transport distributions via ODEs with straight paths. Uses a ‘reflow’ procedure to iteratively straighten trajectories, enabling high-quality 1-step generation with optional lightweight distillation.