A Method for Input-Adaptive Sequence Modeling
This is a Method paper that introduces Liquid-S4, a new state-space model combining the structured state-space framework (S4) with liquid time-constant (LTC) networks. The primary contribution is an input-dependent state transition mechanism that allows the model to adapt its dynamics based on incoming inputs, while retaining the efficient convolutional kernel computation of S4.
Scaling Liquid Networks to Long Sequences
Liquid time-constant (LTC) networks are continuous-time neural networks with input-dependent state transitions, giving them strong generalization and causal modeling properties. However, LTCs rely on ODE solvers that limit their scalability to long sequences. Structured state-space models (S4) solve this scalability problem through HiPPO initialization, diagonal plus low-rank (DPLR) parameterization, and efficient Cauchy kernel computation in the frequency domain, but they use fixed (input-independent) state transitions.
The key question this paper addresses: can the expressivity of LTC networks be combined with the efficiency and scalability of S4 to improve long-range sequence modeling?
The Liquid Kernel: Input-Dependent Convolutions
The core innovation is a linearized LTC state-space model that replaces the standard SSM dynamics:
$$\dot{x}(t) = \mathbf{A}x(t) + \mathbf{B}u(t)$$
with an input-dependent formulation:
$$\dot{x}(t) = \left[\mathbf{A} + \mathbf{B}u(t)\right]x(t) + \mathbf{B}u(t)$$
where $u(t)$ now modulates the state transition matrix itself. After discretization via the bilinear transform, the recurrence becomes:
$$x_{k} = \left(\overline{\mathbf{A}} + \overline{\mathbf{B}}u_{k}\right)x_{k-1} + \overline{\mathbf{B}}u_{k}$$
Unrolling this recurrence reveals that the output $y_{k}$ decomposes into two parts:
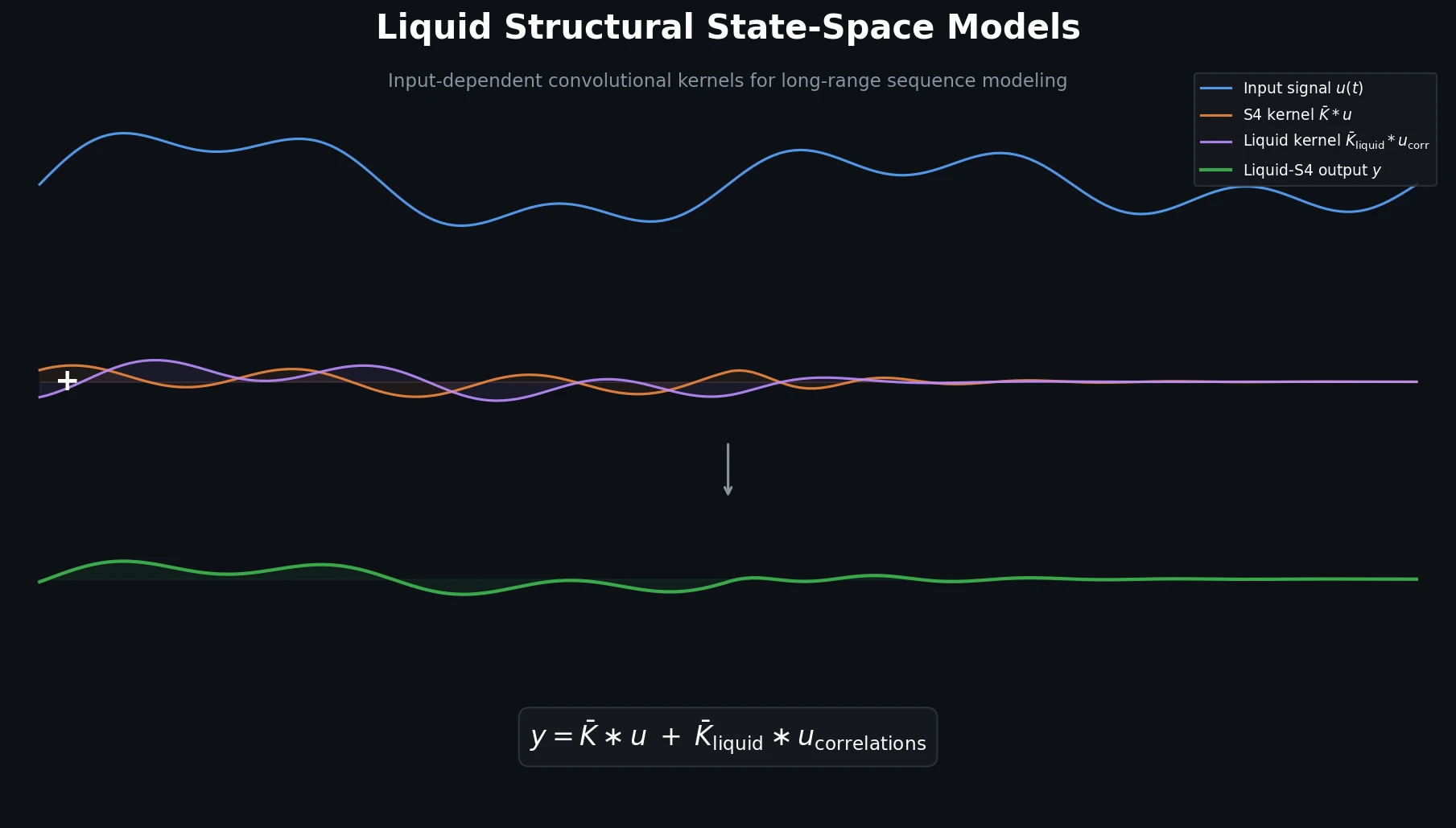
$$y = \overline{\mathbf{K}} * u + \overline{\mathbf{K}}_{\text{liquid}} * u_{\text{correlations}}$$
The first term is the standard S4 convolutional kernel $\overline{\mathbf{K}}$, mapping individual input time steps independently. The second term is a new “liquid kernel” $\overline{\mathbf{K}}_{\text{liquid}}$ that operates on auto-correlation terms of the input signal (products $u_{i}u_{j}$, $u_{i}u_{j}u_{k}$, etc., up to a chosen order $\mathcal{P}$).
Proposition 1 shows that each liquid kernel of order $p$ can be computed from the precomputed S4 kernel via a Hadamard product with $\overline{\mathbf{B}}^{p-1}$ followed by an anti-diagonal transformation (flip):
$$\overline{\mathbf{K}}_{\text{liquid}=p} = \left[\overline{\mathbf{K}}_{(L-\tilde{L},L)} \odot \overline{\mathbf{B}}_{(L-\tilde{L},L)}^{p-1}\right] * \mathbf{J}_{\tilde{L}}$$
This is the KB (Kernel $\times$ B) mode. The authors also propose a simplified PB (Powers of B) mode that sets the transition matrix $\overline{\mathbf{A}}$ to identity for the correlation terms:
$$\overline{\mathbf{K}}_{\text{liquid}=p} = \overline{\mathbf{C}} \odot \overline{\mathbf{B}}^{p-1}$$
The PB kernel is cheaper to compute and performs equally well or better in practice.
The computational complexity is $\tilde{\mathcal{O}}(N + L + p_{\text{max}}\tilde{L})$, where $N$ is the state size, $L$ the sequence length, $p_{\text{max}}$ the maximum liquid order, and $\tilde{L}$ the liquid kernel length (typically two orders of magnitude smaller than $L$).
Benchmarks Across Long-Range Sequence Tasks
Liquid-S4 is evaluated on four benchmark suites with the PB kernel using the S4-LegS (scaled Legendre) parameterization.
Long Range Arena (LRA)
The LRA benchmark contains six tasks with sequence lengths from 1K to 16K. Liquid-S4 achieves state-of-the-art on all six tasks with an average accuracy of 87.32%:
| Task | Input Length | Liquid-S4 | S4-LegS | Improvement |
|---|---|---|---|---|
| ListOps | 2048 | 62.75% | 59.60% | +3.15% |
| Text (IMDB) | 2048 | 89.02% | 86.82% | +2.20% |
| Retrieval (AAN) | 4000 | 91.20% | 90.90% | +0.30% |
| Image (CIFAR) | 1024 | 89.50% | 88.65% | +0.85% |
| Pathfinder | 1024 | 94.80% | 94.20% | +0.60% |
| Path-X | 16384 | 96.66% | 96.35% | +0.31% |
| Average | 87.32% | 86.09% | +1.23% |
Liquid orders $p$ range from 2 to 6 across tasks.
BIDMC Vital Signs
On medical time-series regression (heart rate, respiratory rate, SpO2 prediction from length-4000 biomarker signals):
| Task | Liquid-S4 (RMSE) | S4-LegS (RMSE) | Improvement |
|---|---|---|---|
| Heart Rate | 0.303 | 0.332 | 8.7% |
| Respiratory Rate | 0.158 | 0.247 | 36.0% |
| SpO2 | 0.066 | 0.090 | 26.7% |
Sequential CIFAR (sCIFAR)
Liquid-S4 with $p=3$ achieves 92.02% accuracy on 1-D pixel-level image classification, improving over S4-LegS (91.80%).
Speech Commands (Full 35 Labels)
On the raw 16kHz speech recognition task, Liquid-S4 achieves 96.78% accuracy with only 224K parameters, a 30% reduction compared to S4’s 307K. On the zero-shot 8kHz experiment, performance drops to 90.00% (vs. 91.32% for S4-LegS), which the authors attribute to the liquid kernel’s sensitivity to input covariance structure at different sampling rates.
Consistent Improvements with Smaller Models
Liquid-S4 achieves state-of-the-art performance on every benchmark evaluated: all six LRA tasks (87.32% average), all three BIDMC vital signs tasks, sCIFAR, and full Speech Commands recognition. The gains are particularly large on tasks where input correlation structure matters (ListOps +3.15%, IMDB +2.20%, respiratory rate RMSE improvement of 36%).
A practical advantage is that Liquid-S4 works well with smaller state sizes (as low as 7 units for some tasks), reducing parameter counts. The PB kernel is recommended over KB for its simplicity and competitive performance. Higher liquid orders ($p$) consistently improve performance, though $p=3$ is recommended as a default.
Limitations include degraded performance in zero-shot frequency transfer (8kHz Speech Commands), suggesting the liquid kernel’s input covariance terms may not generalize well across sampling rate changes. The paper also does not compare against non-SSM approaches beyond the LRA benchmark. The causal (unidirectional) configuration works better than bidirectional for Liquid-S4, which may limit applicability to tasks that benefit from bidirectional context.
Reproducibility Details
Classification: Partially Reproducible. Code and all benchmark datasets are publicly available, with complete hyperparameters documented. No pre-trained weights are released and hardware requirements are not specified.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| raminmh/liquid-s4 | Code | Apache-2.0 | Official PyTorch implementation; fork of the S4 repo with KB and PB kernels added |
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Evaluation | Long Range Arena (LRA) | 6 tasks, 1K-16K seq length | ListOps, IMDB, AAN, CIFAR, Pathfinder, Path-X |
| Evaluation | BIDMC Vital Signs | 4000-length biomarker signals | Heart rate, respiratory rate, SpO2 |
| Evaluation | sCIFAR | 1024-length flattened images | 10-class classification |
| Evaluation | Speech Commands | 16kHz raw audio, 35 labels | Full dataset with zero-shot 8kHz test |
Algorithms
The Liquid-S4 kernel computation builds on the S4 kernel pipeline:
- Initialize $\mathbf{A}$ with HiPPO (scaled Legendre) matrix in DPLR form
- Compute S4 kernel $\overline{\mathbf{K}}$ via Cauchy kernel and iFFT
- For each liquid order $p \in {2, \ldots, \mathcal{P}}$, compute $\overline{\mathbf{K}}_{\text{liquid}=p}$ using either KB or PB mode
- Convolve $\overline{\mathbf{K}}_{\text{liquid}}$ with input correlation vector $u_{\text{correlations}}$
The PB kernel mode is used in all reported experiments. The PyKeops package is used for large tensor computations.
Models
| Task | Depth | Features | State Size | Norm | LR | Epochs |
|---|---|---|---|---|---|---|
| ListOps | 9 | 128 | 7 | BN | 0.002 | 30 |
| IMDB | 4 | 128 | 7 | BN | 0.003 | 50 |
| AAN | 6 | 256 | 64 | BN | 0.005 | 20 |
| CIFAR (LRA) | 6 | 512 | 512 | LN | 0.01 | 200 |
| Pathfinder | 6 | 256 | 64 | BN | 0.0004 | 200 |
| Path-X | 6 | 320 | 64 | BN | 0.001 | 60 |
| Speech Commands | 6 | 128 | 7 | BN | 0.008 | 50 |
| BIDMC (HR) | 6 | 128 | 256 | LN | 0.005 | 500 |
| BIDMC (RR) | 6 | 128 | 256 | LN | 0.01 | 500 |
| BIDMC (SpO2) | 6 | 128 | 256 | LN | 0.01 | 500 |
| sCIFAR | 6 | 512 | 512 | LN | 0.01 | 200 |
Liquid-S4 generally requires smaller learning rates than S4/S4D. $\Delta t_{\text{max}} = 0.2$ for all experiments; $\Delta t_{\text{min}} \propto 1/\text{seq_length}$.
Evaluation
All results report validation accuracy (except BIDMC, which reports test RMSE). Experiments use 2-3 random seeds with standard deviations reported.
Hardware
Not specified in the paper.
Paper Information
Citation: Hasani, R., Lechner, M., Wang, T.-H., Chahine, M., Amini, A., & Rus, D. (2022). Liquid Structural State-Space Models. arXiv preprint arXiv:2209.12951.
@misc{hasani2022liquid,
title={Liquid Structural State-Space Models},
author={Hasani, Ramin and Lechner, Mathias and Wang, Tsun-Hsuan and Chahine, Makram and Amini, Alexander and Rus, Daniela},
year={2022},
eprint={2209.12951},
archiveprefix={arXiv},
primaryclass={cs.LG}
}