What kind of paper is this?
This is a method paper that introduces a novel neural network architecture, the 3D Steerable CNN. It provides a comprehensive theoretical derivation for the architecture grounded in group representation theory and demonstrates its practical application.
What is the motivation?
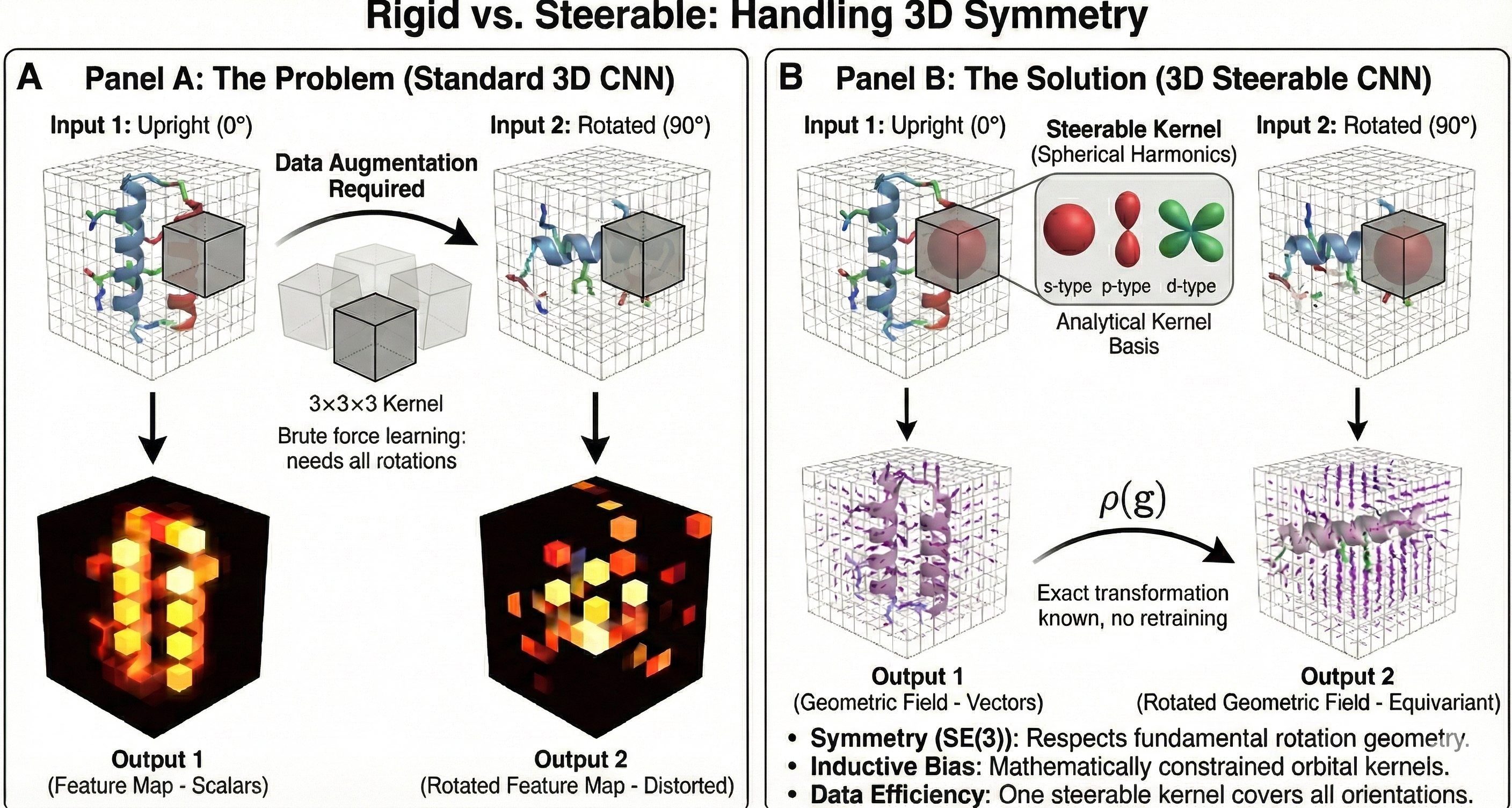
The work is motivated by the prevalence of symmetry in problems from the natural sciences. Standard 3D CNNs lack inherent equivariance to 3D rotations, a fundamental symmetry governed by the SE(3) group in many scientific datasets like molecular or protein structures. Building this symmetry directly into the model architecture as an inductive bias is expected to yield more data-efficient, generalizable, and physically meaningful models.
What is the novelty here?
The core novelty is the rigorous and practical construction of a CNN architecture that is equivariant to 3D rigid body motions (SE(3) group). The key contributions are:
- Geometric Feature Representation: Features are modeled as geometric fields (collections of scalars, vectors, and higher-order tensors) defined over $\mathbb{R}^{3}$. Each type of feature transforms according to an irreducible representation (irrep) of the rotation group SO(3).
- General Equivariant Convolution: The paper proves that the most general form of an SE(3)-equivariant linear map between these fields is a convolution with a rotation-steerable kernel.
- Analytical Kernel Basis: The main theoretical breakthrough is the analytical derivation of a complete basis for these steerable kernels. They solve the kernel’s equivariance constraint, $\kappa(rx) = D^{j}(r)\kappa(x)D^{l}(r)^{-1}$, showing the solutions are functions whose angular components are spherical harmonics. The network’s kernels are then parameterized as a learnable linear combination of these pre-computed basis functions, making the implementation a minor modification to standard 3D convolutions.
- Equivariant Nonlinearity: A novel gated nonlinearity is proposed for non-scalar features. It preserves equivariance by multiplying a feature field by a separately computed, learned scalar field (the gate).
What experiments were performed?
The model’s performance was evaluated on a series of tasks with inherent rotational symmetry:
- Tetris Classification: A toy problem to empirically validate the model’s rotational equivariance by training on aligned blocks and testing on randomly rotated ones.
- SHREC17 3D Model Classification: A benchmark for classifying complex 3D shapes that are arbitrarily rotated.
- Amino Acid Propensity Prediction: A scientific application to predict amino acid types from their 3D atomic environments.
- CATH Protein Structure Classification: A challenging task on a new dataset introduced by the authors, requiring classification of global protein architecture, a problem with full SE(3) invariance.
What outcomes/conclusions?
The 3D Steerable CNN demonstrated clear advantages due to its built-in equivariance:
- It was empirically confirmed to be rotationally equivariant, achieving $99 \pm 2%$ test accuracy on the rotated Tetris dataset (averaged over 17 runs), compared to a standard 3D CNN’s $27 \pm 7%$ accuracy.
- On the amino acid prediction task the model achieves 0.58 accuracy, compared to 0.50 (regular-grid) and 0.56 (concentric-grid) baselines, using roughly half the parameters. On SHREC17 it reaches a total score (micro + macro mAP) of 1.11, compared to 1.13 for the leading contemporary system.
- On the CATH protein classification task, it outperformed a deep 3D CNN baseline while using ~110x fewer parameters. This performance gap widened as the training data was reduced, highlighting the model’s superior data efficiency.
The paper concludes that 3D Steerable CNNs provide a universal and effective framework for incorporating SE(3) symmetry into deep learning models, leading to improved accuracy and efficiency for tasks involving volumetric data, particularly in scientific domains.
Reproducibility Details
Data
- Input Format: All inputs must be voxelized. Point clouds require voxelization before use.
- Proteins (CATH): $50^3$ grid, 0.2 nm voxel size. Simplified to $C_\alpha$ atoms only; Gaussian density placed at each atom position.
- 3D Objects (SHREC17): $64^3$ voxel grids.
- Tetris: $36^3$ input grid.
- Splitting Strategy: CATH used a 10-fold split (7 train, 1 val, 2 test) strictly separated by “superfamily” level to prevent data leakage (<40% sequence identity).
Models
Kernel Basis Construction:
- Constructed from Spherical Harmonics multiplied by Gaussian Radial Shells: $\exp\left(-\frac{1}{2}(|x|-m)^{2}/\sigma^{2}\right)$
- Anti-aliasing: A radially dependent angular frequency cutoff ($J_{\max}$) is applied to prevent aliasing near the origin.
Normalization: Uses Equivariant Batch Norm. Non-scalar fields are normalized by the average of their norms.
Downsampling: Standard strided convolution breaks equivariance. The architecture uses low-pass filtering (Gaussian blur) before downsampling to maintain equivariance.
Exact Architecture Configurations:
Tetris Architecture (4 layers):
| Layer | Field Types | Spatial Size |
|---|---|---|
| Input | 1 scalar | $36^3$ |
| Layer 1 | 4 scalars, 4 vectors ($l=1$), 4 tensors ($l=2$), 1 tensor ($l=3$) | $40^3$ |
| Layer 2 | 16 scalars, 16 vectors, 16 tensors ($l=2$) | $22^3$ (stride 2) |
| Layer 3 | 32 scalars, 16 vectors, 16 tensors ($l=2$) | $13^3$ (stride 2) |
| Layer 4 | 128 scalars | $17^3$ |
| Output | 8 scalars (global average pool) | $1$ |
SHREC17 Architecture (8 layers):
| Layers | Field Types |
|---|---|
| 1-2 | 8 scalars, 4 vectors, 2 tensors ($l=2$) |
| 3-4 | 16 scalars, 8 vectors, 4 tensors |
| 5-7 | 32 scalars, 16 vectors, 8 tensors |
| 8 | 512 scalars |
| Output | 55 scalars (classes) |
CATH Architecture (ResNet34-inspired):
Block progression: (2,2,2,2), (4,4,4,4), (8,8,8,8), (16,16,16,16)
Notation: (a,b,c,d) = $a$ scalars ($l=0$), $b$ vectors ($l=1$), $c$ rank-2 tensors ($l=2$), $d$ rank-3 tensors ($l=3$).
Algorithms
Parameter Counts:
| Task | Model | Parameters |
|---|---|---|
| CATH | 3D Steerable CNN | 143,560 |
| CATH | Baseline (ResNet34-style) | 15,878,764 |
| Amino Acid | 3D Steerable CNN | ~32,600,000 |
| Amino Acid | Regular grid baseline | ~61,100,000 |
| Amino Acid | Concentric grid baseline | ~75,300,000 |
Note: The concentric grid baseline groups voxels by distance from the molecular center, reflecting that atomic interactions are primarily distance-dependent (Torng, W., & Altman, R. B. (2017). 3D deep convolutional neural networks for amino acid environment similarity analysis. BMC Bioinformatics, 18, 302). Amino acid parameter counts are rounded figures as reported in the paper.
Hyperparameters & Training:
| Parameter | Value |
|---|---|
| Optimizer | Adam |
| LR Scheduler | Exponential decay (0.94/epoch) after 40 epoch burn-in |
| Dropout (CATH) | 0.1 (Capsule-wide convolutional dropout) |
| Weight Decay (CATH) | L1 & L2 regularization: $10^{-8.5}$ |
Mathematical Formulations for Equivariance:
Standard operations like Batch Normalization and ReLU break rotational equivariance. The paper derives equivariant alternatives:
Equivariant Batch Normalization:
Standard BN subtracts a mean, which introduces a preferred direction and breaks symmetry. Norm-based normalization scales feature fields by the average of their squared norms to preserve symmetry:
$$f_{i}(x) \mapsto f_{i}(x) \left( \frac{1}{|B|} \sum_{j \in B} \frac{1}{V} \int dx |f_{j}(x)|^{2} + \epsilon \right)^{-1/2}$$
This scales vector lengths to unit variance on average while avoiding mean subtraction, preserving directional information and symmetry.
Equivariant Nonlinearities:
Applying ReLU to vector components independently breaks equivariance (this depends on the coordinate frame). Two approaches:
Norm Nonlinearity (geometric shrinking): Acts on magnitude, preserves direction. Shrinks vectors shorter than learned bias $\beta$ to zero: $$f(x) \mapsto \text{ReLU}(|f(x)| - \beta) \frac{f(x)}{|f(x)|}$$ Note: Found to converge slowly; omitted from final models.
Gated Nonlinearity (used in practice): A separate scalar field $s(x)$ passes through sigmoid to create a gate $\sigma(s(x))$, which multiplies the geometric field: $$f_{\text{out}}(x) = f_{\text{in}}(x) \cdot \sigma(s(x))$$ Architecture implication: Requires extra scalar channels ($l=0$) specifically for gating higher-order channels ($l>0$).
Voxelization Details:
For CATH protein inputs, Gaussian density is placed at each atom position with standard deviation equal to half the voxel width ($0.5 \times 0.2\text{ nm} = 0.1\text{ nm}$).
Evaluation
| Task | Metric | Steerable CNN | Baseline |
|---|---|---|---|
| Tetris (rotated test) | Accuracy | $99 \pm 2%$ | $27 \pm 7%$ (standard 3D CNN) |
| Amino Acid Propensity | Accuracy | 0.58 (32.6M params) | 0.50 (regular grid, 61.1M params); 0.56 (concentric grid, 75.3M params) |
| SHREC17 | micro + macro mAP (higher is better) | 1.11 | 1.13 (SOTA) |
| CATH | Accuracy | Higher across all training set sizes (see Figure 4; not reported as a single value) (143,560 params) | Deep 3D CNN (15,878,764 params; ~110x more) |
Note: On SHREC17, the total score is micro mAP + macro mAP combined (higher is better). From Table 4 in the supplementary material: Steerable CNN micro mAP = 0.661, macro mAP = 0.449, total = 1.11. On CATH, the steerable CNN outperformed the baseline with ~110x fewer parameters, a gap that widened as training data was reduced.
Historical Context (From Peer Reviews)
The NeurIPS peer reviews reveal important context about the paper’s structure and claims:
Evolution of Experiments: The SHREC17 experiment and the arbitrary rotation test in Tetris were added during the rebuttal phase to address reviewer concerns about the lack of standard computer vision benchmarks. This explains why SHREC17 feels somewhat disconnected from the paper’s “AI for Science” narrative.
Continuous vs. Discrete Rotations: The Tetris experiment validates equivariance to continuous ($SO(3)$) rotations alongside discrete 90-degree turns. This distinction is crucial and separates Steerable CNNs from earlier Group CNNs that handled discrete rotation groups exclusively.
Terminology Warning: The use of terms like “fiber” and “induced representation” was critiqued by reviewers as being denser than necessary and inconsistent with related work (e.g., Tensor Field Networks). If you find Section 3 difficult, this is a known barrier of this paper. Focus on the resulting kernel constraints.
Parameter Efficiency Quantified: Reviewers highlighted that the main practical win is parameter efficiency. Standard 3D CNNs hit diminishing returns around $10^7$ parameters, while Steerable CNNs achieve better results with ~110x fewer parameters ($10^5$).
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| se3cnn (GitHub) | Code | MIT | Original implementation; superseded by e3nn for point cloud applications |
| CATH Datasets (GitHub) | Dataset | CC-BY-4.0 | Protein structure classification dataset introduced in this paper |
Pre-trained model weights are not publicly released. Hardware and compute requirements are not specified in the paper.
Paper Information
Citation: Weiler, M., Geiger, M., Welling, M., Boomsma, W., & Cohen, T. S. (2018). 3D steerable CNNs: Learning rotationally equivariant features in volumetric data. Advances in Neural Information Processing Systems, 31. https://proceedings.neurips.cc/paper/2018/hash/488e4104520c6aab692863cc1dba45af-Abstract.html
Publication: NeurIPS 2018
@inproceedings{weiler20183d,
title={3D Steerable CNNs: Learning Rotationally Equivariant Features in Volumetric Data},
author={Weiler, Maurice and Geiger, Mario and Welling, Max and Boomsma, Wouter and Cohen, Taco S},
booktitle={Advances in Neural Information Processing Systems},
volume={31},
year={2018}
}
Additional Resources:
