Many problems in chemistry and biology involve data with geometric structure: molecules have 3D coordinates, proteins have orientations, and physical forces transform predictably under rotation. Geometric deep learning addresses this by building symmetries directly into model architectures. Notes in this section cover equivariant networks, focusing on SO(3) and SE(3) equivariance achieved through group representation theory and spherical harmonics. The goal is to understand how these models work and why the symmetry constraints matter for data efficiency and physical correctness.
| Year | Paper | Key Idea |
|---|---|---|
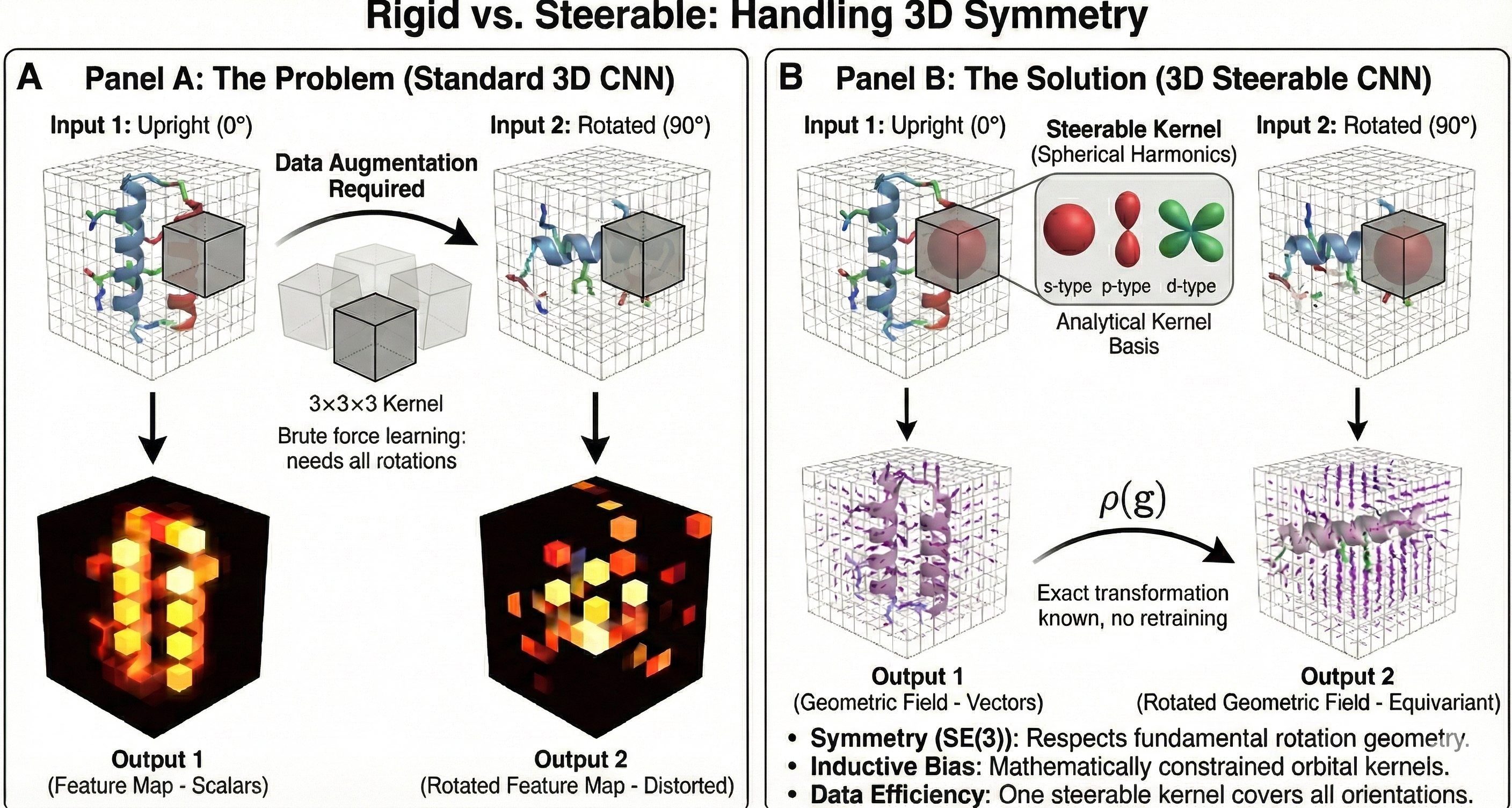
| 2018 | 3D Steerable CNNs: Rotationally Equivariant Features | SE(3)-equivariant features via Wigner D-matrices and spherical harmonics |
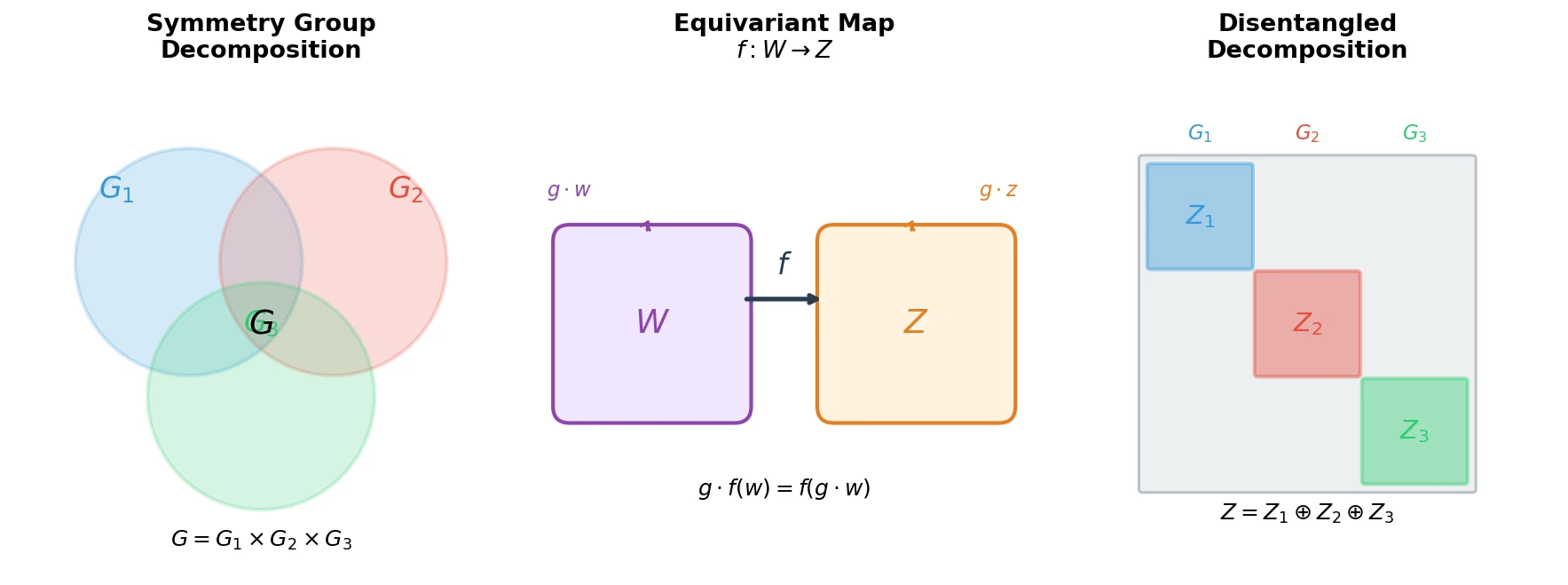
| 2018 | Defining Disentangled Representations via Group Theory | First formal definition of disentanglement using symmetry group decomposition |
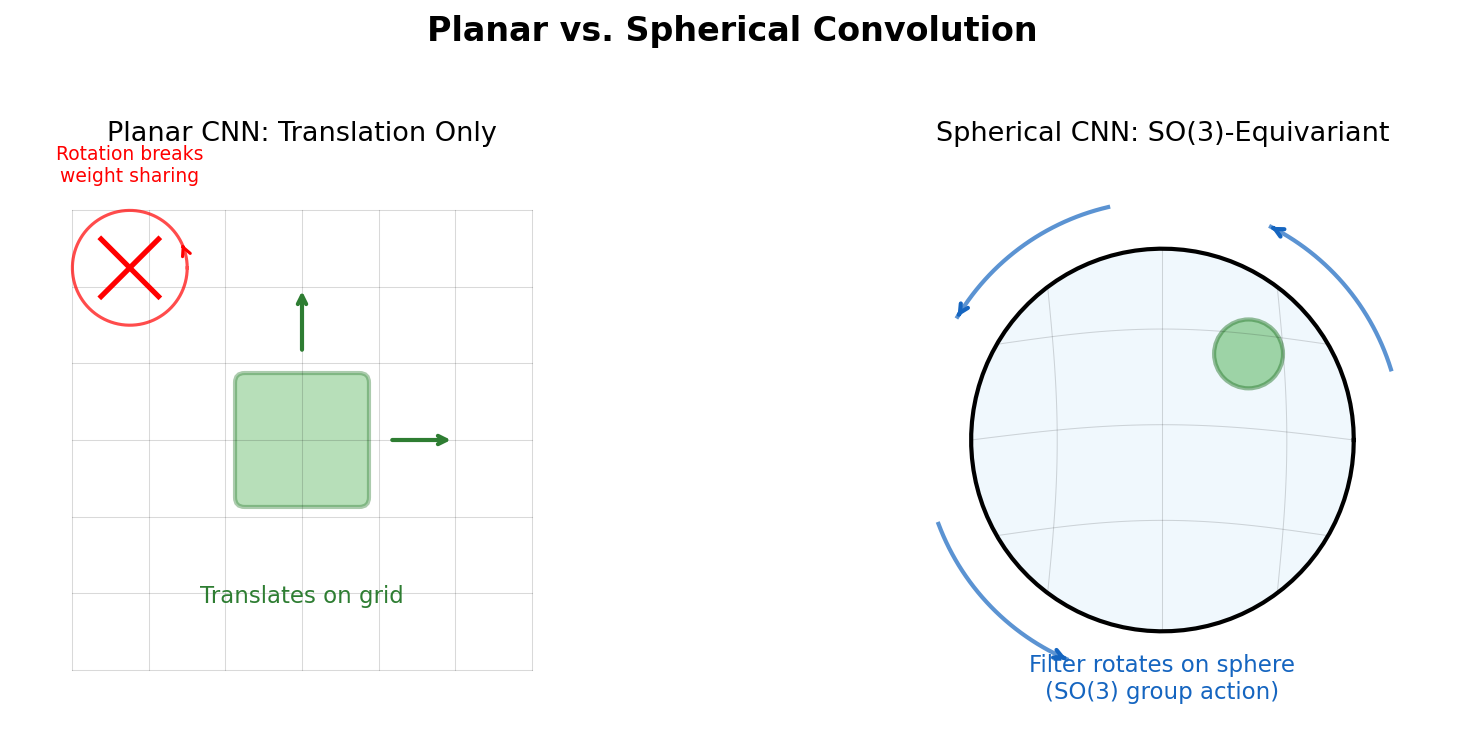
| 2018 | Spherical CNNs: Rotation-Equivariant Networks on the Sphere | SO(3)-equivariance via generalized Fourier transform on the sphere |
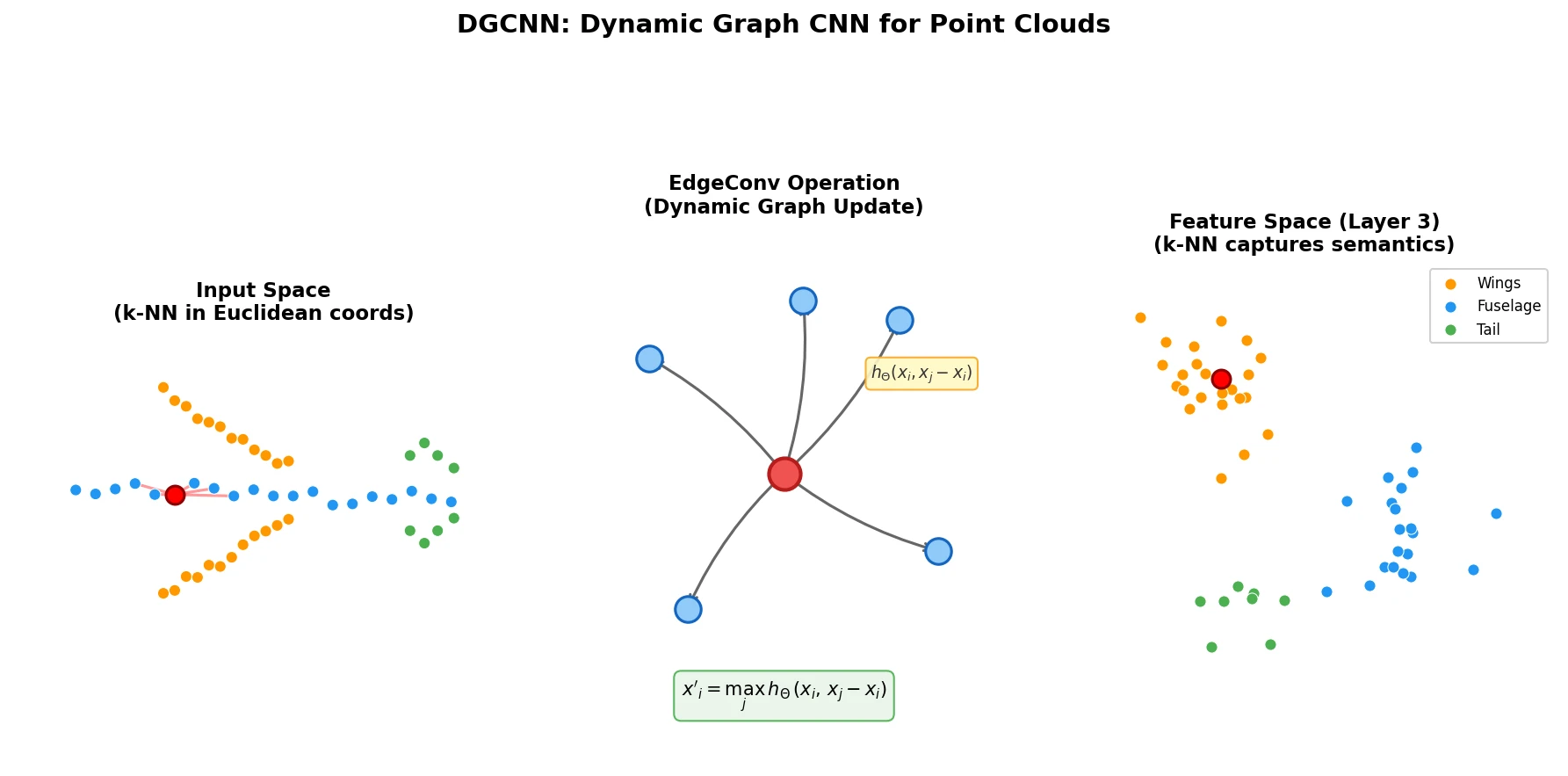
| 2019 | DGCNN: Dynamic Graph CNN for Point Clouds | EdgeConv on dynamically recomputed k-NN graphs in feature space |
| 2020 | SE(3)-Transformers: Equivariant Attention for 3D Data | Self-attention with SE(3)-equivariant type-0/type-1 features |