| GDB-13 | |
|---|---|
| Basic Information | |
| Full Name | Generated Database 13 |
| Domain | Computational Chemistry |
| Year | 2009 |
| Publication & Access | |
| Paper | DOI |
| Dataset | gdb.unibe.ch |
| Dataset Composition | |
| Total Size | 977,468,314 molecules |
| Main C/N/O Set | 910,111,673 molecules |
| Cl/S Set | 67,356,641 molecules |
| Technical Details | |
| Format | SMILES strings |
| Research Context | |
| Authors | Lorenz C. Blum, Jean-Louis Reymond |
| Institution | University of Berne |
GDB Series Overview: The Generated Database (GDB) series represents a systematic exploration of chemical space by generating all possible molecular structures. GDB-11 (26M molecules) established the methodology, GDB-13 (977M molecules) achieved billion-scale generation, and GDB-17 (166B molecules) represents the current limit of systematic chemical space generation.
Dataset Summary
GDB-13 contains nearly one billion small organic molecules created by systematically exploring all possible structures with up to 13 atoms. Building on the approach from GDB-11, this represents a substantial scale-up from millions to nearly a billion molecules while maintaining chemical quality. The dataset covers drug-like chemical space for virtual screening applications. All molecules are provided as SMILES strings.
Related Databases: GDB-13 is part of the Generated Database (GDB) series, following GDB-11 (26 million molecules) and preceding GDB-17 (166 billion molecules), showing the evolution of systematic chemical space exploration.
Key Features
- Large scale: Nearly 1 billion molecules - a significant achievement at the time of publication
- Complete coverage: Systematically covers all possible structures with up to 13 atoms
- Drug-like properties: All molecules follow Lipinski’s Rule of Five for drug-likeness
- Quality filtering: Filtered to remove chemically unrealistic or unstable structures
Dataset Structure
| Composition | Druglike | Fragmentlike | Leadlike | Molecules | Subset |
|---|---|---|---|---|---|
| C, N, O only | 100% | 45.1% | 98.9% | 910M | Main Set |
| 100% | 67M | Cl/S Set | ||
| All molecules | 100% | 45.1% | 98.9% | 977M | Total |
Structural Diversity
- Heterocycles: 71% of molecules contain rings with nitrogen or oxygen atoms
- Small rings: 54% contain strained 3- or 4-membered rings
- Graph types: Most molecules have complex ring systems - 43.8% are polycyclic and 34.6% are tricyclic
Example Sample
CCCC(O)(CO)CC1CC1CN
Visualized with PubChem Sketcher:
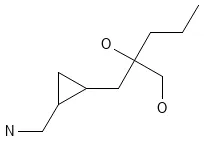
Representative GDB-13 molecule (SMILES: CCCC(O)(CO)CC1CC1CN) demonstrating typical structural features: small rings, heteroatoms, and druglike properties
Use Cases
Primary Applications
- Virtual Screening: Novel drug scaffolds across therapeutic areas
- Fragment-Based Drug Discovery: Comprehensive fragment space coverage
- Chemical Space Exploration: Large-scale structure-property relationship studies
Research Applications
- Algorithm Benchmarking: Virtual screening and molecular descriptor development
- ML Model Development: Training molecular property prediction models
- Structure-Property Studies: Large-scale analysis of chemical relationships
Quality & Limitations
Strengths
- Large Scale: Nearly 1 billion molecules, significant for its time of publication
- Complete Drug-likeness: 100% Lipinski compliance across all molecules
- High Novelty: Novel chemical structures absent from commercial databases
- Coverage: Complete coverage of all possible structures with up to 13 atoms
- Refined Methodology: Improved approach enabling larger databases like GDB-17
Limitations
- Limited Atom Types: Main set only includes carbon, nitrogen, and oxygen (subset includes Cl and S; GDB-17 expanded to include more elements)
- Structural Constraints: Excludes fused small rings and highly strained molecules
- Element Ratio Restrictions: Filters out highly polar molecules with strict heteroatom ratios
- Missing Functional Groups: Excludes peroxides and many unstable intermediates
- Virtual Molecules: Computer-generated without experimental validation or synthetic accessibility scoring
- Size Constraint: Maximum 13 heavy atoms limits coverage of larger drug-like molecules
Generation and Filtering Pipeline
GDB-13 refined and scaled the methodology pioneered in GDB-11, achieving nearly 40-fold increase in database size while maintaining chemical quality. This approach later enabled the massive GDB-17 database. The generation process involves sophisticated multi-step filtering to ensure chemical feasibility and druglikeness:
Graph Generation & Topological Filtering
- Initial enumeration: 27.3M graphs generated using GENG program
- Topo I filter: Removes fused small rings (97.2% rejection rate)
- Topo II filter: Eliminates bridgehead atoms in 3-4 membered rings (56.9% rejection)
- SAV filter: 3D strain analysis using tetrahedron volumes (4.7% rejection)
Chemical Intelligence Filters
- Element ratio limits: N/C < 0.571, O/C < 0.667, (N+O)/C < 1.0
- Heteroatom bonds: Strict rules for N-N, N-O, O-O bond patterns
- Functional group exclusions: Removes unstable groups (hemiacetals, orthoesters, etc.)
- Tautomer standardization: Selects most stable tautomeric forms
Post-Processing Expansions
The Cl/S subset adds chemical diversity through systematic transformations:
- Aromatic nitro groups from carboxylic acids
- Nitriles from aldehydes
- Aromatic chlorines from hydroxyl groups
- Thiophene analogs from oxygen heterocycles
- Sulfonamides and thioureas from carbonyl groups
This rigorous filtering explains the high druglikeness compliance while maintaining chemical diversity. The methodology proved so successful that it enabled the creation of the vastly larger GDB-17 database containing 166 billion molecules.
Citation: Blum, L. C. & Reymond, J.-L. “970 Million Druglike Small Molecules for Virtual Screening in the Chemical Universe Database GDB-13” J. Am. Chem. Soc. 2009, 131 (25), pp 8732–8733.