| GDB-11 | |
|---|---|
| Basic Information | |
| Full Name | Generated Database 11 |
| Domain | Computational Chemistry |
| Year | 2007 |
| Publication & Access | |
| Paper | DOI |
| Dataset | gdb.unibe.ch |
| Dataset Composition | |
| Total Size | 26,434,571 molecules |
| Unique Tautomers (GDB) | 26,434,571 molecules |
| Stereoisomers | 110,979,507 stereoisomers |
| Technical Details | |
| Format | SMILES strings |
| Research Context | |
| Authors | Tobias Fink, Jean-Louis Reymond |
| Institution | University of Berne |
GDB Series Overview: The Generated Database (GDB) series represents a systematic exploration of chemical space by generating all possible molecular structures. GDB-11 (26M molecules) established the methodology, GDB-13 (977M molecules) achieved billion-scale generation, and GDB-17 (166B molecules) represents the current limit of systematic chemical space generation.
Dataset Summary
GDB-11 contains 26.4 million small organic molecules that were systematically generated by exploring all possible structures with up to 11 atoms of carbon, nitrogen, oxygen, and fluorine. This was the founding database in the GDB series and established the approach for exploring chemical space computationally. The dataset is useful for virtual screening and finding new molecular structures for drug discovery. All molecules are provided as SMILES strings.
Related Databases: GDB-11 is part of the Generated Database (GDB) series, which includes the larger GDB-13 (977 million molecules) and GDB-17 (166 billion molecules). Each database expands to larger molecules and includes more atom types.
Key Features
- Complete Coverage: Systematically covers all possible structures with up to 11 atoms
- Drug-like Properties: All molecules follow Lipinski’s Rule of Five, with half meeting the stricter Rule of Three
- Novel Structures: Includes 1,208 unique ring systems, 538 of which were previously unknown
- Quality Filtering: Filtered to remove unstable or chemically unrealistic structures
- Series Foundation: Established the generation methodology for the GDB series
Dataset Structure
| Category | Composition | Count | Druglike rule of 5 | Leadlike rule of 3 |
|---|---|---|---|---|
| Unique Tautomers | C, N, O, F atoms | 26.4 M | 100% | 50% |
| Total Stereoisomers | C, N, O, F atoms | 110.9 M | 100% |
Structural Diversity
- Ring Systems: Most molecules have rings - 43% have one ring, 32% have two rings, 9% have three rings, and 1% have more complex ring systems. Only 15% have no rings at all
- Chirality: Over 70% of molecules are chiral (have handedness), especially larger molecules
- Functional Groups: Covers many different functional groups, limited by the four atom types allowed
Example Sample
FC1C2OC1c3c(F)coc23
Visualized with PubChem Sketcher:
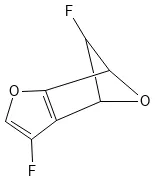
Representative GDB-11 molecule (SMILES: FC1C2OC1c3c(F)coc23)
Use Cases
Primary Applications
- Virtual Screening: Search for new drug candidates like kinase inhibitors and GPCR ligands
- Fragment-Based Drug Discovery: Use the many small, drug-like molecules as starting points
- New Scaffold Discovery: Find novel molecular frameworks from the previously unknown ring systems
Research Applications
- Chemical Space Studies: Understand the landscape of possible small molecules
- Machine Learning: Train and test computational chemistry and cheminformatics models
- Structure-Property Research: Study how molecular shape affects chemical behavior
Quality & Limitations
Strengths
- Complete Coverage: Systematic coverage of all possible structures with up to 11 atoms
- High Drug-likeness: 100% Lipinski compliance, 50% Rule of Three compliance
- Structural Novelty: 538 previously unknown ring systems
- Foundation for Series: Established the generation methodology for the GDB series
Limitations
- Limited Atom Types: Only includes carbon, nitrogen, oxygen, and fluorine (later databases like GDB-13 and GDB-17 include more elements)
- Structural Constraints: Excludes highly strained molecules and some bond patterns
- Missing Functional Groups: Doesn’t include unstable groups like hemiacetals and gem-diols
- Virtual Molecules: These are computer-generated structures, not experimentally validated compounds
- Size Limitation: Maximum of 11 heavy atoms limits complexity compared to many real drugs
Generation and Filtering Pipeline
GDB-11 established the rigorous methodology later refined in GDB-13 and GDB-17. The generation process involves sophisticated multi-step filtering:
Graph Selection and Validation
- Initial Enumeration: 840,000+ mathematical graphs generated
- Topological Filtering: Removal of chemically impossible structures (fused small rings)
- Energy Minimization: MM2 calculations to eliminate high-strain configurations
- Final Selection: 15,726 stable graphs selected
Chemical Intelligence Filters
- Unsaturation Assignment: Systematic addition of double/triple bonds
- Element Substitution: Combinatorial C→N,O,F replacements following valency rules
- Functional Group Validation: Removal of unstable groups (hemiacetals, enols, aminals)
- Tautomer Standardization: Selection of most stable forms
This methodology proved so effective that it was scaled and refined for the much larger GDB-13 (nearly 1 billion molecules) and GDB-17 (166 billion molecules) databases.
Citation: Fink, T. & Reymond, J.-L. “Virtual Exploration of the Chemical Universe up to 11 Atoms of C, N, O, F: Assembly of 26.4 Million Structures (110.9 Million Stereoisomers) and Analysis for New Ring Systems, Stereochemistry, Physicochemical Properties, Compound Classes, and Drug Discovery” J. Chem. Inf. Model. 2007, 47 (2), pp 342–353.