Dataset Examples
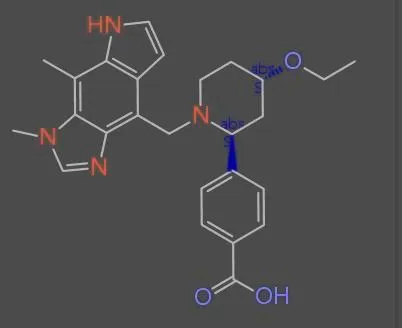
Dataset Subsets
| Subset | Count | Description |
|---|---|---|
| MolParser-7M (Training Set) | 7,740,871 | A large-scale dataset for training OCSR models, split into pre-training and fine-tuning stages. |
| WildMol (Test Set) | 20,000 | A benchmark of 20,000 human-annotated samples cropped from real PDF files to evaluate OCSR models in ‘in-the-wild’ scenarios. Comprises WildMol-10k (10k ordinary molecules) and WildMol-10k-M (10k Markush structures). |
Benchmarks
WildMol-10K Accuracy
Evaluation of OCSR models on 10,000 real-world molecular images cropped from scientific literature and patents
| Rank | Model | Accuracy (%) |
|---|---|---|
| 🥇 1 | MolParser-Base End-to-end visual recognition trained on MolParser-7M | 76.9 |
| 🥈 2 | MolScribe Transformer-based OCSR system | 66.4 |
| 🥉 3 | DECIMER 2.7 Deep learning for chemical image recognition | 56 |
| 4 | MolGrapher Graph-based molecular structure recognition | 45.5 |
| 5 | MolVec 0.9.7 Vector-based structure recognition | 26.4 |
| 6 | OSRA 2.1 Optical Structure Recognition Application | 26.3 |
| 7 | Img2Mol Image-to-molecule translation | 24.4 |
| 8 | Imago 2.0 Chemical structure recognition toolkit | 6.9 |
Key Contribution
Introduces MolParser-7M, the largest open-source Optical Chemical Structure Recognition (OCSR) dataset, uniquely combining diverse synthetic data with a large volume of manually-annotated, “in-the-wild” images from real scientific documents to improve model robustness. Also introduces WildMol, a new challenging benchmark for evaluating OCSR performance on real-world data, including Markush structures.
Overview
The MolParser project addresses the challenge of recognizing molecular structures from images found in real-world scientific documents. Unlike existing OCSR datasets that rely primarily on synthetically generated images, MolParser-7M incorporates 400,000 manually annotated images cropped from actual patents and scientific papers, making it the first large-scale dataset to bridge the gap between synthetic training data and real-world deployment scenarios.
Strengths
- Largest open-source OCSR dataset with over 7.7 million pairs
- The only large-scale OCSR training set that includes a significant amount (400k) of “in-the-wild” data cropped from real patents and literature
- High diversity of molecular structures from numerous sources (PubChem, ChEMBL, polymers, etc.)
- Introduces the WildMol benchmark for evaluating performance on challenging, real-world data, including Markush structures
- The “in-the-wild” fine-tuning data (MolParser-SFT-400k) was curated via an efficient active learning data engine with human-in-the-loop validation
Limitations
- The E-SMILES format cannot represent certain complex cases, such as coordination bonds, dashed abstract rings, Markush structures depicted with special patterns, and replication of long structural segments on the skeleton
- The model and data do not yet fully exploit molecular chirality, which is critical for chemical properties
- Performance could be further improved by scaling up the amount of real annotated training data
Technical Notes
Synthetic Data Generation
To ensure diversity, molecular structures were collected from databases like ChEMBL, PubChem, and Kaggle BMS. A significant number of Markush, polymer, and fused-ring structures were also randomly generated. Images were rendered using RDKit and epam.indigo with randomized parameters (e.g., bond width, font size, rotation) to increase visual diversity. The pretraining dataset is composed of the following subsets:
| Subset | Ratio | Source |
|---|---|---|
| Markush-3M | 40% | Random groups replacement from PubChem |
| ChEMBL-2M | 27% | Molecules selected from ChEMBL |
| Polymer-1M | 14% | Randomly generated polymer molecules |
| PAH-600k | 8% | Randomly generated fused-ring molecules |
| BMS-360k | 5% | Molecules with long carbon chains from BMS |
| MolGrapher-300K | 4% | Training data from MolGrapher |
| Pauling-100k | 2% | Pauling-style images drawn using epam.indigo |
In-the-Wild Data Engine (MolParser-SFT-400k)
A YOLO11 object detection model (MolDet) located and cropped over 20 million molecule images from 1.22 million real PDFs (patents and papers). After de-duplication via p-hash similarity, 4 million unique images remained.
An active learning algorithm was used to select the most informative samples for annotation, targeting images where an ensemble of 5-fold models showed moderate confidence (0.6-0.9 Tanimoto similarity), indicating they were challenging but learnable.
This active learning approach with model pre-annotations reduced manual annotation time per molecule to 30 seconds, approximately 90% savings compared to annotating from scratch. In the final fine-tuning dataset, 56.04% of annotations directly utilized raw model pre-annotations, 20.97% passed review after a single manual correction, 13.87% were accepted after a second round of annotation, and 9.13% required three or more rounds.
The fine-tuning dataset is composed of:
| Subset | Ratio | Source |
|---|---|---|
| MolParser-SFT-400k | 66% | Manually annotated data obtained via data engine |
| MolParser-Gen-200k | 32% | Synthetic data selected from pretraining stage |
| Handwrite-5k | 1% | Handwritten molecules selected from Img2Mol |
E-SMILES Specification
To accommodate complex patent structures that standard SMILES cannot support, the authors introduced an Extended SMILES format (SMILES<sep>EXTENSION). The EXTENSION component uses XML-like tokens to manage complexities:
<a>...</a>encapsulates Markush R-groups and abbreviation groups.<r>...</r>denotes ring attachments with uncertainty positions.<c>...</c>defines abstract rings.<dum>identifies a connection point.
This format enables Markush-molecule matching and LLM integration, while retaining RDKit compatibility for the standard SMILES portion.
Reproducibility
| Artifact | Type | License | Notes |
|---|---|---|---|
| MolParser-7M | Dataset | CC-BY-NC-SA-4.0 | Training and test data on HuggingFace. SFT subset is partially released. |
| MolDet (YOLO11) | Model | Unknown | Molecule detection model on HuggingFace |
| MolParser Demo | Other | N/A | Online OCSR demo using MolParser-Base |
The dataset is publicly available on HuggingFace under a CC-BY-NC-SA-4.0 (non-commercial) license. The MolParser-SFT-400k subset is only partially released. The YOLO11-based MolDet detection model is also available on HuggingFace. No public code repository is provided for the MolParser recognition model itself. All experiments were conducted on 8 NVIDIA RTX 4090D GPUs, and throughput benchmarks were measured on a single RTX 4090D GPU.
