Paper Information
Citation: Wang, J., He, Y., Yang, H., Wu, J., Ge, L., Wei, X., Wang, Y., Li, L., Ao, H., Liu, C., Wang, B., Wu, L., & He, C. (2025). GTR-CoT: Graph Traversal as Visual Chain of Thought for Molecular Structure Recognition (arXiv:2506.07553). arXiv. https://doi.org/10.48550/arXiv.2506.07553
Publication: arXiv preprint (2025)
Additional Resources:
Contribution: Vision-Language Modeling for OCSR
This is a method paper that introduces GTR-VL, a Vision-Language Model for Optical Chemical Structure Recognition (OCSR). The work addresses the persistent challenge of converting molecular structure images into machine-readable formats, with a particular focus on handling chemical abbreviations that cause errors in existing systems.
Motivation: The Abbreviation Bottleneck
The motivation tackles a long-standing bottleneck in chemical informatics: most existing OCSR systems produce incorrect structures when they encounter abbreviated functional groups. When a chemist draws “Ph” for phenyl or “Et” for ethyl, current models fail because they have been trained on data where images contain abbreviations but the ground-truth labels contain fully expanded molecular graphs.
This creates a fundamental mismatch. The model sees “Ph” in the image but is told the “correct” answer is a full benzene ring. The supervision signal is inconsistent with what is actually visible.
Beyond this data problem, existing graph-parsing methods use a two-stage approach: predict all atoms first, then predict all bonds. This is inefficient and ignores the structural constraints that could help during prediction. The authors argue that mimicking how humans analyze molecular structures - following bonds from atom to atom in a connected traversal - would be more effective.
Novelty: Graph Traversal as Visual Chain-of-Thought
The novelty lies in combining two key insights about how to properly train and architect OCSR systems. The main contributions are:
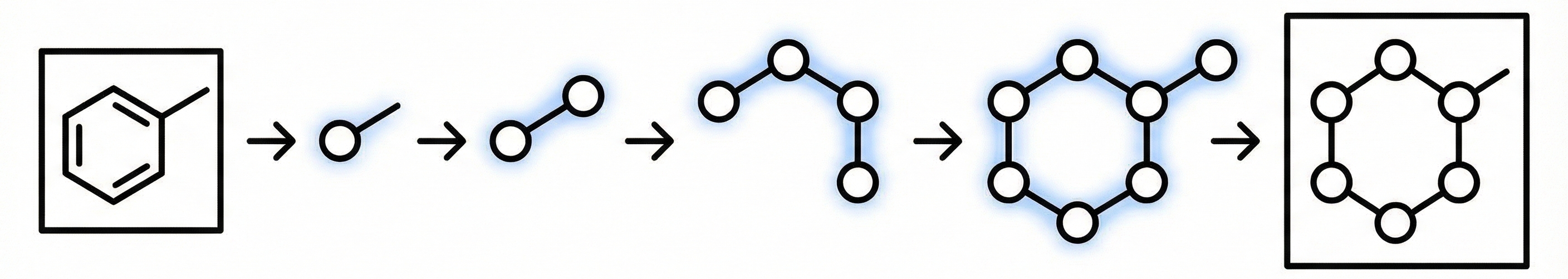
Graph Traversal as Visual Chain of Thought: GTR-VL generates molecular graphs by traversing them sequentially, predicting an atom, then its connected bond, then the next atom, and so on. This mimics how a human chemist would trace through a structure and allows the model to use previously predicted atoms and bonds as context for subsequent predictions.
Formally, the model output sequence for image $I_m$ is generated as:
$$ R_m = \text{concat}(CoT_m, S_m) $$
where $CoT_m$ represents the deterministic graph traversal steps (atoms and bonds) and $S_m$ is the final SMILES representation. This intermediate reasoning step makes the model more interpretable and helps it learn the structural logic of molecules.
“Faithfully Recognize What You’ve Seen” Principle: This addresses the abbreviation problem head-on. The authors correct the ground-truth annotations to match what’s actually visible in the image.
They treat abbreviations like “Ph” as single “superatoms” and build a pipeline to automatically detect and correct training data. Using OCR to extract visible text from molecular images, they replace the corresponding expanded substructures in the ground-truth with the appropriate abbreviation tokens. This ensures the supervision signal is consistent with the visual input.
Large-Scale Dataset (GTR-1.3M): To support this approach, the authors created a large-scale dataset combining 1M synthetic molecules from PubChem with 351K corrected real-world patent images from USPTO. The key innovation is the correction pipeline that identifies abbreviations in patent images and fixes the inconsistent ground-truth labels.
GRPO for Hand-Drawn OCSR: Hand-drawn molecular data lacks fine-grained atom/bond coordinate annotations, making SFT-based graph parsing inapplicable. The authors use Group Relative Policy Optimization (GRPO) with a composite reward function that combines format, SMILES, and graph-level rewards. The graph reward computes the maximum common subgraph (MCS) between predicted and ground-truth molecular graphs:
$$ R_{\text{graph}} = \frac{|N_m^a|}{|N_g^a| + |N_p^a|} + \frac{|N_m^b|}{|N_g^b| + |N_p^b|} $$
where $N_m^a$, $N_g^a$, $N_p^a$ are atom counts in the MCS, ground truth, and prediction, and $N_m^b$, $N_g^b$, $N_p^b$ are the corresponding bond counts.
Two-Stage Training: Stage 1 performs SFT on GTR-1.3M for printed molecule recognition. Stage 2 applies GRPO on a mixture of printed data (GTR-USPTO-4K) and hand-drawn data (DECIMER-HD-Train, 4,070 samples) to extend capabilities to hand-drawn structures.
MolRec-Bench Evaluation: Traditional SMILES-based evaluation fails for molecules with abbreviations because canonicalization breaks down. The authors created a new benchmark that evaluates graph structure directly, providing three metrics: direct SMILES generation, graph-derived SMILES, and exact graph matching.
What experiments were performed?
The evaluation focused on demonstrating that GTR-VL’s design principles solve real problems that plague existing OCSR systems:
Comprehensive Baseline Comparison: GTR-VL was tested against three categories of models:
- Specialist OCSR systems: MolScribe and MolNexTR
- Chemistry-focused VLMs: ChemVLM, ChemDFM-X, OCSU
- General-purpose VLMs: GPT-4o, GPT-4o-mini, Qwen-VL-Max
MolRec-Bench Evaluation: The new benchmark includes two subsets of patent images:
- MolRec-USPTO: 5,423 standard patent images similar to existing benchmarks
- MolRec-Abb: 9,311 molecular images with abbreviated superatoms, derived from MolGrapher’s USPTO 10K abb subset
This design directly tests whether models can handle the abbreviation problem that breaks existing systems.
Ablation Studies: Systematic experiments isolated the contribution of key design choices:
- Chain-of-Thought vs. Direct: Comparing graph traversal CoT against direct SMILES prediction
- Traversal Strategy: Graph traversal vs. the traditional “atoms-then-bonds” approach
- Dataset Quality: Training on corrected vs. uncorrected data
Retraining Experiments: Existing specialist models (MolScribe, MolNexTR) were retrained from scratch on the corrected GTR-1.3M dataset to isolate the effect of data quality from architectural improvements.
Hand-Drawn OCSR Evaluation: GTR-VL was also evaluated on the DECIMER Hand-drawn test set and ChemPix dataset, comparing against DECIMER and AtomLenz+EditKT baselines.
Qualitative Analysis: Visual inspection of predictions on challenging cases with heavy abbreviation usage, complex structures, and edge cases to understand failure modes.
Results & Conclusions: Resolving the Abbreviation Bottleneck
Performance Gains on Abbreviations: On MolRec-Abb, GTR-VL-Stage1 achieves 85.49% Graph accuracy compared to around 20% for MolScribe and MolNexTR with their original checkpoints. On MolRec-USPTO, GTR-VL-Stage1 reaches 93.45% Graph accuracy. Existing specialist models see their accuracy drop below 20% on MolRec-Abb when abbreviations are present.
Data Correction is Critical: When MolScribe and MolNexTR were retrained on GTR-1.3M, their MolRec-Abb Graph accuracy jumped from around 20% to 70.60% and 71.85% respectively. GTR-VL-Stage1 still outperformed these retrained baselines at 85.49%, confirming that both data correction and the graph traversal approach contribute.
Chain-of-Thought Helps: Ablation on GTR-USPTO-351K shows that CoT yields 68.85% Gen-SMILES vs. 66.54% without CoT, a 2.31 percentage point improvement.
Graph Traversal Beats Traditional Parsing: Graph traversal achieves 83.26% Graph accuracy vs. 80.15% for the atoms-then-bonds approach, and 81.88% vs. 79.02% on Gra-SMILES.
General VLMs Still Struggle: General-purpose VLMs like GPT-4o scored near 0% on MolRec-Bench across all metrics, highlighting the importance of domain-specific training for OCSR.
Hand-Drawn Recognition via GRPO: GTR-VL-Stage1 (SFT only) achieves only 9.53% Graph accuracy on DECIMER-HD-Test, but after GRPO training in Stage 2, performance jumps to 75.44%. On ChemPix, Graph accuracy rises from 22.02% to 86.13%. The graph reward is essential: GRPO without graph supervision achieves only 11.00% SMILES on DECIMER-HD-Test, while adding graph reward reaches 75.64%.
Evaluation Methodology Matters: The new graph-based evaluation metrics revealed problems with traditional SMILES-based evaluation that previous work had missed. Many “failures” in existing benchmarks were actually correct graph predictions that got marked wrong due to canonicalization issues with abbreviations.
The work establishes that addressing the abbreviation problem requires both correcting the training data and rethinking the model architecture. The combination of faithful data annotation and sequential graph generation improves OCSR performance on molecules with abbreviations by a large margin over previous methods.
Reproducibility Details
Models
Base Model: GTR-VL fine-tunes Qwen2.5-VL.
Input/Output Mechanism:
- Input: The model takes an image $I_m$ and a text prompt
- Output: The model generates $R_m = \text{concat}(CoT_m, S_m)$, where it first produces the Chain-of-Thought (the graph traversal steps) followed immediately by the final SMILES string
- Traversal Strategy: Uses depth-first traversal to alternately predict atoms and bonds
Prompt Structure: The model is prompted to “list the types of atomic elements… the coordinates… and the chemical bonds… then… output a canonical SMILES”. The CoT output is formatted as a JSON list of atoms (with coordinates) and bonds (with indices referring to previous atoms), interleaved.
Data
Training Dataset (GTR-1.3M):
- Synthetic Component: 1 million molecular SMILES from PubChem, converted to images using Indigo
- Real Component: 351,000 samples from USPTO patents (filtered from an original 680,000)
- Processed using an OCR pipeline to detect abbreviations (e.g., “Ph”, “Et”)
- Ground truth expanded structures replaced with superatoms to match visible abbreviations in images
- This “Faithfully Recognize What You’ve Seen” correction ensures training supervision matches visual input
Evaluation Dataset (MolRec-Bench):
- MolRec-USPTO: 5,423 molecular images from USPTO patents
- MolRec-Abb: 9,311 molecular images with abbreviated superatoms, derived from MolGrapher’s USPTO 10K abb subset
Algorithms
Graph Traversal Algorithm:
- Depth-first traversal strategy
- Alternating atom-bond prediction sequence
- Each step uses previously predicted atoms and bonds as context
Two-Stage Training:
- Stage 1 (SFT): Train on GTR-1.3M to learn visual CoT mechanism for printed molecules (produces GTR-VL-Stage1)
- Stage 2 (GRPO): Apply GRPO on GTR-USPTO-4K + DECIMER-HD-Train (4,070 samples) for hand-drawn recognition (produces GTR-VL-Stage2, i.e., GTR-VL)
Training Procedure:
- Optimizer: AdamW
- Learning Rate (SFT): Peak learning rate of $1.6 \times 10^{-4}$ with cosine decay
- Learning Rate (GRPO): Peak learning rate of $1 \times 10^{-5}$ with cosine decay
- Warm-up: Linear warm-up for the first 10% of iterations
- Batch Size (SFT): 2 per GPU with gradient accumulation over 16 steps, yielding effective batch size of 1024
- Batch Size (GRPO): 4 per GPU with gradient accumulation of 1, yielding effective batch size of 128
Evaluation
Metrics (three complementary measures to handle abbreviation issues):
- Gen-SMILES: Exact match ratio of SMILES strings directly generated by the VLM (image-captioning style)
- Gra-SMILES: Exact match ratio of SMILES strings derived from the predicted graph structure (graph-parsing style)
- Graph: Exact match ratio between ground truth and predicted graphs (node/edge comparison, bypassing SMILES canonicalization issues)
Baselines Compared:
- Specialist OCSR systems: MolScribe, MolNexTR
- Chemistry-focused VLMs: ChemVLM, ChemDFM-X, OCSU
- General-purpose VLMs: GPT-4o, GPT-4o-mini, Qwen-VL-Max
Hardware
Compute: Training performed on 32 NVIDIA A100 GPUs
Reproducibility Status
Status: Closed. As of the paper’s publication, no source code, pre-trained model weights, or dataset downloads (GTR-1.3M, MolRec-Bench) have been publicly released. The paper does not mention plans for open-source release. The training data pipeline relies on PubChem SMILES (public), USPTO patent images (publicly available through prior work), the Indigo rendering tool (open-source), and an unspecified OCR system for abbreviation detection. Without the released code and data corrections, reproducing the full pipeline would require substantial re-implementation effort.