Paper Information
Citation: Wang, H., Yu, Y., & Liu, J.-C. (2026). GraphReco: Probabilistic Structure Recognition for Chemical Molecules. ChemistryOpen, e202500537. https://doi.org/10.1002/open.202500537
Publication: ChemistryOpen 2026 (Open Access)
A Rule-Based OCSR System with Probabilistic Graph Assembly
GraphReco tackles a challenge that is rarely addressed explicitly in rule-based OCSR: the ambiguity that arises during graph assembly when lower-level component extraction results are imprecise. Small deviations in bond endpoint locations, false positive detections, and spatial proximity between elements all create uncertainty about which atoms and bonds should be connected, merged, or discarded.
The system introduces two main contributions:
- Fragment Merging (FM) line detection: An adaptive three-stage algorithm for precise bond line identification across images of variable resolution
- Probabilistic ambiguity resolution: A Markov network that infers the most likely existence and merging state of atom and bond candidates
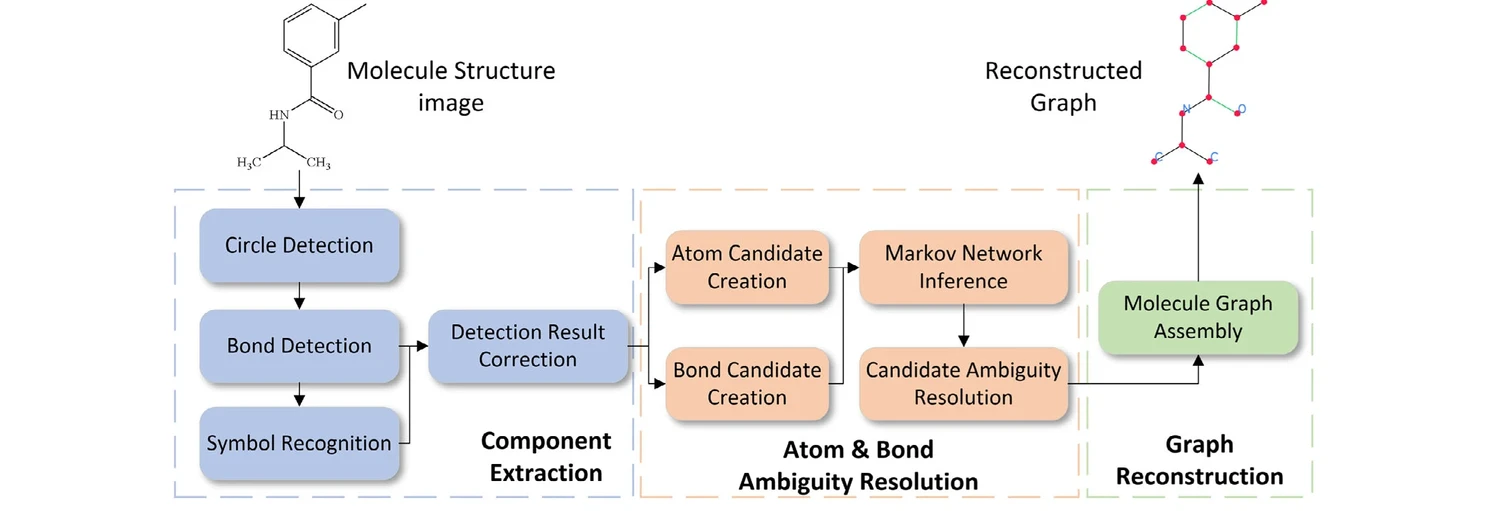
Three-Stage Pipeline
GraphReco follows a three-stage workflow:
Component Extraction: Detects circles (aromatic bonds), bond lines (via the FM algorithm), and chemical symbols (via Tesseract OCR). Includes detection of solid wedge, dashed wedge, dashed line, and wavy bond styles. A semi-open-loop correction step resolves cases where symbols are misclassified as bonds and vice versa.
Atom and Bond Ambiguity Resolution: Creates atom and bond candidates from detected components, builds a Markov network to infer their most probable states, and resolves candidates through existence and merging decisions.
Graph Reconstruction: Assembles resolved atoms and bonds into a molecule graph, selects the largest connected component, and exports as MDL Molfile.
Fragment Merging Line Detection
Classical Line Hough Transform (LHT) struggles with chemical structure images because bond lines suffer from pixelization, and algorithm parameters that work for one image resolution fail at others. The FM algorithm addresses this with three stages:
Fragment extraction: Apply LHT with high-resolution parameters (resolution $r = 2$, resolution $\theta = 2°$) to detect fine line fragments. Walk along detected theoretical lines to find actual black pixels and group them by connectivity.
Fragment grouping: Pair fragments that share similar angles, are close in the perpendicular direction, and are either overlapping or connected by a path of black pixels.
Fragment merging: Merge grouped fragments into single line segments using the two border pixels farthest from the centroid.
The FM algorithm effectively handles the tradeoff that plagues standard LHT: coarse parameters miss short lines and produce overlaps, while fine parameters return many fragments shorter than actual bonds.
Probabilistic Ambiguity Resolution via Markov Network
After component extraction, GraphReco creates atom and bond candidates rather than directly assembling the graph. Each bond endpoint generates an atom candidate with a circular bounding area of radius:
$$r_b = \min(l_{\text{bond}}, l_{\text{med}}) / 4$$
where $l_{\text{bond}}$ is the bond length and $l_{\text{med}}$ is the median bond length.
A Markov network is constructed with four types of nodes:
- Atom nodes: Boolean existence variables for each atom candidate
- Bond nodes: Boolean existence variables for each bond candidate
- Atom merge nodes: Boolean variables for pairs of overlapping atom candidates
- Bond merge nodes: Boolean variables for pairs of nearby bond candidates
Potential functions encode rules about when candidates should exist or merge, with merging likelihood between two bond-ending atom candidates defined as a piecewise function of center distance $d$:
$$P(a_1, a_2) = \begin{cases} 0.9, & \text{if } d \leq Q \\ 0.7 - 0.4(d - Q)/(R - Q), & \text{if } Q < d \leq R \\ 0.1, & \text{if } d > R \end{cases}$$
where $Q = \max(r_1, r_2)$ and $R = \min(1.5Q, r_1 + r_2)$. MAP inference determines the final state of all candidates.
Evaluation Results
GraphReco is evaluated on USPTO benchmarks with InChI string comparison (stereochemistry removed):
| System | USPTO-10K | USPTO-10K-Abb | USPTO |
|---|---|---|---|
| GraphReco | 94.2 | 86.7 | 89.9 |
| MolVec 0.9.7 | 92.4 | 70.3 | 89.1 |
| Imago 2.0 | 89.9 | 63.0 | 89.4 |
| OSRA 2.1 | 89.7 | 63.9 | 89.3 |
| MolGrapher | 93.3 | 82.8 | 91.5 |
| Img2Mol | 35.4 | 13.8 | 25.2 |
GraphReco outperforms all rule-based systems and most ML systems, with a particularly large margin on USPTO-10K-Abb (abbreviation-heavy molecules). MolGrapher achieves slightly higher accuracy on the USPTO dataset.
Robustness on Perturbed Images
On USPTO-perturbed (rotation and shearing applied), rule-based methods degrade substantially:
| System | USPTO-perturbed |
|---|---|
| MolGrapher | 86.7 |
| Img2Mol | 42.3 |
| GraphReco | 40.6 |
| MolVec 0.9.7 | 30.7 |
| OSRA 2.1 | 6.4 |
| Imago 2.0 | 5.1 |
GraphReco performs better than other rule-based systems on perturbed inputs (40.6% vs. under 31%) thanks to its probabilistic assembly, but still falls far behind MolGrapher (86.7%), demonstrating the robustness advantage of learned approaches.
Ablation Study
Each component contributes substantially to overall performance on USPTO-10K:
| Configuration | USPTO-10K | USPTO-10K-Abb | USPTO |
|---|---|---|---|
| Full system | 94.2 | 86.7 | 89.9 |
| Without FM line detection | 2.9 | 5.5 | 4.8 |
| Without atom candidates | 9.8 | 0.4 | 5.0 |
| Without bond candidates | 79.1 | 75.8 | 75.0 |
| Without Markov network | 88.2 | 81.4 | 84.2 |
The FM algorithm and atom candidate mechanism are both critical (accuracy drops below 10% without either). Bond candidates provide a moderate improvement (~15 percentage points), and the Markov network adds ~6 points over hard-threshold alternatives.
Limitations
- Deterministic expert rules limit robustness on perturbed or noisy images, as evidenced by the large accuracy gap with MolGrapher on USPTO-perturbed
- The system relies on Tesseract OCR for symbol recognition, which may struggle with unusual fonts or degraded image quality
- Only handles single 2D molecule structures per image
- Stereochemistry is removed during evaluation, so performance on stereo-bond recognition is not assessed
Reproducibility
GraphReco is implemented in Python and relies on Tesseract OCR, OpenCV, and RDKit. The authors provide an online demo for testing but have not released the source code or a public repository.
| Artifact | Type | License | Notes |
|---|---|---|---|
| Online Demo | Other | Unknown | Google Cloud Run deployment (no longer available) |
Missing components for full reproduction:
- Source code is not publicly available
- No pre-built package or installable library
- Hyperparameters for Markov network potential functions are given in the paper (Equations 8-11), but full implementation details are not released
Hardware/compute requirements: Not specified in the paper. The system uses classical computer vision (Hough transforms, thinning) and probabilistic inference (Markov networks), so GPU hardware is likely not required.