A Multimodal Method for Markush Structure Recognition
This is a Method paper that introduces MarkushGrapher-2, a universal encoder-decoder model for recognizing both standard molecular structures and multimodal Markush structures from chemical images. The primary contribution is a dual-encoder architecture that fuses a pretrained OCSR (Optical Chemical Structure Recognition) vision encoder with a Vision-Text-Layout (VTL) encoder, connected through a dedicated ChemicalOCR module for end-to-end processing. The paper also introduces two new resources: a large-scale training dataset (USPTO-MOL-M) of real-world Markush structures extracted from USPTO patent MOL files, and IP5-M, a manually annotated benchmark of 1,000 Markush structures from five major patent offices.
Why Markush Structure Recognition Remains Challenging
Markush structures are compact representations used in patent documents to describe families of related molecules. They combine a visual backbone (atoms, bonds, variable regions) with textual definitions of substituents that can replace those variable regions. This multimodal nature makes them harder to parse than standard molecular diagrams.
Three factors limit automatic Markush recognition. First, visual styles vary across patent offices and publication years. Second, textual definitions lack standardization and often contain conditional or recursive descriptions. Third, real-world training data with comprehensive annotations is scarce. As a result, Markush structures are currently indexed only in two proprietary, manually curated databases: MARPAT and DWPIM.
Prior work, including the original MarkushGrapher, required pre-annotated OCR outputs at inference time, limiting practical deployment. General-purpose models like GPT-5 and DeepSeek-OCR produce mostly chemically invalid outputs on Markush images, suggesting these lie outside their training distribution.
Dual-Encoder Architecture with Dedicated ChemicalOCR
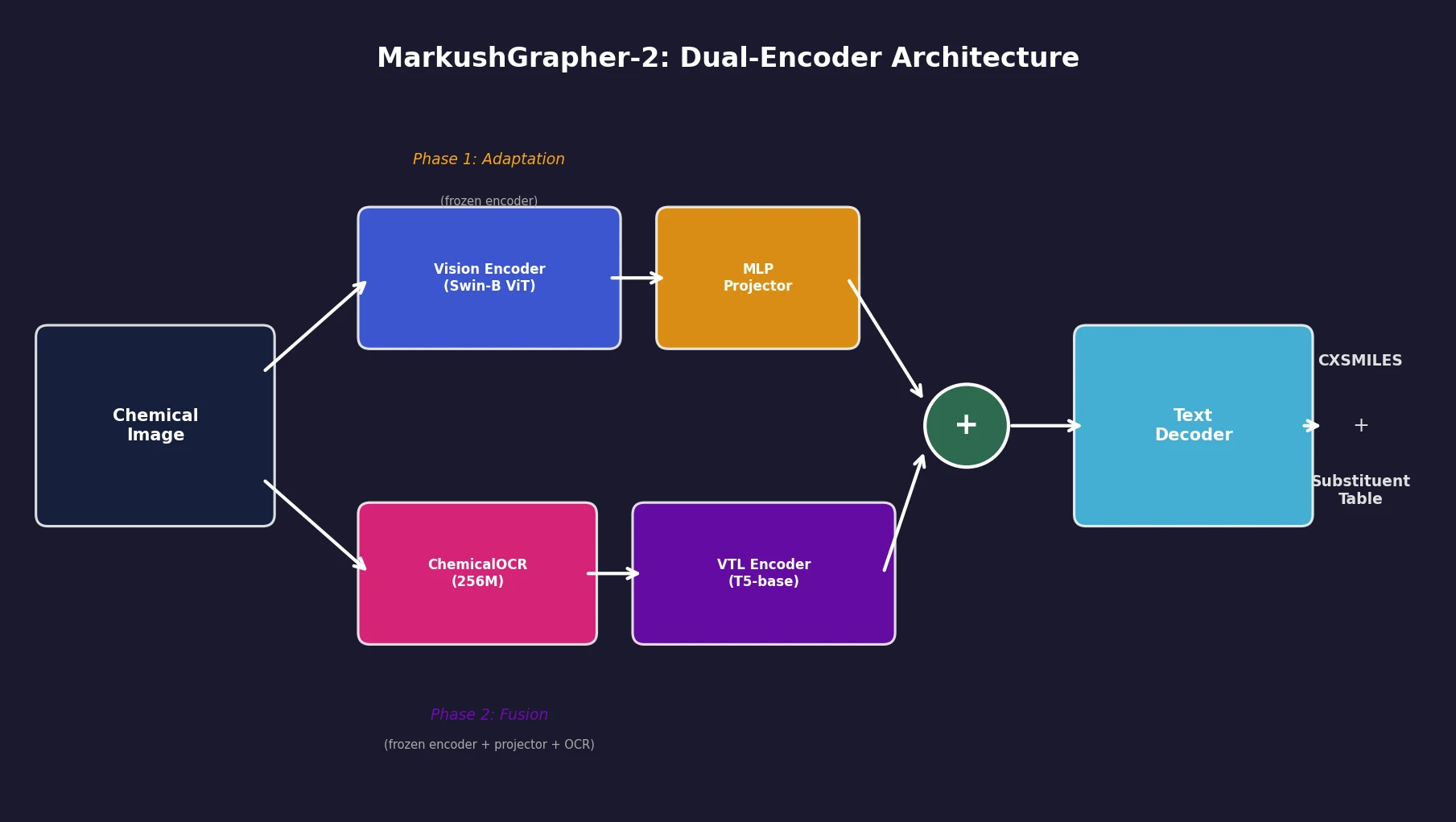
MarkushGrapher-2 uses two complementary encoding pipelines:
Vision encoder pipeline: The input image passes through a Swin-B Vision Transformer (taken from MolScribe) pretrained for OCSR. This encoder extracts visual features representing molecular structures and remains frozen during training.
Vision-Text-Layout (VTL) pipeline: The same image goes through ChemicalOCR, a compact 256M-parameter vision-language model fine-tuned from SmolDocling for OCR on chemical images. ChemicalOCR extracts character-level text and bounding boxes. These, combined with image patches, feed into a T5-base VTL encoder following the UDOP fusion paradigm, where visual and textual tokens are spatially aligned by bounding box overlap.
The VTL encoder output is concatenated with projected embeddings from the vision encoder. This joint representation feeds a text decoder that auto-regressively generates a CXSMILES (ChemAxon Extended SMILES) string describing the backbone structure and a substituent table listing variable group definitions.
Two-Stage Training Strategy
Training proceeds in two phases:
Phase 1 (Adaptation): The vision encoder is frozen. The MLP projector and text decoder train on 243K real-world image-SMILES pairs from MolScribe’s USPTO dataset (3 epochs). This aligns the decoder to the pretrained OCSR feature space.
Phase 2 (Fusion): The vision encoder, projector, and ChemicalOCR are all frozen. The VTL encoder and text decoder train on a mix of 235K synthetic and 145K real-world Markush samples (2 epochs). The VTL encoder learns the features needed for CXSMILES and substituent table prediction without disrupting the established OCSR representations.
The total model has 831M parameters, of which 744M are trainable.
Datasets and Evaluation Benchmarks
Training Data
| Purpose | Dataset | Size | Source |
|---|---|---|---|
| OCR pretraining | Synthetic chemical structures | 235K | PubChem SMILES augmented to CXSMILES, rendered with annotations |
| OCR fine-tuning | Manual OCR annotations | 7K | IP5 patent document crops |
| Phase 1 (OCSR) | MolScribe USPTO | 243K | Real image-SMILES pairs |
| Phase 2 (MMSR) | Synthetic CXSMILES | 235K | Same as OCR pretraining set |
| Phase 2 (MMSR) | MolParser dataset | 91K | Real-world Markush, converted to CXSMILES |
| Phase 2 (MMSR) | USPTO-MOL-M | 54K | Real-world, auto-extracted from USPTO MOL files (2010-2025) |
Evaluation Benchmarks
Markush benchmarks: M2S (103 samples), USPTO-M (74), WildMol-M (10K, semi-manual), and the new IP5-M (1,000 manually annotated from USPTO, JPO, KIPO, CNIPA, and EPO patents, 1980-2025).
OCSR benchmarks: USPTO (5,719), JPO (450), UOB (5,740), WildMol (10K).
The primary metric is CXSMILES Accuracy (A): a prediction is correct when (1) the predicted SMILES matches the ground truth by InChIKey equivalence, and (2) all Markush features (variable groups, positional and frequency variation indicators) are correctly represented. Stereochemistry is ignored during evaluation.
Results: Markush Structure Recognition
| Model | M2S | USPTO-M | WildMol-M | IP5-M |
|---|---|---|---|---|
| MolParser-Base | 39 | 30 | 38.1 | 47.7 |
| MolScribe | 21 | 7 | 28.1 | 22.3 |
| GPT-5 | 3 | 0 | - | - |
| DeepSeek-OCR | 0 | 0 | 1.9 | 0.0 |
| MarkushGrapher-1 | 38 | 10 | 32 | - |
| MarkushGrapher-2 | 56 | 13 | 55 | 48.0 |
On M2S, MarkushGrapher-2 achieves 56% CXSMILES accuracy vs. 38% for MarkushGrapher-1, a relative improvement of 47%. On WildMol-M (the largest benchmark at 10K samples), MarkushGrapher-2 reaches 55% vs. 38.1% for MolParser-Base and 32% for MarkushGrapher-1. GPT-5 and DeepSeek-OCR generate mostly chemically invalid outputs on Markush images: only 30% and 15% of their predictions are valid CXSMILES on M2S, respectively.
Results: Standard Molecular Structure Recognition
| Model | WildMol | JPO | UOB | USPTO |
|---|---|---|---|---|
| MolParser-Base | 76.9 | 78.9 | 91.8 | 93.0 |
| MolScribe | 66.4 | 76.2 | 87.4 | 93.1 |
| MolGrapher | 45.5 | 67.5 | 94.9 | 91.5 |
| DeepSeek-OCR | 25.8 | 31.6 | 78.7 | 36.9 |
| MarkushGrapher-2 | 68.4 | 71.0 | 96.6 | 89.8 |
MarkushGrapher-2 achieves the highest score on UOB (96.6%) and remains competitive on other OCSR benchmarks, despite being primarily optimized for Markush recognition.
ChemicalOCR vs. General OCR
| Model | M2S F1 | USPTO-M F1 | IP5-M F1 |
|---|---|---|---|
| PaddleOCR v5 | 7.7 | 1.2 | 1.9 |
| EasyOCR | 10.2 | 18.0 | 18.4 |
| ChemicalOCR | 87.2 | 93.0 | 86.5 |
General-purpose OCR tools fail on chemical images because they misinterpret bonds as characters and cannot parse chemical abbreviations. ChemicalOCR outperforms both by a large margin.
Ablation Results and Key Findings
OCR input is critical for Markush features. Without OCR, CXSMILES accuracy drops from 56% to 4% on M2S, and from 53.7% to 15.4% on IP5-M. The backbone structure accuracy ($A_{\text{InChIKey}}$) also drops substantially (from 80% to 39% on M2S), though the vision encoder alone can still recover some structural information. This confirms that textual cues (brackets, indices, variable definitions) are essential for Markush feature prediction.
Two-phase training improves both tasks. Compared to single-phase (fusion only) training, the two-phase strategy improves CXSMILES accuracy from 44% to 50% on M2S and from 53.0% to 61.5% on JPO after the same number of epochs. Adapting the decoder to OCSR features before introducing the VTL encoder prevents the fusion process from degrading learned visual representations.
Frequency variation indicators remain the hardest feature. On IP5-M, the per-feature breakdown shows 73.3% accuracy for backbone InChI, 74.8% for variable groups, 78.8% for positional variation, but only 30.7% for frequency variation (Sg groups). These repeating structural units are particularly challenging to represent and predict.
Limitations: The model relies on accurate OCR as a prerequisite. Performance on USPTO-M (13% CXSMILES accuracy) lags behind other benchmarks, likely due to the older patent styles in that dataset. The paper does not report inference latency.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| OCR pretraining | Synthetic chemical images | 235K | Generated from PubChem SMILES, augmented to CXSMILES |
| OCR fine-tuning | IP5 patent crops | 7K | Manually annotated |
| Phase 1 training | MolScribe USPTO | 243K | Public, real image-SMILES pairs |
| Phase 2 training | Synthetic + MolParser + USPTO-MOL-M | 380K | Mix of synthetic (235K), MolParser (91K), USPTO-MOL-M (54K) |
| Evaluation | M2S, USPTO-M, WildMol-M, IP5-M | 103 to 10K | Markush benchmarks |
| Evaluation | WildMol, JPO, UOB, USPTO | 450 to 10K | OCSR benchmarks |
Models
| Component | Architecture | Parameters | Status |
|---|---|---|---|
| Vision encoder | Swin-B ViT (from MolScribe) | ~87M | Frozen |
| VTL encoder + decoder | T5-base | ~744M trainable | Trained |
| ChemicalOCR | SmolDocling-based VLM | 256M | Fine-tuned, frozen in Phase 2 |
| MLP projector | Linear projection | - | Trained in Phase 1, frozen in Phase 2 |
| Total | 831M |
Evaluation
| Metric | Definition |
|---|---|
| CXSMILES Accuracy (A) | Percentage of samples where InChIKey matches AND all Markush features correct |
| $A_{\text{InChIKey}}$ | Backbone structure accuracy only (ignoring Markush features) |
| Table Accuracy | Percentage of correctly predicted substituent tables |
| Markush Accuracy | Joint CXSMILES + Table accuracy |
| OCR F1 | Bounding-box-level precision/recall at IoU > 0.5 |
Hardware
- Training: NVIDIA A100 GPU
- Phase 1: 3 epochs, Adam optimizer, lr 5e-4, 1000 warmup steps, batch size 10, weight decay 1e-3
- Phase 2: 2 epochs, batch size 8
Paper Information
Citation: Strohmeyer, T., Morin, L., Meijer, G. I., Weber, V., Nassar, A., & Staar, P. (2026). MarkushGrapher-2: End-to-end Multimodal Recognition of Chemical Structures. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR).
Publication: CVPR 2026
Additional Resources:
@misc{strohmeyer2026markushgrapher,
title={MarkushGrapher-2: End-to-end Multimodal Recognition of Chemical Structures},
author={Strohmeyer, Tim and Morin, Lucas and Meijer, Gerhard Ingmar and Weber, Val\'{e}ry and Nassar, Ahmed and Staar, Peter},
year={2026},
eprint={2603.28550},
archiveprefix={arXiv},
primaryclass={cs.CV}
}