Paper Information
Citation: Morin, L., Weber, V., Nassar, A., Meijer, G. I., Van Gool, L., Li, Y., & Staar, P. (2025). MarkushGrapher: Joint Visual and Textual Recognition of Markush Structures. 2025 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 14505-14515. https://doi.org/10.1109/CVPR52734.2025.01352
Publication: CVPR 2025
Additional Resources:
Overcoming Unimodal Limitations for Markush Structures
The automated analysis of chemical literature, particularly patents, is critical for drug discovery and material science. A major bottleneck is the extraction of Markush structures, which are complex chemical templates that represent families of molecules using a core backbone image and textual variable definitions. Existing methods are limited because they either rely solely on images (OCSR) and miss the textual context, or focus solely on text and miss the structural backbone. This creates a practical need for a unified, multi-modal approach that jointly interprets visual and textual data to accurately extract these structures for prior-art search and database construction. This paper proposes a Method and introduces a new Resource (M2S dataset) to bridge this gap.
MarkushGrapher: The Multi-Modal Architecture
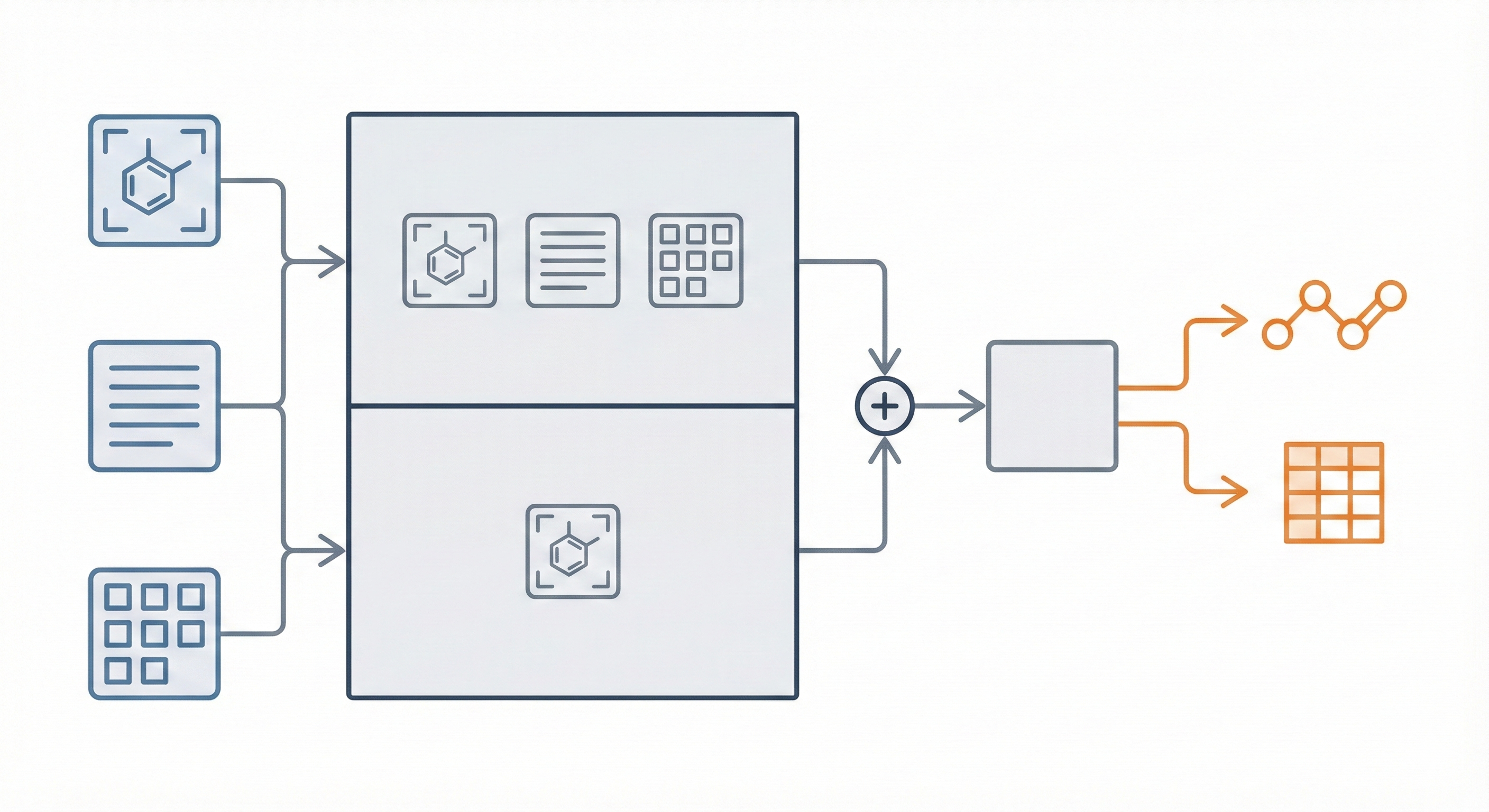
The core innovation is MarkushGrapher, a multi-modal architecture that jointly encodes image, text, and layout information. Key contributions include:
- Dual-Encoder Architecture: Combines a Vision-Text-Layout (VTL) encoder (based on UDOP) with a specialized, pre-trained Optical Chemical Structure Recognition (OCSR) encoder (MolScribe). Let $E_{\text{VTL}}$ represent the combined sequence embedding and $E_{\text{OCSR}}$ represent the domain-specific visual embeddings.
- Joint Recognition: The model autoregressively generates a sequential graph representation (Optimized CXSMILES) and a substituent table simultaneously. It uses cross-modal dependencies, allowing text to clarify ambiguous visual details like bond types.
- Synthetic Data Pipeline: A comprehensive pipeline generates realistic synthetic Markush structures (images and text) from PubChem data, overcoming the lack of labeled training data.
- Optimized Representation: A compacted version of CXSMILES moves variable groups into the SMILES string and adds explicit atom indexing to handle complex “frequency” and “position” variation indicators.
Experimental Validation on the New M2S Benchmark
The authors validated their approach using the following setup:
- Baselines: Compared against image-only chemistry models (DECIMER, MolScribe) and general-purpose multi-modal models (Uni-SMART, GPT-4o, Pixtral, Llama-3.2).
- Datasets: Evaluated on three benchmarks:
- MarkushGrapher-Synthetic: 1,000 generated samples.
- M2S: A new benchmark of 103 manually annotated real-world patent images.
- USPTO-Markush: 74 Markush backbone images from USPTO patents.
- Ablation Studies: Analyzed the impact of the OCSR encoder, late fusion strategies, and the optimized CXSMILES format. Late fusion improved USPTO-Markush EM from 23% (VTL only) to 32% (Table 3). Removing R-group compression dropped M2S EM from 38% to 30%, and removing atom indexing dropped USPTO-Markush EM from 32% to 24% (Table 4).
Key Results
- Performance: MarkushGrapher outperformed all baselines. On the M2S benchmark, it achieved 38% Exact Match on CXSMILES (compared to 21% for MolScribe) and 29% Exact Match on tables. On USPTO-Markush, it reached 32% CXSMILES EM versus 7% for MolScribe.
- Markush Feature Recognition: The model can recognize complex Markush features like frequency variation (‘Sg’) and position variation (’m’) indicators. DECIMER and MolScribe scored 0% on both ’m’ and ‘Sg’ sections (Table 2), while MarkushGrapher achieved 76% on ’m’ and 31% on ‘Sg’ sections on M2S.
- Cross-Modal Reasoning: Qualitative analysis showed the model can correctly infer visual details (such as bond order) that appear ambiguous in the image but become apparent with the text description.
- Robustness: The model generalizes well to real-world data despite being trained purely on synthetic data. On augmented versions of M2S and USPTO-Markush simulating low-quality scanned documents, it maintained 31% and 32% CXSMILES EM respectively (Table 6).
Limitations
The authors note several limitations:
- MarkushGrapher does not currently handle abbreviations in chemical structures (e.g., ‘OG’ for oxygen connected to a variable group).
- The model relies on ground-truth OCR cells as input, requiring an external OCR model for practical deployment.
- Substituent definitions that combine text with interleaved chemical structure drawings are not supported.
- The model is trained to predict ’m’ sections connecting to all atoms in a cycle, which can technically violate valence constraints, though the output contains enough information to reconstruct only valid connections.
Reproducibility Details
Data
Training Data
- Source: Synthetic dataset generated from PubChem SMILES.
- Size: 210,000 synthetic images.
- Pipeline:
- Selection: Sampled SMILES from PubChem based on substructure diversity.
- Augmentation: SMILES augmented to artificial CXSMILES using RDKit (inserting variable groups, frequency indicators).
- Rendering: Images rendered using Chemistry Development Kit (CDK) with randomized drawing parameters (font, bond width, spacing).
- Text Generation: Textual definitions generated using manual templates extracted from patents; 10% were paraphrased using Mistral-7B-Instruct-v0.3 to increase diversity.
- OCR: Bounding boxes extracted via a custom SVG parser aligned with MOL files.
Evaluation Data
- M2S Dataset: 103 images from USPTO, EPO, and WIPO patents (1999-2023), manually annotated with CXSMILES and substituent tables.
- USPTO-Markush: 74 images from USPTO patents (2010-2016).
- MarkushGrapher-Synthetic: 1,000 samples generated via the pipeline.
Algorithms
- Optimized CXSMILES:
- Compression: Variable groups moved from the extension block to the main SMILES string as special atoms to reduce sequence length.
- Indexing: Atom indices appended to each atom (e.g.,
C:1) to explicitly link the graph to the extension block (crucial formandSgsections). - Vocabulary: Specific tokens used for atoms and bonds.
- Augmentation: Standard image augmentations (shift, scale, blur, pepper noise, random lines) and OCR text augmentations (character substitution/insertion/deletion).
Models
- Architecture: Encoder-Decoder Transformer.
- VTL Encoder: T5-large encoder (initialized from UDOP) that processes image patches, text tokens, and layout (bounding boxes).
- OCSR Encoder: Vision encoder from MolScribe (Swin Transformer), frozen during training.
- Text Decoder: T5-large decoder.
- Fusion Strategy: Late Fusion. The core multi-modal alignment combines the textual layout features with specialized chemical vision explicitly. The fused representation relies on the VTL output $e_1$ concatenated with the MLP-projected OCSR output $e_2$ before decoding: $$ e = e_1(v, t, l) \oplus \text{MLP}(e_2(v)) $$
- Parameters: 831M total (744M trainable).
Evaluation
Metrics:
- CXSMILES Exact Match (EM): Requires perfect match of SMILES string, variable groups,
msections, andSgsections (ignoring stereochemistry). - Tanimoto Score: Similarity of RDKit DayLight fingerprints (Markush features removed).
- Table Exact Match: All variable groups and substituents must match.
- Table F1-Score: Aggregated recall and precision of substituents per variable group.
Hardware
- Compute: Trained on a single NVIDIA H100 GPU.
- Training Config: 10 epochs, batch size of 10, ADAM optimizer, learning rate 5e-4, 100 warmup steps, weight decay 1e-3.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| MarkushGrapher | Code | MIT | Official implementation |
Citation
@inproceedings{morinMarkushGrapherJointVisual2025,
title = {MarkushGrapher: Joint Visual and Textual Recognition of Markush Structures},
shorttitle = {MarkushGrapher},
booktitle = {2025 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR)},
author = {Morin, Lucas and Weber, Valéry and Nassar, Ahmed and Meijer, Gerhard Ingmar and Van Gool, Luc and Li, Yawei and Staar, Peter},
year = {2025},
month = jun,
pages = {14505--14515},
doi = {10.1109/CVPR52734.2025.01352}
}