Paper Information
Citation: Zhang, D., Zhao, D., Wang, Z., Li, J., & Li, J. (2024). MMSSC-Net: multi-stage sequence cognitive networks for drug molecule recognition. RSC Advances, 14(26), 18182-18191. https://doi.org/10.1039/D4RA02442G
Publication: RSC Advances 2024
Contribution: A Multi-Stage Architectural Pipeline
Methodological Paper ($\Psi_{\text{Method}}$). The paper proposes a deep learning architecture (MMSSC-Net) for Optical Chemical Structure Recognition (OCSR). It focuses on architectural innovation, specifically combining a SwinV2 visual encoder with a GPT-2 decoder, and validates this method through extensive benchmarking against existing rule-based and deep-learning baselines. It includes ablation studies to justify the choice of the visual encoder.
Motivation: Addressing Noise and Rigid Image Recognition
- Data Usage Gap: Drug discovery relies heavily on scientific literature, but molecular structures are often locked in vector graphics or images that computers cannot easily process.
- Limitations of Prior Work: Existing Rule-based methods are rigid and sensitive to noise. Previous Deep Learning approaches (Encoder-Decoder “Image Captioning” styles) often lack precision, interpretability, and struggle with varying image resolutions or large molecules.
- Need for “Cognition”: The authors argue that treating the image as a single isolated whole is insufficient; a model needs to “perceive” fine-grained details (atoms and bonds) to handle noise and varying pixel qualities effectively.
Novelty: A Fine-Grained Perception Pipeline
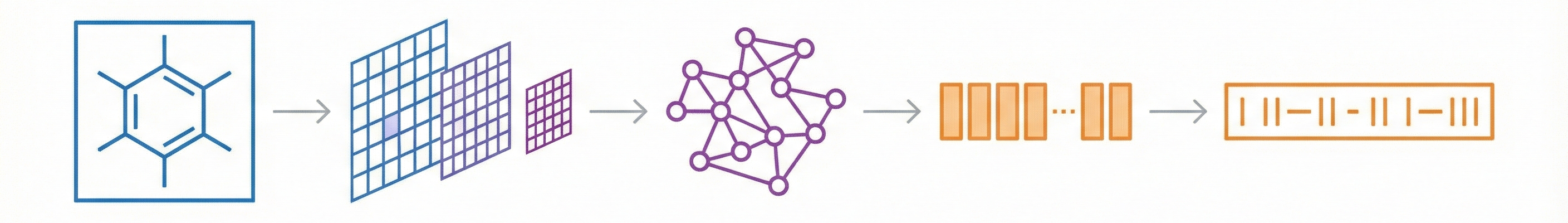
- Multi-Stage Cognitive Architecture: MMSSC-Net splits the task into stages:
- Fine-grained Perception: Detecting atom and bond sequences (including spatial coordinates) using SwinV2.
- Graph Construction: Assembling these into a molecular graph.
- Sequence Evolution: converting the graph into a machine-readable format (SMILES).
- Hybrid Transformer Model: It combines a hierarchical vision transformer (SwinV2) for encoding with a generative pre-trained transformer (GPT-2) and MLPs for decoding atomic and bond targets.
- Robustness Mechanisms: The inclusion of random noise sequences during training to improve generalization to new molecular targets.
Methodology and Benchmarks
- Baselines: compared against 8 other tools:
- Rule-based: MolVec, OSRA.
- Image-Smiles (DL): ABC-Net, Img2Mol, MolMiner.
- Image-Graph-Smiles (DL): Image-To-Graph, MolScribe, ChemGrapher.
- Datasets: Evaluated on 5 diverse datasets: STAKER (synthetic), USPTO, CLEF, JPO, and UOB (real-world).
- Metrics:
- Accuracy: Exact string match of the predicted SMILES.
- Tanimoto Similarity: Chemical similarity using Morgan fingerprints.
- Ablation Study: Tested different visual encoders (Swin Transformer, ViT-B, ResNet-50) to validate the choice of SwinV2.
- Resolution Sensitivity: Tested model performance across image resolutions from 256px to 2048px.
Results and Core Outcomes
- Strong Performance: MMSSC-Net achieved 75-98% accuracy across datasets, outperforming baselines on most benchmarks. The first three intra-domain and real datasets achieved above 94% accuracy.
- Resolution Robustness: The model maintained relatively stable accuracy across varying image resolutions, whereas baselines like Img2Mol showed greater sensitivity to resolution changes (Fig. 4 in the paper).
- Efficiency: The SwinV2 encoder was noted to be more efficient than ViT-B in this context.
- Limitations: The model struggles with stereochemistry, specifically confusing dashed wedge bonds with solid wedge bonds and misclassifying single bonds as solid wedge bonds. It also has difficulty with “irrelevant text” noise (e.g., unexpected symbols in JPO and DECIMER datasets).
Reproducibility Details
Data
The model was trained on a combination of PubChem and USPTO data, augmented to handle visual variability.
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training | PubChem | 1,000,000 | Converted from InChI to SMILES; random sampling. |
| Training | USPTO | 600,000 | Patent images; converted from MOL to SMILES. |
| Evaluation | STAKER | 40,000 | Synthetic; Avg res $256 \times 256$. |
| Evaluation | USPTO | 4,862 | Real; Avg res $721 \times 432$. |
| Evaluation | CLEF | 881 | Real; Avg res $1245 \times 412$. |
| Evaluation | JPO | 380 | Real; Avg res $614 \times 367$. |
| Evaluation | UOB | 5,720 | Real; Avg res $759 \times 416$. |
Augmentation:
- Image: Random perturbations using RDKit/Indigo (rotation, filling, cropping, bond thickness/length, font size, Gaussian noise).
- Molecular: Introduction of functional group abbreviations and R-substituents (dummy atoms) using SMARTS templates.
Algorithms
- Target Sequence Formulation: The model predicts a sequence containing bounding box coordinates and type labels: ${y_{\text{min}}, x_{\text{min}}, y_{\text{max}}, x_{\text{max}}, C_{n}}$.
- Loss Function: Cross-entropy loss with maximum likelihood estimation. $$ \max \sum_{i=1}^{N} \sum_{j=1}^{L} \omega_{j} \log P(t_{j}^{i} \mid x_{1}^{i}, x_{2}^{i}, \dots, x_{M}^{i}, t_{1}^{i}, \dots, t_{j-1}^{i}) $$
- Noise Injection: A random sequence $T_r$ is appended to the target sequence during training to improve generalization to new goals.
- Graph Construction: Atoms ($v$) and bonds ($e$) are recognized separately; bonds are defined by connecting spatial atomic coordinates.
Models
- Encoder: Swin Transformer V2.
- Pre-trained on ImageNet-1K.
- Window size: $16 \times 16$.
- Parameters: 88M.
- Input resolution: $256 \times 256$.
- Features: Scaled cosine attention; log-space continuous position bias.
- Decoder: GPT-2 + MLP.
- GPT-2: Used for recognizing atom types.
- Layers: 24.
- Attention Heads: 12.
- Hidden Dimension: 768.
- Dropout: 0.1.
- MLP: Used for classifying bond types (single, double, triple, aromatic, solid wedge, dashed wedge).
- GPT-2: Used for recognizing atom types.
- Vocabulary:
- Standard: 95 common numbers/characters ([0], [C], [=], etc.).
- Extended: 2000 SMARTS-based characters for isomers/groups (e.g., “[C2F5]”, “[halo]”).
Evaluation
Metrics:
- Accuracy: Exact match of the generated SMILES string.
- Tanimoto Similarity: Similarity of Morgan fingerprints between predicted and ground truth molecules.
Key Results (Accuracy):
| Dataset | MMSSC-Net | MolVec (Rule) | ABC-Net (DL) | MolScribe (DL) |
|---|---|---|---|---|
| Indigo | 98.14 | 95.63 | 96.4 | 97.5 |
| RDKit | 94.91 | 86.7 | 98.3 | 93.8 |
| USPTO | 94.24 | 88.47 | * | 92.6 |
| CLEF | 91.26 | 81.61 | * | 86.9 |
| UOB | 92.71 | 81.32 | 96.1 | 87.9 |
| Staker | 89.44 | 4.49 | * | 86.9 |
| JPO | 75.48 | 66.8 | * | 76.2 |
Hardware
- Training Configuration:
- Batch Size: 128.
- Learning Rate: $4 \times 10^{-5}$.
- Epochs: 40.
- Inference Speed: The SwinV2 encoder demonstrated higher efficiency (faster inference time) compared to ViT-B and ResNet-50 baselines during ablation.
Reproducibility
| Artifact | Type | License | Notes |
|---|---|---|---|
| MMSSCNet (GitHub) | Code | Unknown | Official implementation; includes training and prediction scripts |
The paper is published in RSC Advances (open access). Source code is available on GitHub, though the repository has minimal documentation and no explicit license. The training data comes from PubChem (public) and USPTO (public patent data). Pre-trained model weights do not appear to be released. No specific GPU hardware or training time is reported in the paper.
Citation
@article{zhangMMSSCNetMultistageSequence2024,
title = {MMSSC-Net: Multi-Stage Sequence Cognitive Networks for Drug Molecule Recognition},
shorttitle = {MMSSC-Net},
author = {Zhang, Dehai and Zhao, Di and Wang, Zhengwu and Li, Junhui and Li, Jin},
year = 2024,
journal = {RSC Advances},
volume = {14},
number = {26},
pages = {18182--18191},
publisher = {Royal Society of Chemistry},
doi = {10.1039/D4RA02442G},
url = {https://pubs.rsc.org/en/content/articlelanding/2024/ra/d4ra02442g}
}