Contribution: Early OCSR Pipeline Methodology
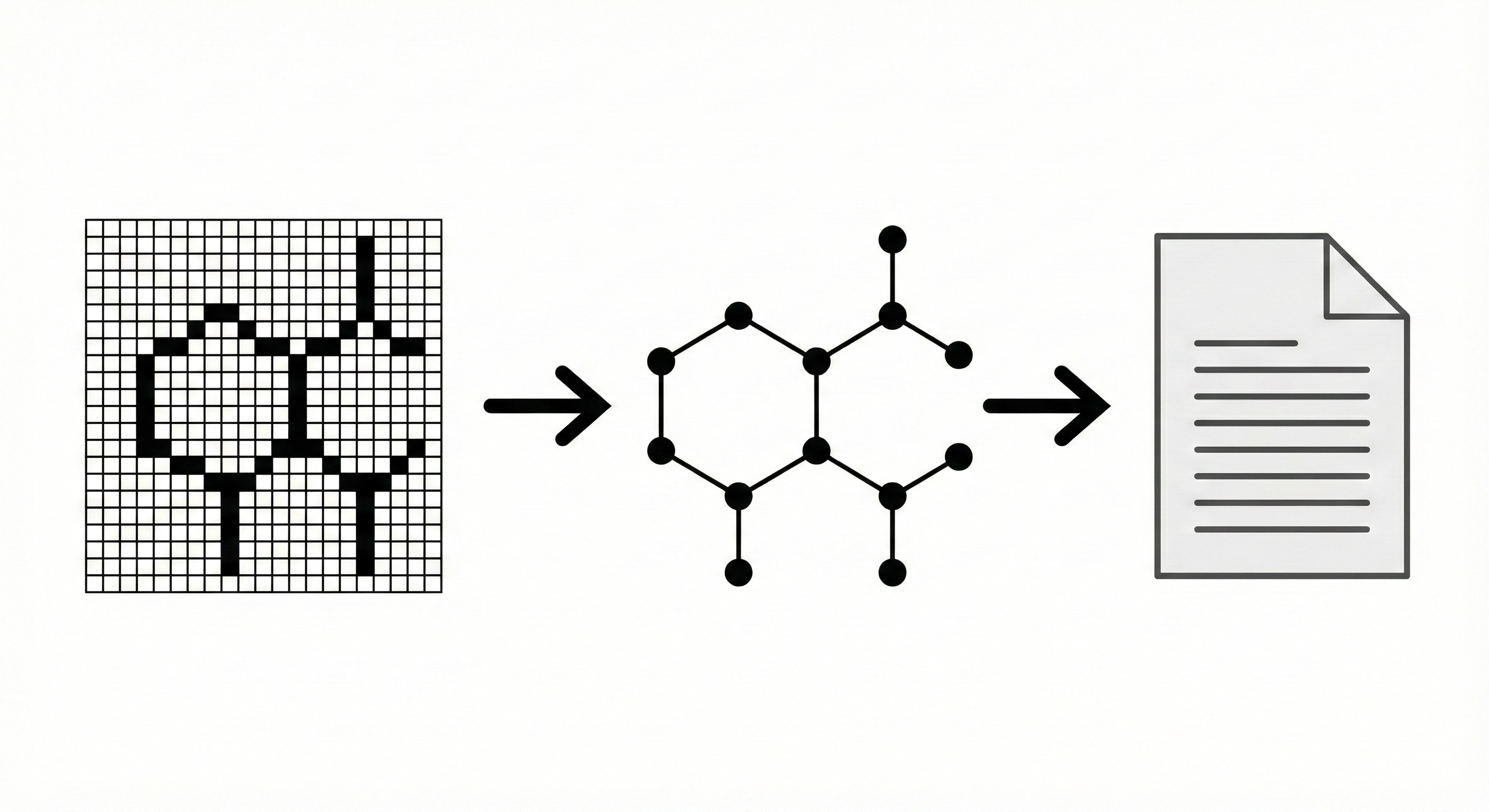
Method. This paper proposes a novel architectural pipeline for the automatic recognition of chemical structure diagrams. It defines a specific sequence of algorithmic steps, including diagram separation, vectorization, segmentation, and structural analysis, which converts pixel data into a semantic chemical representation (MDL Molfile).
Motivation: Digitizing Legacy Chemical Data
Problem: In 1993, vast databases of chemical information existed, but the entry of graphical data was significantly less advanced than the facilities for manipulating it.
Gap: Creating digital chemical structures required trained operators to manually redraw diagrams that already existed in printed journals and catalogs, leading to a costly duplication of effort.
Goal: To automate the creation of coded representations (connection tables) directly from optically scanned diagrams on printed pages.
Novelty: General Document Analysis Integrated with Chemical Rules
Pipeline Approach: The authors present a complete end-to-end system that integrates general document analysis with domain-specific chemical rules.
Convex Bounding Separation: A novel use of “bounding polygons” defined by 8 fixed-direction bands to distinguish diagram components from text with linear computational cost.
Vector-Based Segmentation: The system uses the output of a vectorizer (GIFTS) to classify diagram elements. It relies on the observation that vectorizers approximate characters with sets of short vectors to distinguish them from bonds.
Methodology and System Evaluation
System Implementation: The algorithm was implemented in ‘C’ on IBM PS/2 personal computers running OS/2 Presentation Manager.
Input Specification: The system was tested on documents scanned at 300 dpi using an IBM 3119 scanner.
Qualitative Evaluation: The authors evaluated the system on “typical scanned structures” and “simple planar diagrams”. Large-scale quantitative benchmarking was not conducted in this work.
Results, Performance, and Limitations
Performance: The prototype processes a typical structure (after extraction) in less than one minute.
Accuracy: It is reported to be accurate for simple planar diagrams.
Output Format: The system successfully generates MDL Molfiles that interface with standard chemistry software like REACCS, MACCS, and modeling tools.
Limitations: The system struggles with broken lines, characters touching bond structures, and requires manual intervention for complex errors.
Reproducibility Details
Status: Closed (Historical). As an early prototype from 1993, no source code, datasets, or digital models were publicly released. Reproducing this exact system would require recreating the pipeline from the described heuristics and sourcing vintage OCR software.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| None available | N/A | N/A | No digital artifacts were released with this 1993 publication. |
Data
The paper does not release a dataset but specifies the input requirements for the system.
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Input | Scanned Documents | N/A | Black ink on white paper; scanned at 300 dpi bi-level. |
Algorithms
The paper relies on a pipeline of specific heuristics and geometric rules.
1. Diagram Separation (Region Growing)
- Bounding Polygons: Uses convex polygons defined by pairs of parallel sides in 8 fixed directions. This approximation improves distance estimation compared to bounding rectangles.
- Seed Detection: Finds a connected component with bounding dimension $D > d_{\text{max\_char\_size}}$.
- Aggregation: Iteratively searches for neighboring components within a specific distance threshold $d_t$ (where $d_t$ is smaller than the whitespace margin) and merges them into the bounding polygon.
2. Vectorization & Segmentation
- Vectorization: Uses the GIFTS system (IBM Tokyo) to fit lines to pixels.
- Classification Heuristics:
- Ratio Test: If the ratio of a group’s dimension to the full diagram dimension is below a threshold $\tau$, it is classified as a Symbol: $$ \frac{D_{\text{group}}}{D_{\text{diagram}}} < \tau $$
- Context Rule: Small vector groups near letters are classified as Characters (handles ’l’ in ‘Cl’).
- Circle Rule: A group is a Circle (aromatic ring) if it contains $N \ge 8$ vectors in a roughly circular arrangement.
- Default: Otherwise, classified as Bond Structure.
3. Cleanup & Structure Recognition
- Short Vector Removal: Vectors shorter than a fraction of the median line length $L_{\text{median}}$ are shrunk to their midpoint (fixing broken junctions).
- Vertex Merging: If two vectors meet at an angle $\theta < 35^{\circ}$, the vertex is removed (fixing single lines broken into two).
- Aromatic Processing: If a circle is detected, the system identifies the 6 closest atoms and adds double bonds to every second bond in the ring.
Models
OCR:
- The system uses a feature-based, single-font OCR engine.
- It assumes non-serif, plain styles typical of drafting standards.
- Character images are normalized for size before recognition.
Hardware
- Scanner: IBM 3119 (300 dpi).
- Compute: IBM PS/2 series running OS/2.
Paper Information
Citation: Casey, R., et al. (1993). Optical Recognition of Chemical Graphics. Proceedings of 2nd International Conference on Document Analysis and Recognition (ICDAR ‘93), 627-631. https://doi.org/10.1109/ICDAR.1993.395658
Publication: ICDAR 1993
@inproceedings{caseyOpticalRecognitionChemical1993,
title = {Optical Recognition of Chemical Graphics},
booktitle = {Proceedings of 2nd {{International Conference}} on {{Document Analysis}} and {{Recognition}} ({{ICDAR}} '93)},
author = {Casey, R. and Boyer, S. and Healey, P. and Miller, A. and Oudot, B. and Zilles, K.},
year = 1993,
pages = {627--631},
publisher = {IEEE Comput. Soc. Press},
address = {Tsukuba Science City, Japan},
doi = {10.1109/ICDAR.1993.395658}
}