Paper Information
Citation: Fang, X., Tao, H., Yang, S., Huang, C., Zhong, S., Lu, H., Lyu, H., Li, X., Zhang, L., & Ke, G. (2025). Uni-Parser Technical Report. arXiv preprint arXiv:2512.15098. https://arxiv.org/abs/2512.15098
Publication: arXiv 2025
Additional Resources:
An Industrial-Grade Multi-Modal Document Parser
Uni-Parser is a modular, loosely coupled PDF parsing engine built for scientific literature and patents. It routes different content types (text, equations, tables, figures, chemical structures) to specialized expert models, then reassembles the parsed outputs into structured formats (JSON, Markdown, HTML) for downstream consumption by LLMs and other applications.
The system processes up to 20 PDF pages per second on 8 NVIDIA RTX 4090D GPUs and supports over 80 languages for OCR.
A Five-Stage Pipeline Architecture
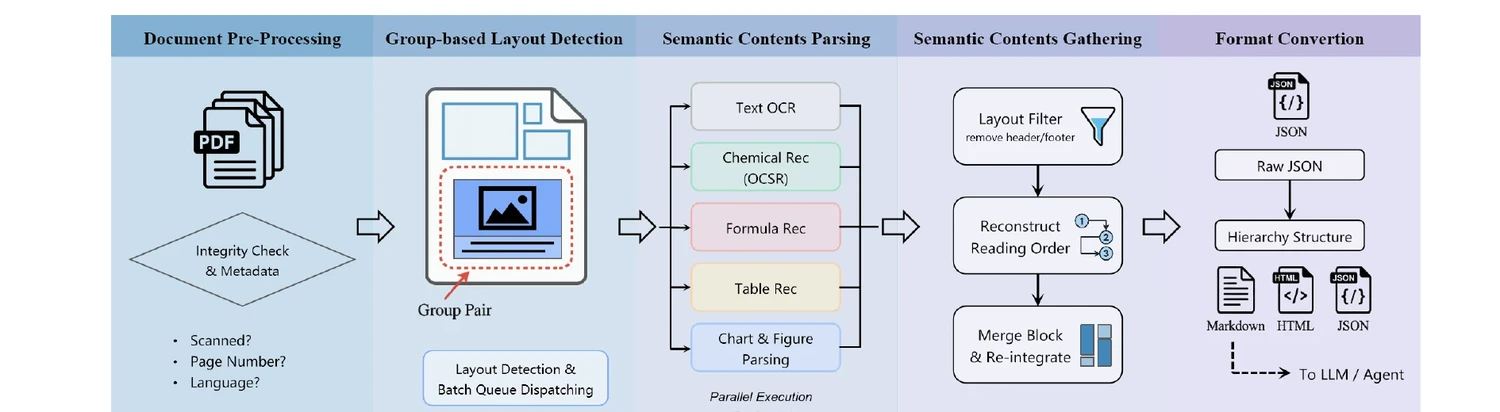
The system is organized into five sequential stages:
- Document Pre-Processing: Validates PDFs, extracts metadata, checks text accessibility, and identifies language.
- Group-based Layout Detection: Locates semantic blocks and identifies their categories using a novel tree-structured layout representation. Groups naturally paired elements (image-caption, table-title, molecule-identifier).
- Semantic Contents Parsing: Routes each block to a specialized model: OCR for text, formula recognition for equations, table structure recognition, OCSR for chemical structures, reaction extraction, and chart parsing. Over ten sub-models operate in parallel.
- Semantic Contents Gathering: Filters non-essential elements, reconstructs reading order, merges cross-page and multi-column content, and reintegrates inline multimodal elements.
- Output Formatting and Semantic Chunking: Exports parsed documents in task-specific formats with proper chunking for RAG and other downstream tasks.
Group-Based Layout Detection
A key contribution is the group-based layout detection model (Uni-Parser-LD), which uses a hierarchical tree structure to represent page layouts. Elements are organized into a bottom layer (parent nodes like paragraphs, tables, images) and a top layer (child nodes like captions, footnotes, identifiers). This preserves semantic associations between paired elements, such as molecules and their identifiers.
The model is trained on 500k pages, including 220k human-annotated pages from scientific journals and patents across 85 languages. A modified DETR-based architecture was selected as the backbone after finding that RT-DETRv2, YOLOv12, and D-FINE exhibited training instability for this task.
Chemical Structure Recognition with MolParser 1.5
Uni-Parser integrates MolParser 1.5 for OCSR, an end-to-end model that directly generates molecular representations from images. The authors explicitly note that graph-based (atom-bond) methods were the first direction they explored but ultimately abandoned because of:
- Strong reliance on rigid, hand-crafted rules that limit scalability
- Substantially higher annotation costs (over 20x compared to end-to-end approaches)
- Lower performance ceilings despite increasing training data
Molecule Localization
Uni-Parser-LD achieves strong molecule detection performance:
| Model | mAP@50 | mAP@50-95 |
|---|---|---|
| Uni-Parser-LD (Uni-Parser Bench) | 0.994 | 0.968 |
| MolDet-Doc-L | 0.983 | 0.919 |
| MolDet-General-L | 0.974 | 0.815 |
| Uni-Parser-LD (BioVista Bench) | 0.981 | 0.844 |
| MolDet-Doc-L | 0.961 | 0.871 |
| MolDet-General-L | 0.945 | 0.815 |
| BioMiner | 0.929 | - |
| MolMiner | 0.899 | - |
OCSR Accuracy
MolParser 1.5 consistently outperforms prior methods across molecule types:
| Model | Full | Chiral | Markush | All |
|---|---|---|---|---|
| MolParser 1.5 (Uni-Parser Bench) | 0.979 | 0.809 | 0.805 | 0.886 |
| MolParser 1.0 | 0.953 | 0.676 | 0.664 | 0.800 |
| MolScribe | 0.617 | 0.274 | 0.168 | 0.417 |
| MolParser 1.5 (BioVista Bench) | 0.795 | 0.604 | 0.761 | 0.780 |
| MolParser 1.0 | 0.669 | 0.352 | 0.733 | 0.703 |
| MolMiner | 0.774 | 0.497 | 0.185 | 0.507 |
| MolScribe | 0.703 | 0.481 | 0.156 | 0.455 |
| MolNexTR | 0.695 | 0.419 | 0.045 | 0.401 |
| DECIMER | 0.545 | 0.326 | 0.000 | 0.298 |
Chiral molecule recognition remains a significant challenge and is identified as a key area for future work.
Document Parsing Benchmarks
On the Uni-Parser Benchmark (150 PDFs, 2,887 pages from patents and scientific articles), Uni-Parser (HQ mode) achieves an overall score of 89.74 (excluding molecules), outperforming both pipeline tools (MinerU, PP-StructureV3) and specialized VLMs (MinerU2-VLM, DeepSeek-OCR, PaddleOCR-VL). Competing systems score zero on molecule localization and OCSR because they lack molecular recognition capabilities.
On the general-document OmniDocBench-1.5, a variant (Uni-Parser-G) using a swapped layout module achieves 89.75 overall, competitive with top-performing specialized VLMs.
Comparison with OCSR-Enabled PDF Parsers
On a controlled test set of 141 simple molecules, Uni-Parser outperforms other PDF parsing systems with OCSR support:
| Method | Recall | OCSR Success | OCSR Acc | Id Match | Time |
|---|---|---|---|---|---|
| Uni-Parser | 100% | 100% | 96.5% | 100% | 1.8s |
| MathPix | 100% | 75.9% | 59.6% | - | 66.1s |
| MinerU.Chem | 66.7% | 63.1% | 22.7% | - | ~7 min |
Reproducibility
| Artifact | Type | License | Notes |
|---|---|---|---|
| HuggingFace Models | Model/Dataset | Unknown | MolDet models and MolParser-7M dataset available |
| Project Page | Other | Unknown | Project website with documentation |
The Uni-Parser system is deployed on a cluster of 240 NVIDIA L40 GPUs (48 GB each) with 22 CPU cores and 90 GB of host memory per GPU. The reference throughput benchmark (20 pages/second) uses 8 NVIDIA RTX 4090D GPUs. The HuggingFace organization hosts MolDet detection models and several datasets (MolParser-7M, RxnBench, OmniScience), but the full Uni-Parser system code and end-to-end inference pipeline do not appear to be publicly released. MolParser 1.5 model weights are not publicly available as of this writing.
Limitations and Future Directions
- Chiral molecule recognition remains a challenge for end-to-end OCSR models
- Chemical reaction understanding in real-world literature has substantial room for improvement
- Layout models are primarily tailored to scientific and patent documents, with plans to expand to newspapers, slides, books, and financial statements
- Chart parsing falls short of industrial-level requirements across the diversity of chart types in scientific literature