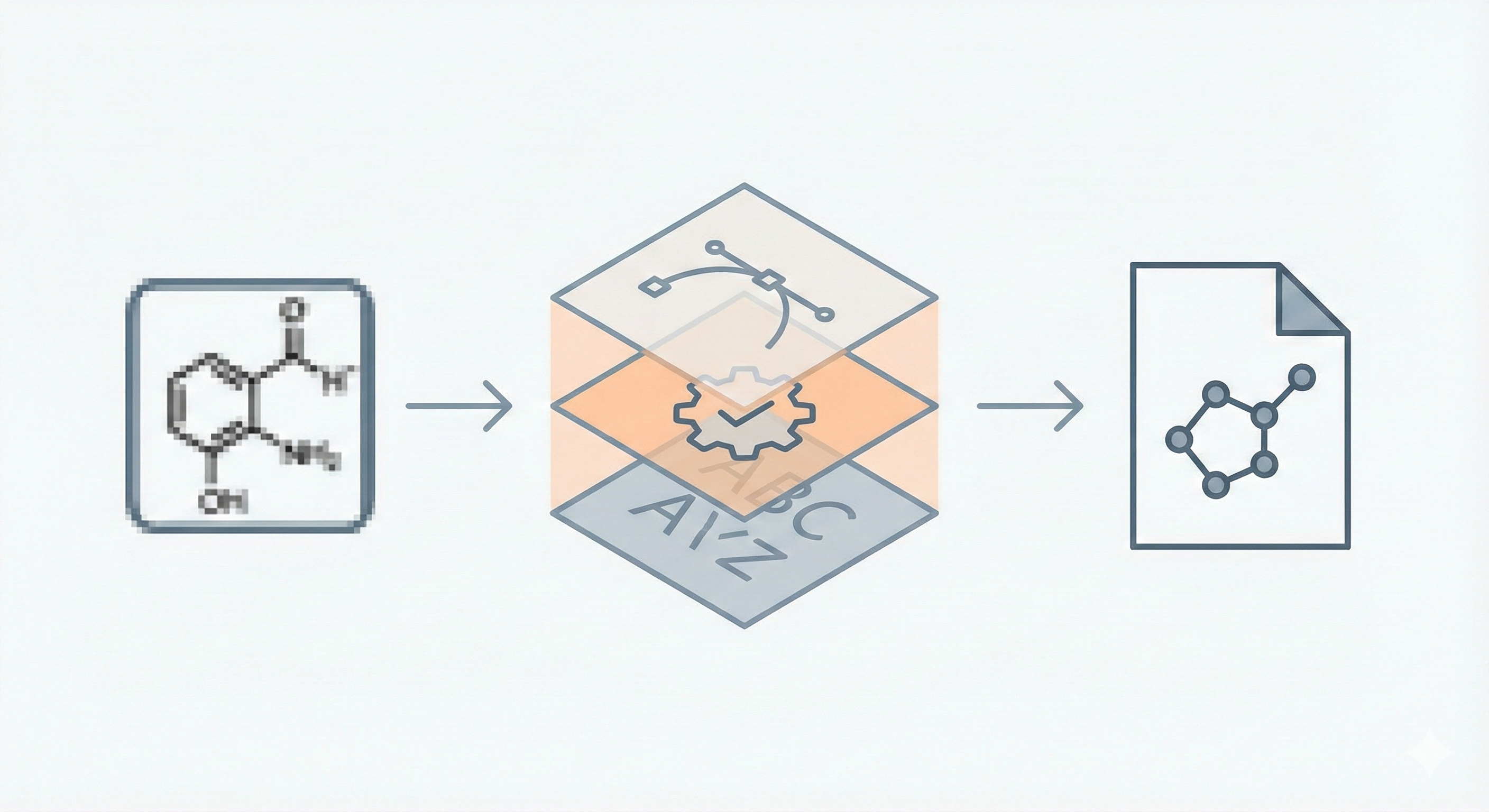
Contribution: Rule-Based Image Mining Architecture
$\Psi_{\text{Method}}$ (Methodological Basis)
This is a methodological paper describing a system architecture for image mining in the chemical domain. It focuses on the engineering challenge of converting rasterized depictions of molecules into computer-readable SDF files. The paper details the algorithmic pipeline and validates it through quantitative benchmarking against a commercial alternative.
Motivation: Digitizing Chemical Literature
- Loss of Information: Chemical software creates images. The chemical significance is lost when published in scientific literature, making the data “dead” to computers.
- Gap in Technology: Image mining lags behind advances in text mining. Existing commercial solutions (like CLIDE) faded away or remained limited.
- Scale of Problem: The colossal production of chemical documents requires automated tools to exploit this information at large scale.
Core Innovation: Graph-Preserving Vectorization
- Graph-Preserving Vectorization: The system uses a custom vectorizer designed to preserve the “graph characteristics” of chemical diagrams (1 vector = 1 line), which avoids creating spurious vectors at thick joints. It aims to generate a mathematical graph, $G = (V, E)$, mapped geometrically to the image lines.
- Chemical Knowledge Integration: A distinct module validates the reconstructed graph against chemical rules (valences, charges) to ensure the output is chemically valid.
- Hybrid Processing: The system splits the image into “connected components” for an OCR path (text/symbols) and a “body” path (bonds), reassembling them later.
Methodology & Experiments: Benchmark Validation
The authors performed a quantitative validation using three different databases where ground-truth SDF files were available. They also compared their system against the commercial tool CLIDE (Chemical Literature Data Extraction).
- Database 1: 100 images (varied line widths/fonts)
- Database 2: 100 images
- Database 3: 7,604 images (large-scale batch processing)
Results & Conclusions: Superior Accuracy over Baselines
- High Accuracy: The system achieved 94% correct reconstruction on Database 1 and 77% on Database 2. Accuracy was measured as correct recovery of identical geometry and connections.
$$ \text{Acc} = \frac{\text{Correct Images}}{\text{Total Images}} $$
- Baseline Superiority: The commercial tool CLIDE only successfully reconstructed ~50% of images in Database 1 (compared to the authors’ 94%).
- Scalability: On the large dataset (Database 3), the system achieved 67% accuracy in batch mode.
- Robustness: The authors state the system uses a handful of parameters and works robustly across different image types. CLIDE lacked flexibility and required manual intervention.
Reproducibility Details
Reproducibility Status: Closed / Not Formally Reproducible. As is common with applied research from this era, the source code, training models (SVM), and specific datasets used for benchmarking do not appear to be publicly maintained or available.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| None available | N/A | Unknown | No public code, models, or datasets were released with this 2007 publication. |
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Evaluation | Database 1 | 100 Images | Used for comparison with CLIDE; 94% success rate |
| Evaluation | Database 2 | 100 Images | 77% success rate |
| Evaluation | Database 3 | 7,604 Images | Large-scale test; 67% success rate |
Algorithms
The paper outlines a 5-module pipeline:
- Pre-processing: Adaptive histogram binarization and non-recursive connected component labeling using RLE segments.
- OCR: A “chemically oriented OCR” using wavelet functions for feature extraction and a Support Vector Machine (SVM) for classification. It distinguishes characters from molecular structure.
- Vectorizer: Assigns local directions to RLE segments and groups them into patterns. Crucially, it enforces a one-to-one mapping between image lines and graph vectors.
- Reconstruction: A rule-based module that annotates vectors:
- Stereochemistry: Registers vectors against original pixels; thick geometric forms (triangles) become chiral wedges.
- Dotted Bonds: Identifies isolated vectors and clusters them using quadtree clustering.
- Multi-bonds: Identifies parallel vectors within a dilated bounding box (factor of 2).
- Chemical Knowledge: Validates the graph valences and properties before exporting SDF.
Models
- SVM: Used in the OCR module to classify text/symbols. It supports dynamic training to correct classification mistakes.
Evaluation
The primary metric is the percentage of correctly reconstructed images (generating a valid, matching SDF file).
| Metric | System Value (DB1) | Baseline (CLIDE) | Notes |
|---|---|---|---|
| Reconstruction Accuracy | 94% | ~50% | CLIDE noted as unsuitable for batch processing |
Paper Information
Citation: Algorri, M.-E., Zimmermann, M., & Hofmann-Apitius, M. (2007). Automatic Recognition of Chemical Images. Eighth Mexican International Conference on Current Trends in Computer Science, 41-46. https://doi.org/10.1109/ENC.2007.25
Publication: ENC 2007 (IEEE Computer Society)
@inproceedings{algorriAutomaticRecognitionChemical2007,
title = {Automatic {{Recognition}} of {{Chemical Images}}},
booktitle = {Eighth {{Mexican International Conference}} on {{Current Trends}} in {{Computer Science}} ({{ENC}} 2007)},
author = {Algorri, Maria-Elena and Zimmermann, Marc and {Hofmann-Apitius}, Martin},
year = {2007},
pages = {41--46},
publisher = {IEEE},
doi = {10.1109/ENC.2007.25}
}