Atom Pair Encoding for Chemical Language Modeling
This is a Method paper that introduces Atom Pair Encoding (APE), a tokenization algorithm designed specifically for chemical string representations (SMILES and SELFIES). The primary contribution is demonstrating that a chemistry-aware tokenizer, which preserves atomic identity during subword merging, leads to improved molecular property classification accuracy in transformer-based models compared to the standard Byte Pair Encoding (BPE) approach.
Why Tokenization Matters for Chemical Strings
Existing chemical language models based on BERT/RoBERTa architectures have typically relied on BPE for tokenizing SMILES and SELFIES strings. Byte Pair Encoding (BPE) was originally designed for natural language and data compression, where it excels at breaking words into meaningful subword units. When applied to chemical strings, BPE operates at the character level without understanding chemical semantics, leading to several problems:
- Stray characters: BPE may create tokens like “C)(” that have no chemical meaning.
- Element splitting: Multi-character elements like chlorine (“Cl”) can be split into “C” and “l”, causing the model to misinterpret carbon and a dangling character.
- Lost structural context: BPE compresses sequences without considering how character position encodes molecular structure.
Previous work on SMILES Pair Encoding (SPE) attempted to address this by iteratively merging SMILES substrings into chemically meaningful tokens. However, SPE had practical limitations: its Python implementation did not support SELFIES, and it produced a smaller vocabulary (~3000 tokens) than what the data could support. These gaps motivated the development of APE.
The APE Tokenizer: Chemistry-Aware Subword Merging
APE draws inspiration from both BPE and SPE but addresses their shortcomings. The key design decisions are:
Atom-level initialization: Instead of starting from individual characters (as BPE does), APE begins with chemically valid atomic units. For SMILES, this means recognizing multi-character elements (e.g., “Cl”, “Br”) as single tokens. For SELFIES, each bracketed string (e.g., [C], [Ring1], [=O]) serves as the fundamental unit.
Iterative pair merging: Like BPE, APE iteratively merges the most frequent adjacent token pairs. The difference is that the initial tokenization preserves atomic boundaries, so merged tokens always represent valid chemical substructures.
Larger vocabulary: Using the same minimum frequency threshold of 2000, APE generates approximately 5300 unique tokens from the PubChem dataset, compared to SPE’s approximately 3000. This richer vocabulary provides more expressive power for representing chemical substructures.
SELFIES compatibility: APE natively supports both SMILES and SELFIES, using the bracketed token structure of SELFIES as its starting point for that representation.
The tokenizer was trained on a subset of 2 million molecules from PubChem (10 million SMILES total). This produced four tokenizer variants: SMILES-BPE, SMILES-APE, SELFIES-BPE, and SELFIES-APE.
Pre-training and Evaluation on MoleculeNet Benchmarks
Model architecture
All four models use the RoBERTa architecture with 6 hidden layers, a hidden size of 768, an intermediate size of 1536, and 12 attention heads. Pre-training used masked language modeling (MLM) with 15% token masking on 1 million molecules from PubChem, with a validation set of 100,000 molecules. Each model was pre-trained for 20 epochs using AdamW, with hyperparameter optimization via Optuna.
Downstream tasks
The models were fine-tuned on three MoleculeNet classification tasks:
| Dataset | Category | Compounds | Tasks | Metric |
|---|---|---|---|---|
| BBBP | Physiology | 2,039 | 1 | ROC-AUC |
| HIV | Biophysics | 41,127 | 1 | ROC-AUC |
| Tox21 | Physiology | 7,831 | 12 | ROC-AUC |
Data was split 80/10/10 (train/validation/test) following MoleculeNet recommendations. Models were fine-tuned for 5 epochs with early stopping based on validation ROC-AUC.
Baselines
Results were compared against two text-based models (ChemBERTa-2 MTR-77M and SELFormer) and two graph-based models (D-MPNN from Chemprop and MoleculeNet Graph-Conv).
Main results
| Model | BBBP ROC | HIV ROC | Tox21 ROC |
|---|---|---|---|
| SMILYAPE-1M | 0.754 +/- 0.006 | 0.772 +/- 0.010 | 0.838 +/- 0.002 |
| SMILYBPE-1M | 0.746 +/- 0.006 | 0.754 +/- 0.015 | 0.849 +/- 0.002 |
| SELFYAPE-1M | 0.735 +/- 0.015 | 0.768 +/- 0.012 | 0.842 +/- 0.002 |
| SELFYBPE-1M | 0.676 +/- 0.014 | 0.709 +/- 0.012 | 0.825 +/- 0.001 |
| ChemBERTa-2-MTR-77M | 0.698 +/- 0.014 | 0.735 +/- 0.008 | 0.790 +/- 0.003 |
| SELFormer | 0.716 +/- 0.021 | 0.769 +/- 0.010 | 0.838 +/- 0.005 |
| MoleculeNet-Graph-Conv | 0.690 | 0.763 | 0.829 |
| D-MPNN | 0.737 | 0.776 | 0.851 |
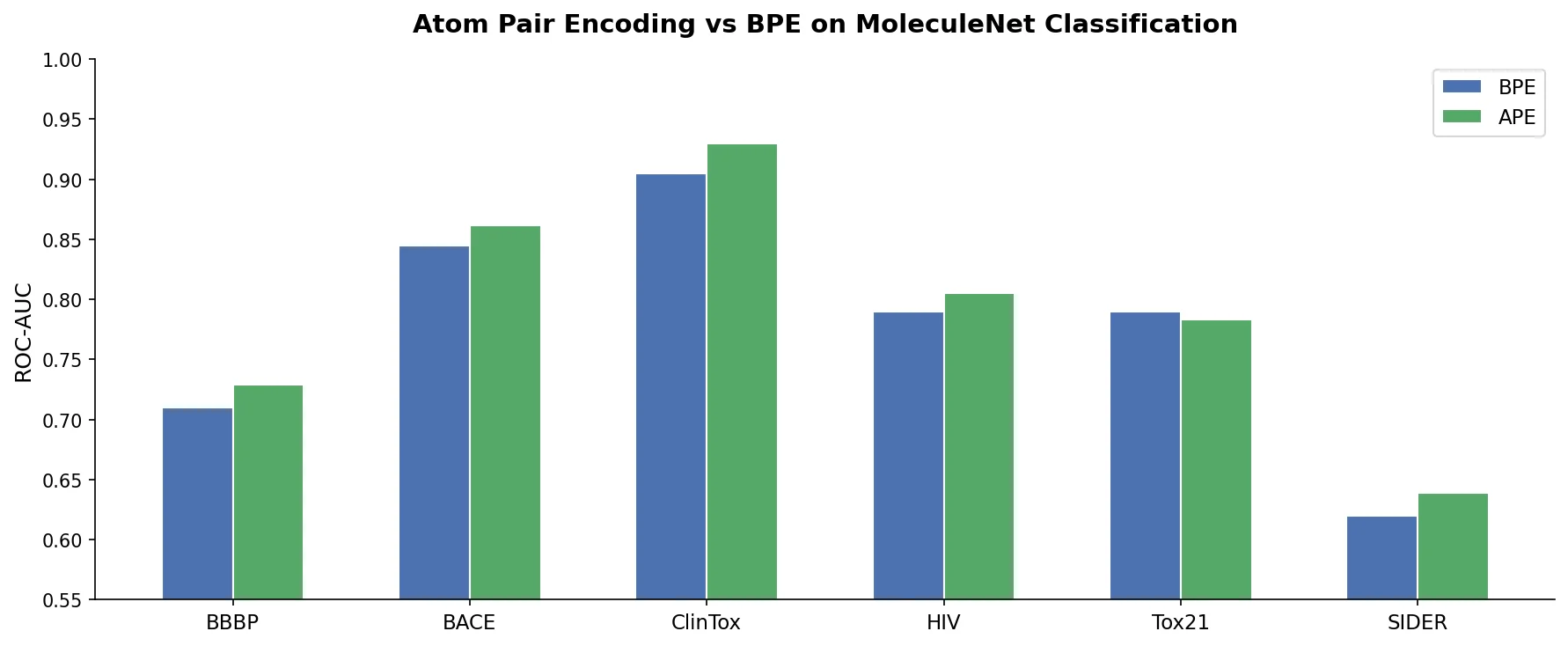
APE consistently outperforms BPE for both SMILES and SELFIES. SMILYAPE achieves the best BBBP score (0.754), beating D-MPNN (0.737). On HIV, SMILYAPE (0.772) is competitive with D-MPNN (0.776). On Tox21, D-MPNN (0.851) leads, with SMILYBPE (0.849) and SELFYAPE (0.842) close behind.
Statistical significance
Mann-Whitney U tests confirmed statistically significant differences between SMILYAPE and SMILYBPE (p < 0.05 on all datasets). Cliff’s delta values indicate large effect sizes: 0.74 (BBBP), 0.70 (HIV), and -1.00 (Tox21, favoring BPE). For SELFIES models, SELFYAPE achieved Cliff’s delta of 1.00 across all three datasets, indicating complete separation from SELFYBPE.
Key Findings and Limitations
APE outperforms BPE by preserving atomic identity
The consistent advantage of APE over BPE stems from APE’s atom-level initialization. By starting with chemically valid units rather than individual characters, APE avoids creating nonsensical tokens that break chemical elements or mix structural delimiters with atoms.
SMILES outperforms SELFIES with APE tokenization
SMILYAPE generally outperforms SELFYAPE across tasks. Attention weight analysis revealed that SMILYAPE assigns more weight to immediate neighboring tokens (0.108 vs. 0.096) and less to distant tokens (0.030 vs. 0.043). This pattern aligns with chemical intuition: bonding is primarily determined by directly connected atoms. SMILYAPE also produces more compact tokenizations (8.6 tokens per molecule vs. 11.9 for SELFYAPE), potentially allowing more efficient attention allocation.
SELFIES models show higher inter-tokenizer agreement
On the BBBP dataset, all true positives identified by SELFYBPE were also captured by SELFYAPE, with SELFYAPE achieving higher recall (61.68% vs. 55.14%). In contrast, SMILES-based models shared only 29.3% of true positives between APE and BPE variants, indicating that tokenization choice has a larger impact on SMILES models.
Limitations
- Pre-training used only 1 million molecules, compared to 77 million for ChemBERTa-2. Despite this, APE models were competitive or superior, but scaling effects remain unexplored.
- Evaluation was limited to three binary classification tasks from MoleculeNet. Regression tasks, molecular generation, and reaction prediction were not tested.
- The Tox21 result is notable: SMILYBPE outperforms SMILYAPE (0.849 vs. 0.838), suggesting APE’s advantage may be task-dependent.
- No comparison with recent atom-level tokenizers like Atom-in-SMILES or newer approaches beyond SPE.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Tokenizer training | PubChem subset | 2M molecules | SMILES strings converted to SELFIES via selfies library |
| Pre-training | PubChem subset | 1M molecules | 100K validation set |
| Evaluation | BBBP | 2,039 compounds | 80/10/10 split |
| Evaluation | HIV | 41,127 compounds | 80/10/10 split |
| Evaluation | Tox21 | 7,831 compounds | 80/10/10 split, 12 tasks |
Algorithms
- Tokenizers: BPE (via Hugging Face), APE (custom implementation, minimum frequency 2000)
- Pre-training: Masked Language Modeling (15% masking) for 20 epochs
- Optimizer: AdamW with Optuna hyperparameter search
- Fine-tuning: 5 epochs with early stopping on validation ROC-AUC
Models
- Architecture: RoBERTa with 6 layers, hidden size 768, intermediate size 1536, 12 attention heads
- Four variants: SMILYAPE, SMILYBPE, SELFYAPE, SELFYBPE
Evaluation
| Metric | SMILYAPE | SMILYBPE | SELFYAPE | SELFYBPE |
|---|---|---|---|---|
| BBBP ROC-AUC | 0.754 | 0.746 | 0.735 | 0.676 |
| HIV ROC-AUC | 0.772 | 0.754 | 0.768 | 0.709 |
| Tox21 ROC-AUC | 0.838 | 0.849 | 0.842 | 0.825 |
Hardware
- NVIDIA RTX 3060 GPU with 12 GiB VRAM
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| APE Tokenizer | Code | Other (unspecified SPDX) | Official APE tokenizer implementation |
| PubChem10M SMILES/SELFIES | Dataset | Not specified | 10M SMILES with SELFIES conversions |
| Pre-trained and fine-tuned models | Model | Not specified | All four model variants on Hugging Face |
Paper Information
Citation: Leon, M., Perezhohin, Y., Peres, F., Popovič, A., & Castelli, M. (2024). Comparing SMILES and SELFIES tokenization for enhanced chemical language modeling. Scientific Reports, 14(1), 25016. https://doi.org/10.1038/s41598-024-76440-8
@article{leon2024comparing,
title={Comparing SMILES and SELFIES tokenization for enhanced chemical language modeling},
author={Leon, Miguelangel and Perezhohin, Yuriy and Peres, Fernando and Popovi{\v{c}}, Ale{\v{s}} and Castelli, Mauro},
journal={Scientific Reports},
volume={14},
number={1},
pages={25016},
year={2024},
publisher={Nature Publishing Group},
doi={10.1038/s41598-024-76440-8}
}