A Hybrid Molecular Representation Combining SMILES and Chemical-Environment Tokens
This is a Method paper that introduces SMI+AIS(N), a hybrid molecular string representation combining standard SMILES tokens with Atom-In-SMILES (AIS) tokens. AIS tokens encode local chemical environment information (central atom, ring membership, and neighboring atoms) into a single token. The key contribution is a systematic hybridization strategy that selectively replaces the most frequent SMILES tokens with AIS equivalents, preserving SMILES grammar compatibility while enriching token diversity. The method is validated on molecular structure generation via latent space optimization for drug design.
Limitations of Standard SMILES for Machine Learning
SMILES is the most widely adopted string-based molecular representation, used in major databases like ZINC and PubChem. Despite this ubiquity, SMILES has several well-known limitations for machine learning applications:
- Non-unique representations: The same molecule can be encoded as multiple distinct SMILES strings.
- Invalid string generation: Generative models can produce syntactically invalid SMILES that do not correspond to any molecule.
- Limited token diversity: SMILES tokens map one-to-one to atoms or bonds, so the token vocabulary is restricted to the available atom and bond types.
- Insufficient chemical context: Individual SMILES tokens carry no information about the local chemical environment of an atom.
Alternative representations like SELFIES (guaranteeing validity) and InChI (guaranteeing uniqueness) address some of these issues but share the same fundamental limitation of low token diversity. The Atom-In-SMILES (AIS) representation (Ucak et al., 2023) enriches tokens with neighboring atom and ring information, but using AIS exclusively produces a large vocabulary with many infrequent tokens that can cause data sparsity problems. The authors aim to find a middle ground: adding chemical context to the most common tokens while keeping the vocabulary manageable.
Core Innovation: Selective Token Hybridization with AIS
The SMI+AIS(N) representation hybridizes standard SMILES with AIS tokens through a frequency-based selection process:
AIS Token Structure
Each AIS token encodes three pieces of information about an atom, delimited by semicolons:
$$ \lbrack \text{central atom} ; \text{ring info} ; \text{neighbor atoms} \rbrack $$
For example, the oxygen in a carboxyl group of benzoic acid is represented as [O;!R;C], meaning: oxygen atom, not in a ring, bonded to carbon. In standard SMILES, this would simply be O.
Hybridization Procedure
- Convert all SMILES strings in the ZINC database to their full AIS representations.
- Count the frequency of each AIS token across the database.
- Select the top-N most frequent AIS tokens to form the hybrid vocabulary.
- In the hybrid representation, atoms matching these top-N AIS tokens are written in AIS notation; all other atoms use standard SMILES notation.
For benzoic acid, the hybridization produces:
$$ \text{SMI}: \texttt{O=C(O)c1ccccc1} $$
$$ \text{SMI+AIS}: \texttt{\lbrack O;!R;C\rbrack=\lbrack C;!R;COO\rbrack(\lbrack OH;!R;C\rbrack)c1ccccc1} $$
The parameter N controls vocabulary size. The authors test N = 50, 100, 150, and 200, finding that N = 100-150 provides the best balance for the ZINC database.
Token Frequency Rebalancing
A key benefit of hybridization is mitigating the severe token frequency imbalance in standard SMILES. Carbon (C), the most frequent element with ~184 million occurrences in ZINC, is represented by only 16 token types in SMILES. With SMI+AIS(200), carbon is distinguished into 145 token types based on chemical environment, with 74% of carbon occurrences represented by AIS tokens. Less common elements like halogens see minimal change (only 2% AIS representation), which avoids introducing unnecessarily rare tokens.
| Element | Frequency | SMILES Types | SMI+AIS(100) Types (AIS %) | SMI+AIS(200) Types (AIS %) |
|---|---|---|---|---|
| C | 183,860,954 | 16 | 78 (73%) | 145 (74%) |
| O | 27,270,229 | 8 | 16 (11%) | 24 (11%) |
| N | 26,022,928 | 11 | 32 (1%) | 46 (10%) |
| X (halogens) | 6,137,030 | 7 | 10 (2%) | 11 (2%) |
| S | 4,581,307 | 12 | 17 (2%) | 24 (2%) |
Latent Space Optimization for Molecular Generation
Model Architecture
The evaluation uses a conditional variational autoencoder (CVAE) with:
- Encoder: BERT-style architecture with entity and positional embeddings, 4 multi-head attention layers (8 heads each), producing mean and standard deviation vectors in latent space.
- Decoder: 4 stacked gated recurrent unit (GRU) layers that transform sampled latent vectors (conditioned) back into token sequences.
- Training: 20 epochs on 9 million compounds from the ZINC database (8:1:1 train/valid/test split) under identical conditions for all representations.
Optimization Setup
Bayesian Optimization (BO) via BoTorch is applied to the CVAE latent space, maximizing a multi-objective function:
$$ \text{Obj} = -\text{BA} - 0.5 \times \text{SA}^2 $$
where BA is binding affinity (docking score from QuickVina 2, lower is stronger) and SA is synthetic accessibility score (from RDKit, lower is more synthesizable). Each BO iteration generates 800 candidate latent vectors. Invalid strings receive a penalty objective value of -100.
Protein Targets
Four diverse targets were used to assess generalizability:
- PDK4 (Pyruvate Dehydrogenase Kinase 4): narrow, deep binding pocket
- 5-HT1B (Serotonin Receptor 1B): shallow, open GPCR conformation
- PARP1 (Poly ADP-ribose Polymerase 1): small, flexible molecule binding site
- CK1d (Casein Kinase I Delta): broad, accessible conformation
Protein structures were obtained from the Protein Data Bank (PDB IDs: 4V26, 4IAQ, 6I8M, 4TN6). Each optimization was run 10 times independently from the same 5 initial compounds selected from BindingDB.
Key Results
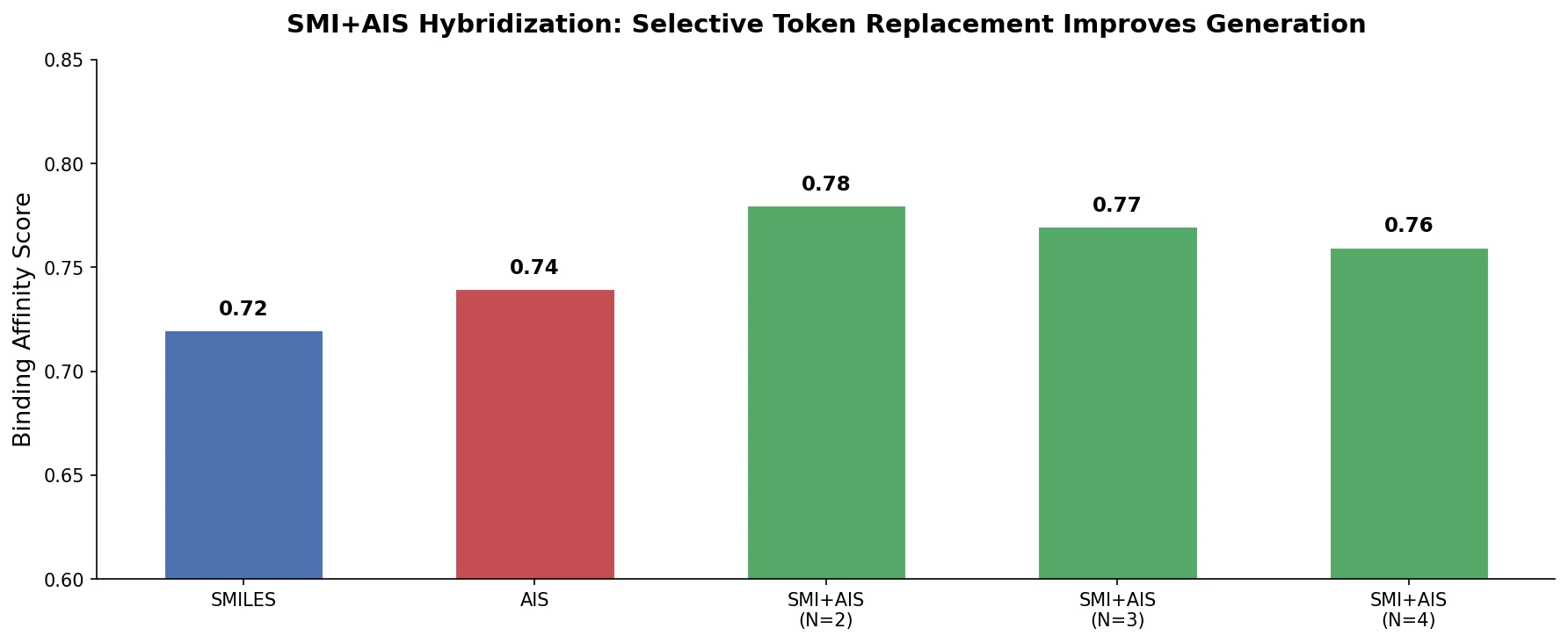
SMI+AIS(100) consistently achieved the highest objective values across protein targets.
PDK4 Optimization (Top-1 results over 10 independent runs):
- SMI+AIS(100) achieved approximately 12% improvement over standard SMILES and 28% improvement over SELFIES based on median Top-1 objective values.
- Generated structures exhibited BA scores between -10 and -9 and SA scores between 2.0 and 2.3.
- Molecular weights clustered around 400 amu, consistent with the CVAE conditioning.
Validity Ratios: Standard SMILES produced approximately 40% valid structures. SMI+AIS representations showed significant improvement as N increased, though SMI+AIS(200) showed slight saturation, likely from insufficiently trained infrequent tokens.
SELFIES: Despite achieving the highest validity ratio, SELFIES failed to generate chemically meaningful structures with desirable BA and SA scores. The authors attribute this to SELFIES grammar where token meaning is highly context-dependent, causing minor latent space variations to produce large structural changes.
Cross-target consistency: Improvements were observed across all four protein targets, with slight variation (5-HT1B showed smaller differences between SMI and SMI+AIS(100) for Top-1, while other targets showed significant improvements).
Improved Molecular Generation Through Chemical Context Enrichment
The SMI+AIS(N) representation achieves consistent improvements in molecular generation quality compared to both standard SMILES and SELFIES. The core findings are:
- Binding affinity improvement: Approximately 7% improvement over standard SMILES for the PDK4 target.
- Synthesizability improvement: Approximately 6% increase in synthetic accessibility scores.
- Target independence: Performance gains transfer across four structurally diverse protein targets.
- Preserved structural motifs: The generative model retains chemically meaningful fragments (e.g., acetamide and piperidine) from initial compounds without explicit fragment constraints.
Limitations
The authors acknowledge several limitations:
- Stereochemistry: SMI+AIS inherits the limited stereochemistry handling of standard SMILES.
- Evaluation scope: Only molecular generation was tested; property prediction and other ML tasks remain unexplored.
- Compute constraints: The study was limited to molecular generation due to computing power and time.
- Single optimization strategy: Only latent space optimization with Bayesian optimization was evaluated; other generative approaches were not compared.
Future Directions
The authors suggest extending SMI+AIS to diverse benchmarking tests including molecular property prediction, experimental validation, and broader applications of chemical language models.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training/Vocab | ZINC Database | 9M compounds | Canonicalized, deduplicated, split 8:1:1 |
| Binding targets | BindingDB | 5 initial compounds per target | Selected for each protein target |
| Protein structures | PDB | 4 structures | IDs: 4V26, 4IAQ, 6I8M, 4TN6 |
Algorithms
- Tokenization: AIS token frequency counting on full ZINC database, top-N selection
- Generative model: Conditional VAE with BERT encoder (4 layers, 8 heads) and GRU decoder (4 layers)
- Optimization: Bayesian Optimization via BoTorch (800 candidates per iteration)
- Docking: QuickVina 2 with 25 A pocket size, 10 docking simulations per ligand
- SA scoring: RDKit SA score
- Training: 20 epochs for all representations under identical conditions
Models
- CVAE architecture details in supplementary (Fig. S9, Tables S2, S4)
- No pre-trained weights released
Evaluation
| Metric | SMI+AIS(100) vs SMILES | SMI+AIS(100) vs SELFIES | Notes |
|---|---|---|---|
| Median Top-1 Obj. Value | +12% | +28% | PDK4 target |
| Validity Ratio | Higher than ~40% (SMILES) | Lower than SELFIES | SMI+AIS improves with N |
| BA (binding affinity) | ~7% improvement | Substantial | Lower (more negative) is better |
| SA (synthesizability) | ~6% improvement | Substantial | Lower is more synthesizable |
Hardware
Hardware details are not specified in the main text. Optimization wall times are reported in supplementary Table S5.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| AIS-Drug-Opt | Code | Not specified | Source code and datasets for reproduction |
Reproducibility Status: Partially Reproducible. Code and processed data are publicly available on GitHub, but no pre-trained model weights are released, the license is unspecified, and hardware requirements are not documented in the main text.
Paper Information
Citation: Han, H., Yeom, M. S., & Choi, S. (2025). Hybridization of SMILES and chemical-environment-aware tokens to improve performance of molecular structure generation. Scientific Reports, 15, 16892. https://doi.org/10.1038/s41598-025-01890-7
@article{han2025hybridization,
title={Hybridization of SMILES and chemical-environment-aware tokens to improve performance of molecular structure generation},
author={Han, Herim and Yeom, Min Sun and Choi, Sunghwan},
journal={Scientific Reports},
volume={15},
number={1},
pages={16892},
year={2025},
publisher={Springer Nature},
doi={10.1038/s41598-025-01890-7}
}