A Systematization of Molecular Representation Foundation Models
This paper is a Systematization that provides the first comprehensive review of foundation models for molecular representation learning (MRL). The authors classify existing models by their input modality (unimodal vs. multimodal), analyze four mainstream pretraining strategies, survey five downstream application domains, and propose practical guidelines for model selection. The review covers over 35 representative models published between 2020 and 2024, with parameter counts ranging from 2 million to over 1 trillion.
Why a Systematic Review of MRL Foundation Models Is Needed
Molecular representation learning transforms molecular structures and properties into numerical vectors that serve as inputs for machine learning models. The field has evolved rapidly from molecular fingerprints through SMILES-based sequence models to graph neural networks and 3D geometry-aware architectures. Foundation models, characterized by large-scale pretraining on unlabeled molecular data followed by fine-tuning on downstream tasks, have introduced new opportunities for generalizability and transfer learning in drug discovery.
Despite this rapid progress, the authors identify a gap: no prior work has systematically reviewed MRL foundation models across all input modalities and pretraining paradigms. Existing surveys tend to focus on specific representations (e.g., graph-based methods) or specific applications (e.g., property prediction) without providing the cross-cutting perspective needed to guide model selection. This review fills that gap by offering a unified taxonomy and practical guidelines.
Taxonomy of Molecular Descriptors and Model Architectures
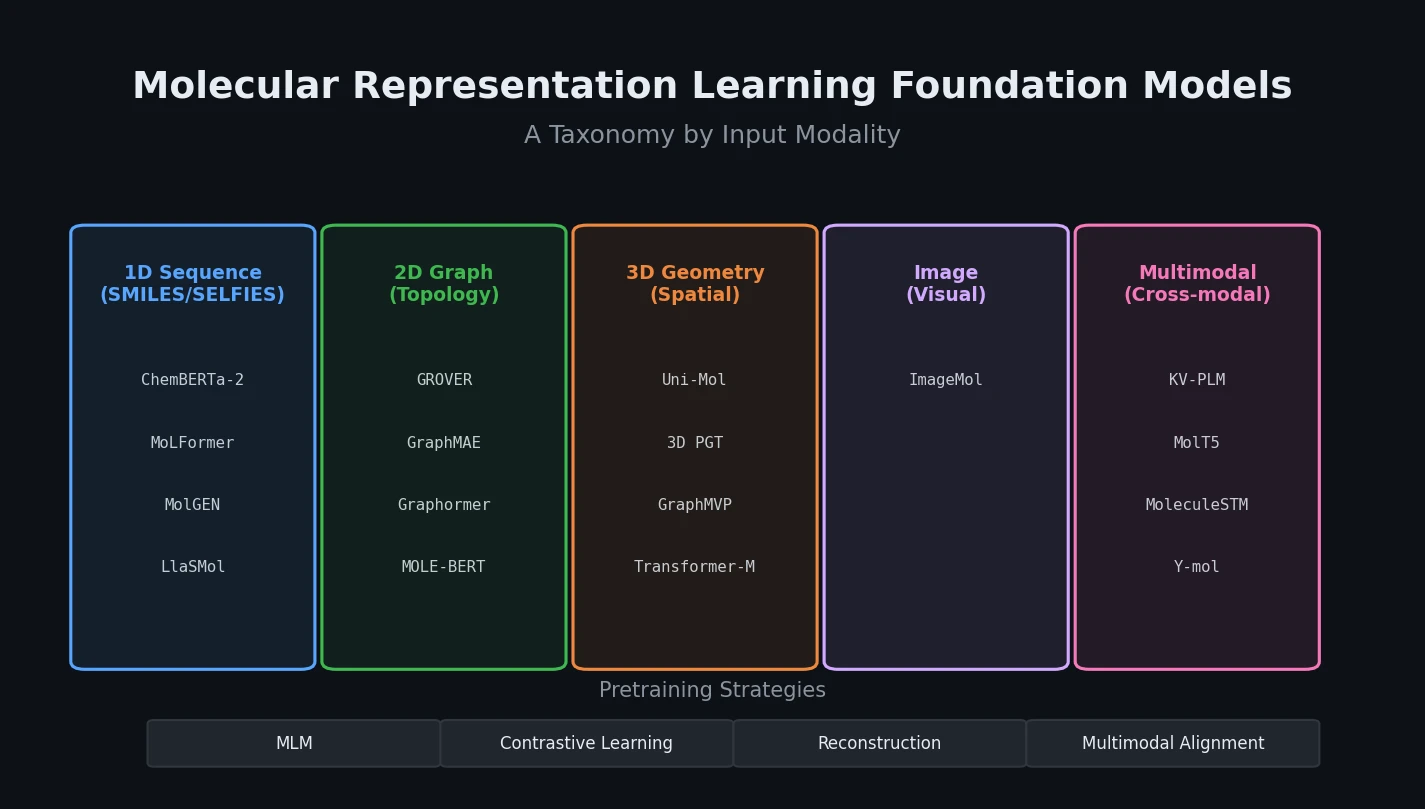
The core organizational framework classifies models along two axes: the molecular descriptor used as input and the backbone architecture.
Molecular Descriptors
The review identifies five primary descriptor types:
- Molecular fingerprints: Binary vectors encoding structural features (e.g., Morgan fingerprints). Rarely used in foundation models due to information loss and dimensional complexity.
- 1D sequences: SMILES and SELFIES string representations. SMILES is compact and widely used but can produce invalid molecules. SELFIES guarantees valid molecular strings by construction.
- 2D topological graphs: Atoms as nodes, bonds as edges. Can be derived from SMILES via RDKit, making graph datasets effectively interchangeable with SMILES datasets.
- 3D geometry: Spatial coordinates capturing conformational information, energy states, and stereochemistry. Experimentally expensive to obtain, limiting dataset availability.
- Multimodal: Combinations of the above with text, IUPAC names, knowledge graphs, and molecular images.
The paper also discusses mathematically abstract molecular representations. For example, the Wiener index quantifies structural complexity:
$$ W = \frac{1}{2} \sum_{i < j} d_{ij} $$
where $d_{ij}$ is the topological distance (shortest bonding path length) between atoms $i$ and $j$.
Degree centrality captures local connectivity:
$$ C_{D}(v_{i}) = \sum_{j=1}^{n} A_{ij} $$
where $A \in \mathbb{R}^{n \times n}$ is the molecular graph adjacency matrix.
Model Architectures
Models are classified into two primary categories:
Unimodal-based models:
- Sequence-based: Transformer models operating on SMILES/SELFIES (e.g., ChemBERTa-2, MoLFormer, MolGEN, LlaSMol). These capture syntactic patterns but miss spatial and topological features.
- Topological graph-based: GNN variants (GIN, GCN, GAT) and Transformer-based graph models (Graphormer). GNNs capture local topology through message passing; Transformers overcome locality limitations through global self-attention.
- 3D geometry-based: Models like Uni-Mol and 3D PGT that incorporate spatial coordinates. Uni-Mol uses distance-aware self-attention with an SE(3)-equivariant coordinate head for rotation/translation invariance.
- Image-based: CNN-based models (ImageMol) that process 2D molecular images using visual representation learning.
Multimodal-based models:
- Sequence + Graph: DVMP, PanGu Drug Model. Combines the strengths of string and topological representations.
- Graph + 3D Geometry: GraphMVP, Transformer-M. Enriches topological features with spatial information.
- Text + Molecular Structure: KV-PLM, MolT5, MoleculeSTM, MolReGPT, Y-mol. Aligns molecular structural information with biomedical text through cross-modal learning.
Four Pretraining Paradigms for MRL
The review systematically categorizes pretraining strategies into four paradigms:
Masked Language Modeling (MLM)
The cornerstone strategy for sequence-based models. Randomly masks tokens in molecular sequences and trains the model to predict them. ChemBERTa pretrained on 77 million SMILES sequences from PubChem achieves 5-10% improvement in AUC-ROC on property prediction tasks compared to task-specific models. MLM captures local dependencies and global sequence patterns but cannot model spatial or topological features, making it best suited for unimodal sequence inputs.
Contrastive Learning (CL)
The dominant strategy for multimodal models. Constructs positive-negative sample pairs to align features across modalities or views. In unimodal settings, CL generates negative samples by perturbing molecular graphs. In multimodal settings, it aligns features from different modalities. GraphMVP, which contrasts 2D topological features with 3D spatial features, reduces RMSE by 15% on QM9 energy prediction compared to unimodal models. Performance depends heavily on the quality of positive sample construction.
Reconstruction-Based Pretraining (RBP)
Learns global molecular features by reconstructing original data from corrupted inputs. Tasks include node feature reconstruction, graph structure reconstruction, and coordinate/energy reconstruction. MGMAE masks more than 50% of nodes and edges in molecular graphs and trains the model to reconstruct them, achieving 94.2% AUC-ROC on BBBP. RBP captures global structural patterns but requires high model complexity and training cost.
Multimodal Alignment Pretraining (MAP)
Designed for multimodal inputs, aligning and fusing features from different modalities through cross-modal tasks. KV-PLM uses SMILES-to-text matching to align molecular structure and functional information. MAP fuses structural information (SMILES, graphs) with semantic information (text) but requires large-scale cross-modal labeled data, posing significant data acquisition challenges.
Downstream Applications and Performance Benchmarks
The review evaluates MRL foundation models across five application domains.
Molecular Property Prediction
The most common benchmark for MRL models. The review provides comprehensive ROC-AUC comparisons across eight MoleculeNet classification datasets:
| Model | Type | BBBP | BACE | ClinTox | Tox21 | SIDER | HIV |
|---|---|---|---|---|---|---|---|
| MGMAE | Graph | 94.2 | 92.7 | 96.7 | 86.0 | 66.4 | - |
| MPG | Graph | 92.2 | 92.0 | 96.3 | 83.7 | 66.1 | - |
| GROVER | Graph+Trans. | 94.0 | 89.4 | 94.4 | 83.1 | 65.8 | - |
| MoLFormer | Sequence | 93.7 | 88.2 | 94.8 | 84.7 | 69.0 | 82.2 |
| MM-Deacon | Seq.+IUPAC | 78.5 | - | 99.5 | - | 69.3 | 80.1 |
| Uni-Mol | 3D | 72.9 | 85.7 | 91.9 | 79.6 | 65.9 | 80.8 |
| DVMP | Seq.+Graph | 77.8 | 89.4 | 95.6 | 79.1 | 69.8 | 81.4 |
| TxD-T-LLM | Seq.+Text | - | - | 86.3 | 88.2 | - | 73.2 |
The table shows that no single architecture dominates across all datasets. Transformer- and GIN-based architectures with graph inputs generally perform well. The review notes that model effectiveness depends heavily on the dataset, with Mole-BERT encountering negative transfer due to a small and unbalanced atomic vocabulary.
Molecular Generation
MolGEN (SELFIES-based, 8B parameters) achieves 100% validity on synthetic molecules. MolT5 excels at text-to-molecule generation. Uni-Mol generates 3D conformations with 97.95% coverage on QM9.
Drug-Drug Interaction Prediction
MPG achieves 96.6% AUC-ROC on BIOSNAP by combining unsupervised pretraining with supervised fine-tuning and multi-task learning.
Retrosynthesis Prediction
DVMP achieves 66.5% top-1 accuracy on USPTO-50K when reaction types are provided as priors (54.2% without).
Drug Synergy Prediction
SynerGPT (GPT-based) achieves 77.7% AUC-ROC in few-shot settings for novel drug combinations, outperforming baselines through contextual learning.
Guidelines, Limitations, and Future Directions
Model Selection Guidelines
The authors provide structured guidelines for choosing MRL foundation models based on:
- Task objective: Property prediction favors GNNs or large pretrained frameworks (ChemBERTa-2, Uni-Mol). Generation tasks favor GPT-style autoregressive models (MolGEN). Retrosynthesis benefits from multimodal architectures.
- Data characteristics: SMILES/graph representations suit generation tasks. Knowledge graph-enhanced models benefit interaction and synergy prediction. Transfer learning helps data-limited scenarios.
- Interpretability needs: Transformer architectures are preferred when interpretability is required, as attention matrices enable visualization of learned molecular features.
- Computational budget: GIN-based models have $\mathcal{O}(|V| + |E|)$ complexity, while Transformer-based models scale as $\mathcal{O}(n^2 \cdot d)$.
Limitations and Future Directions
The review identifies five key challenges:
- Multimodal data integration: Each representation paradigm has distinct limitations (1D neglects spatial configuration, 2D omits conformational details, 3D faces rotational invariance challenges). The authors propose incorporating molecular dynamics trajectories as a dynamic modality and using cross-modal data augmentation.
- Data scarcity: Semi-supervised learning can achieve more than 90% of fully supervised performance using only 10% labeled data on QM9. Cross-modal augmentation (e.g., 3D InfoMax) can generate plausible 3D conformers from 2D graphs.
- Interpretability: Current methods rely primarily on attention-based visualization, which is insufficient for multimodal models. The authors suggest assessing decision consistency across modalities and incorporating chemical knowledge graphs.
- Training efficiency: Large parameter counts demand distributed parallel training techniques, with data parallelism being the most common approach.
- Robustness and generalization: Strategies include data augmentation (multiple SMILES representations, 3D conformer generation), meta-learning for rapid adaptation, and sparse attention mechanisms to reduce sensitivity to irrelevant long-range interactions.
Reproducibility Details
This is a review paper, so standard reproducibility criteria for experimental papers do not directly apply. The review compiles results from the original publications of each surveyed model.
Data
The review catalogs 28 representative molecular datasets used by the surveyed foundation models:
| Dataset | Size | Descriptor | Primary Use |
|---|---|---|---|
| PubChem | ~118M | SMILES, 3D, Image, IUPAC | Pretraining |
| ZINC15 | ~980M | SMILES | Pretraining |
| ChEMBL | ~2.4M | SMILES | Pretraining |
| QM9 | 133,884 | SMILES | Property prediction |
| GEOM | 450,000 | 3D coordinates | Property prediction |
| USPTO-full | 950,000 | SMILES | Reaction prediction |
| Molecule3D | 4M | 3D coordinates | Property prediction |
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| Review Materials (GitHub) | Code/Data | Not specified | Code and data tables for figures |
| Paper (PMC) | Paper | CC-BY | Open access via PubMed Central |
Evaluation
All performance metrics reported in the review are directly cited from the original studies. The evaluation protocols follow each model’s original setup. The review covers:
- ROC-AUC for classification tasks (property prediction, DDI, synergy)
- RMSE/MAE for regression tasks
- Validity and novelty for molecular generation
- Top-k accuracy for retrosynthesis
- COV and MAT for conformation generation
Paper Information
Citation: Song, B., Zhang, J., Liu, Y., Liu, Y., Jiang, J., Yuan, S., Zhen, X., & Liu, Y. (2025). A systematic review of molecular representation learning foundation models. Briefings in Bioinformatics, 27(1), bbaf703. https://doi.org/10.1093/bib/bbaf703
@article{song2025systematic,
title={A systematic review of molecular representation learning foundation models},
author={Song, Bosheng and Zhang, Jiayi and Liu, Ying and Liu, Yuansheng and Jiang, Jing and Yuan, Sisi and Zhen, Xia and Liu, Yiping},
journal={Briefings in Bioinformatics},
volume={27},
number={1},
pages={bbaf703},
year={2025},
publisher={Oxford University Press},
doi={10.1093/bib/bbaf703}
}