A New Molecular String Notation for Generative Models
This is a Method paper that introduces DeepSMILES, a modified SMILES syntax designed to reduce the rate of syntactically invalid strings produced by machine-learning generative models. The primary contribution is a pair of string-level transformations (for ring closures and for branches) that can be applied independently and interconverted with standard SMILES without loss of information, including stereochemistry.
The Problem of Invalid SMILES in Molecular Generation
Deep neural networks for de novo molecular design commonly operate on SMILES strings. Variational autoencoders (Gomez-Bombarelli et al., 2018), recurrent neural networks with LSTM (Segler et al., 2018; Olivecrona et al., 2017), and grammar-based approaches (Kusner et al., 2017) all generate molecules by sampling character sequences. A persistent problem is that many generated strings are syntactically invalid SMILES, with reported validity rates ranging from 7% to 80%.
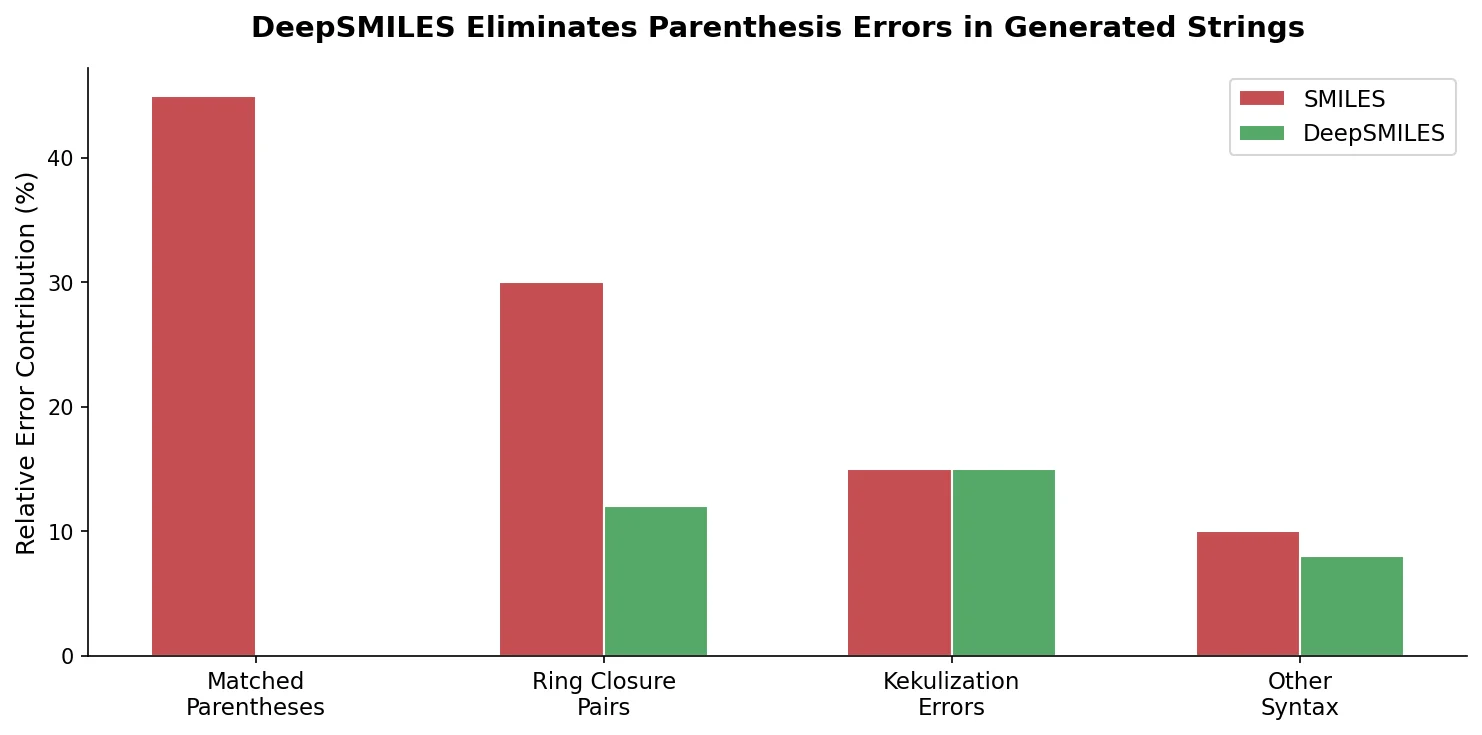
Two structural features of SMILES syntax are responsible for most invalid strings:
- Balanced parentheses: Branches require matched open/close parenthesis pairs. A generative model must track nesting state across long sequences to produce valid brackets.
- Paired ring closure symbols: Rings require two identical digit tokens at corresponding positions. The model must remember which digits are “open” and close them appropriately.
Grammar-based approaches (e.g., Grammar VAE) can enforce balanced parentheses through a context-free grammar, but they cannot enforce the ring closure pairing constraint because that constraint is context-sensitive. Syntax-directed approaches (Dai et al., 2018) add explicit ring closure constraints but at the cost of significantly more complex decoder architectures.
Core Innovation: Postfix Branch Notation and Single Ring Closure Symbols
DeepSMILES addresses both syntax problems through two independent string transformations.
Ring closure transformation
Standard SMILES uses a pair of identical digits to mark ring openings and closings (e.g., c1ccccc1 for benzene). DeepSMILES eliminates the ring-opening digit and replaces the ring-closing digit with the ring size, counting back along the tree path to the ring-opening atom. Benzene becomes cccccc6, where 6 means “connect to the atom 6 positions back.”
This transformation has three key properties:
- Every ring of a given size always uses the same digit, regardless of context. A phenyl ring is always
cccccc6in DeepSMILES, whereas in SMILES it might bec1ccccc1,c2ccccc2,c3ccccc3, etc. - A single symbol cannot be “unmatched” since there is no corresponding opening symbol.
- For double-digit ring sizes, the
%Nnotation is used (and%(N)for sizes above 99).
Bond stereochemistry is preserved by moving any explicit or stereo bond from the eliminated ring-opening symbol to the ring-closing symbol, with direction adjusted as needed.
Branch (parenthesis) transformation
Standard SMILES uses matched open/close parenthesis pairs for branches (e.g., C(OC)(SC)F). DeepSMILES replaces this with a postfix notation inspired by Reverse Polish Notation (RPN). Only close parentheses are used, and the number of consecutive close parentheses indicates how far back on the current branch the next atom attaches.
For example, C(OC)(SC)F becomes COC))SC))F. The interpretation uses a stack: atoms are pushed onto the stack as they are read, each close parenthesis pops one atom from the stack, and the next atom connects to whatever is on top of the stack.
Stereochemistry preservation
Tetrahedral stereochemistry is fully preserved through the transformations. When ring closure symbol reordering would change the stereo configuration, the @/@@ annotation is inverted during encoding to compensate.
Independence of transformations
The two transformations are independent and can be applied separately or together. Any application of DeepSMILES should specify which transformations were applied.
Roundtrip Validation on ChEMBL 23
The authors validated DeepSMILES by roundtripping all entries in the ChEMBL 23 database through SMILES-to-DeepSMILES-to-SMILES conversion. Canonical SMILES (including stereochemistry) were generated by four independent cheminformatics toolkits: CDK, OEChem, Open Babel, and RDKit. Using multiple toolkits ensures coverage of different traversal orders and ring closure ordering conventions.
All SMILES strings roundtripped without error across all three configurations (branches only, rings only, both). The exact string representation may differ in ring closure digit assignment or digit ordering, sometimes with an associated stereo inversion at tetrahedral centers, but the canonical SMILES of the original and roundtripped molecules are identical.
Performance characteristics
The following table shows the effect of DeepSMILES conversion on string length and throughput, measured on canonical SMILES from Open Babel for ChEMBL 23:
| Transformation | Mean % change in length | Encoding (per sec) | Decoding (per sec) |
|---|---|---|---|
| Branches only | +8.2% | 32,000 | 16,000 |
| Rings only | -6.4% | 26,000 | 24,000 |
| Both | +1.9% | 26,000 | 17,500 |
The ring transformation slightly shortens strings (by removing one digit per ring), while the branch transformation slightly lengthens them (additional close parentheses). Combined, the net effect is a small increase of about 2%. Throughput is in the tens of thousands of conversions per second in pure Python.
Limitations and Future Directions
DeepSMILES does not eliminate all invalid strings. Invalid DeepSMILES can still be generated, for example when there are more close parentheses than atoms on the stack, or when a ring size exceeds the number of available atoms. The reference implementation raises a DecodeError in these cases, though the authors note that a more tolerant decoder (ignoring extra parentheses or defaulting to the first atom for oversized rings) could be used during generation.
The paper assumes that input SMILES are generated by a standard cheminformatics toolkit as a depth-first traversal of the molecular graph. Non-standard SMILES (e.g., CC(C1)CCCC1) cannot be directly encoded.
The authors suggest several directions for future work:
- Investigating whether a preferred traversal order (e.g., shorter branches first) would make DeepSMILES even easier for models to learn.
- Exploring notations where atoms in the organic subset explicitly list their hydrogen count, which would allow a fully parenthesis-free representation.
- Using SMILES augmentation with random traversal orders (as explored by Bjerrum and Threlfall, 2017) in combination with DeepSMILES.
- Designing entirely new line notations optimized for ML, where every string maps to a valid molecule, there are few duplicate representations, small string changes produce small structural changes, and string length correlates with pharmaceutical relevance.
The fused ring case presents additional complexity: a bicyclic system has three cycles, and depending on traversal order, the ring size digit may not directly correspond to the ring size of any individual ring. This is an inherent limitation of depth-first traversal-based notations.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Validation | ChEMBL 23 | ~1.7M compounds | Canonical SMILES from CDK, OEChem, Open Babel, RDKit |
Algorithms
The DeepSMILES encoder and decoder are pure string-processing algorithms with no machine-learning components. The transformations operate on SMILES syntax tokens (atoms, bonds, parentheses, ring closure digits) without chemical interpretation.
Evaluation
| Metric | Value | Notes |
|---|---|---|
| Roundtrip accuracy | 100% | All ChEMBL 23 entries across 4 toolkits |
| Encoding throughput | 26,000-32,000/s | Pure Python, varies by transformation |
| Decoding throughput | 16,000-24,000/s | Pure Python, varies by transformation |
Hardware
No specific hardware requirements. The implementation is a pure Python module with no GPU dependencies.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| deepsmiles | Code | MIT | Pure Python encoder/decoder |
Paper Information
Citation: O’Boyle, N. M., & Dalke, A. (2018). DeepSMILES: An Adaptation of SMILES for Use in Machine-Learning of Chemical Structures. ChemRxiv. https://doi.org/10.26434/chemrxiv.7097960.v1
@article{oboyle2018deepsmiles,
title={DeepSMILES: An Adaptation of SMILES for Use in Machine-Learning of Chemical Structures},
author={O'Boyle, Noel M. and Dalke, Andrew},
journal={ChemRxiv},
year={2018},
doi={10.26434/chemrxiv.7097960.v1}
}