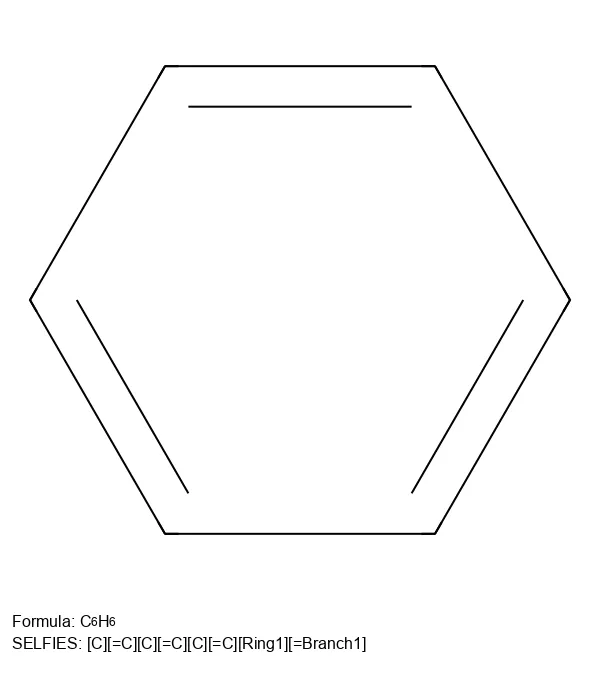
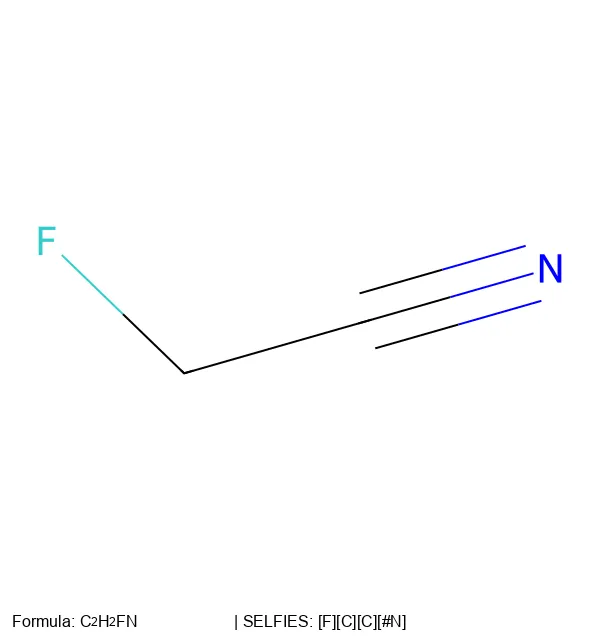
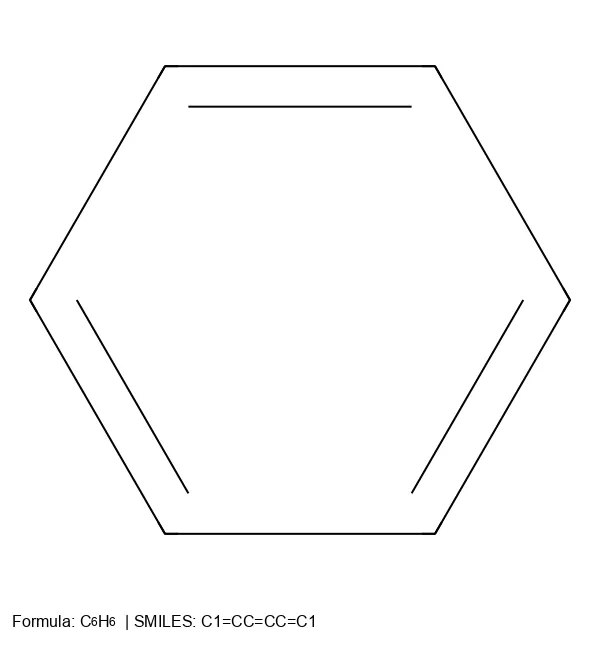
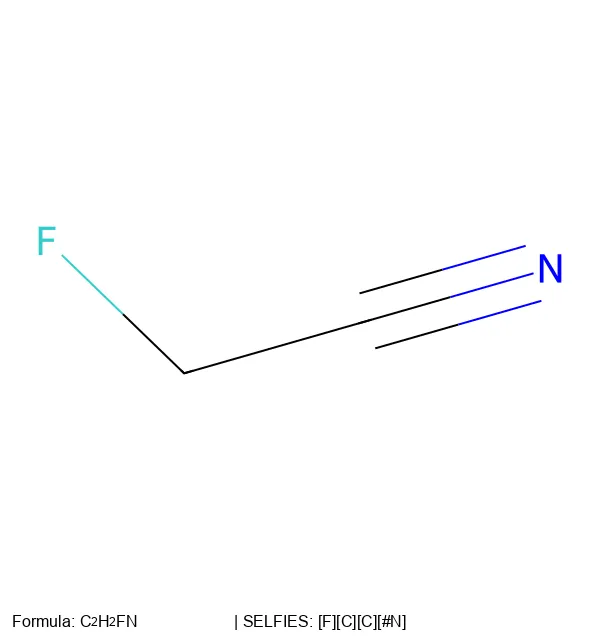
How a molecule is encoded as a string or graph determines what a model can learn from it. This section traces the evolution of molecular string representations: from the original SMILES notation and InChI identifiers, through the SELFIES format designed to guarantee every generated string maps to a valid molecule, to newer extensions like RInChI for reactions and MInChI for mixtures. Notes cover both the technical specifications of each format and the practical tradeoffs (validity, expressiveness, canonicality) that matter when choosing a representation for machine learning.
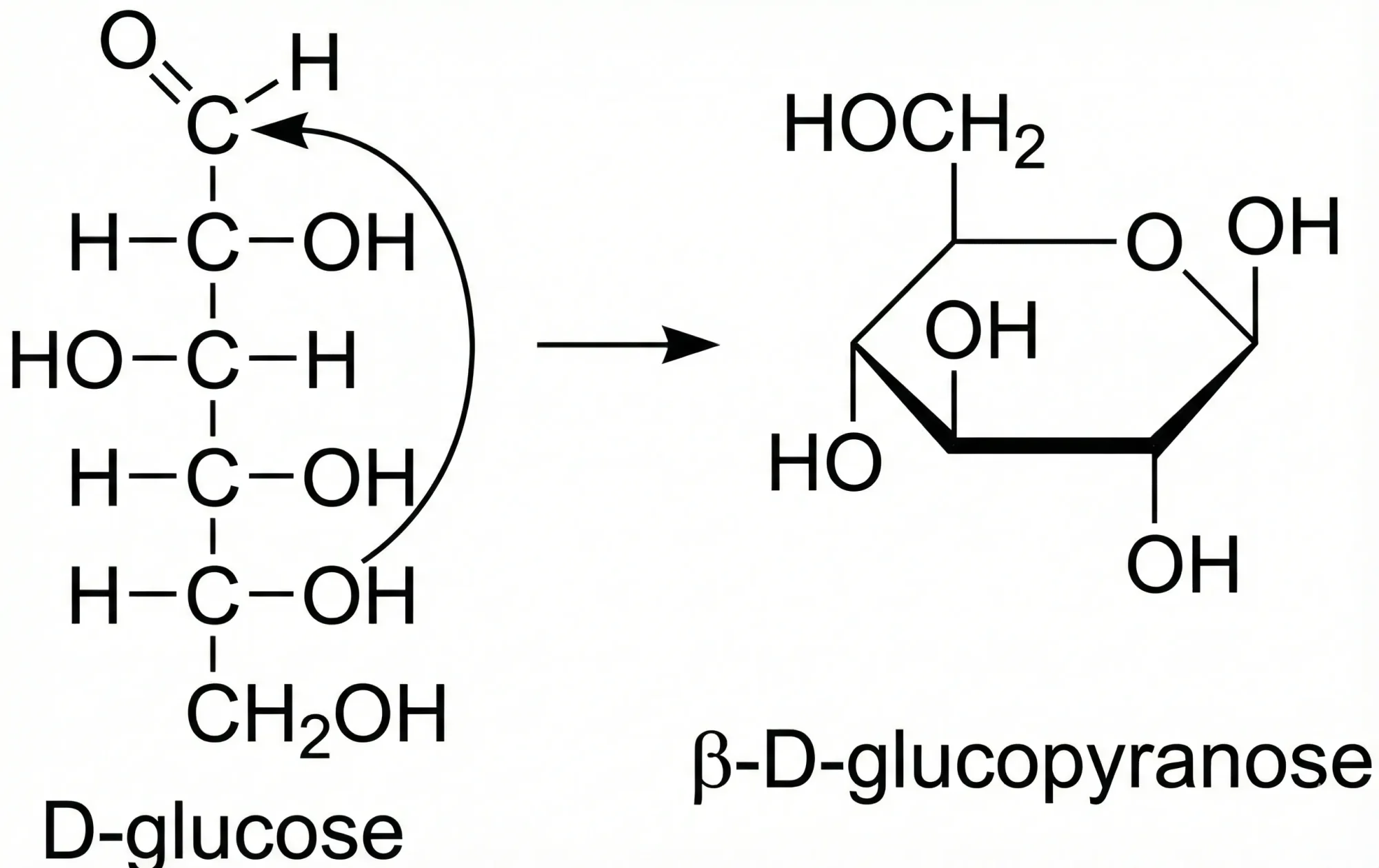
InChI and Tautomerism: Toward Comprehensive Treatment
A comprehensive 2020 analysis of the tautomerism problem in chemical databases, compiling 86 tautomeric transformation rules (20 existing, 66 new) and validating them across 400M+ structures to inform algorithmic improvements for InChI V2.