A Graph-Augmented Transformer for Molecular Property Prediction
This is a Method paper that proposes the Molecule Attention Transformer (MAT), a Transformer-based architecture adapted for molecular property prediction. The primary contribution is a modified self-attention mechanism that incorporates inter-atomic distances and molecular graph structure alongside the standard query-key attention. Combined with self-supervised pretraining on 2 million molecules from ZINC15, MAT achieves competitive performance across seven diverse molecular property prediction tasks while requiring minimal hyperparameter tuning.
Challenges in Deep Learning for Molecular Properties
Predicting molecular properties is central to drug discovery and materials design, yet deep neural networks have struggled to consistently outperform shallow methods like random forests and SVMs on these tasks. Wu et al. (2018) demonstrated through the MoleculeNet benchmark that graph neural networks do not reliably beat classical models. Two recurring problems compound this:
- Underfitting: Graph neural networks tend to underfit training data, with performance failing to scale with model complexity (Ishiguro et al., 2019).
- Hyperparameter sensitivity: Deep models for molecule property prediction require extensive hyperparameter search (often 500+ configurations) to achieve competitive results, making them impractical for many practitioners.
Concurrent work explored using vanilla Transformers on SMILES string representations of molecules (Honda et al., 2019; Wang et al., 2019), but these approaches discard the explicit structural information encoded in molecular graphs and 3D conformations. The motivation for MAT is to combine the flexibility of the Transformer architecture with domain-specific inductive biases from molecular structure.
Molecule Self-Attention: Combining Attention, Distance, and Graph Structure
The core innovation is the Molecule Self-Attention layer, which replaces standard Transformer self-attention. In a standard Transformer, head $i$ computes:
$$ \mathcal{A}^{(i)} = \rho\left(\frac{\mathbf{Q}_{i} \mathbf{K}_{i}^{T}}{\sqrt{d_{k}}}\right) \mathbf{V}_{i} $$
MAT augments this with two additional information sources. Let $\mathbf{A} \in {0, 1}^{N_{\text{atoms}} \times N_{\text{atoms}}}$ denote the molecular graph adjacency matrix and $\mathbf{D} \in \mathbb{R}^{N_{\text{atoms}} \times N_{\text{atoms}}}$ denote the inter-atomic distance matrix. The modified attention becomes:
$$ \mathcal{A}^{(i)} = \left(\lambda_{a} \rho\left(\frac{\mathbf{Q}_{i} \mathbf{K}_{i}^{T}}{\sqrt{d_{k}}}\right) + \lambda_{d}, g(\mathbf{D}) + \lambda_{g}, \mathbf{A}\right) \mathbf{V}_{i} $$
where $\lambda_{a}$, $\lambda_{d}$, and $\lambda_{g}$ are scalar hyperparameters weighting each component, and $g$ is either a row-wise softmax or an element-wise exponential decay $g(d) = \exp(-d)$.
Key architectural details:
- Atom embedding: Each atom is represented as a 26-dimensional vector encoding atomic identity (one-hot over B, N, C, O, F, P, S, Cl, Br, I, dummy, other), number of heavy neighbors, number of hydrogens, formal charge, ring membership, and aromaticity.
- Dummy node: An artificial disconnected node (distance $10^{6}$ from all atoms) is added to each molecule, allowing the model to “skip” attention heads when no relevant pattern exists, similar to how BERT uses the separation token.
- 3D conformers: Distance matrices are computed from RDKit-generated 3D conformers using the Universal Force Field (UFF).
- Pretraining: Node-level masked atom prediction on 2 million ZINC15 molecules (following Hu et al., 2019), where 15% of atom features are masked and the model predicts them.
Benchmark Evaluation and Ablation Studies
Experimental setup
MAT is evaluated on seven molecular property prediction datasets spanning regression and classification:
| Dataset | Task | Size | Metric | Split |
|---|---|---|---|---|
| FreeSolv | Regression (hydration free energy) | 642 | RMSE | Random |
| ESOL | Regression (log solubility) | 1,128 | RMSE | Random |
| BBBP | Classification (BBB permeability) | 2,039 | ROC AUC | Scaffold |
| Estrogen-alpha | Classification (receptor activity) | 2,398 | ROC AUC | Scaffold |
| Estrogen-beta | Classification (receptor activity) | 1,961 | ROC AUC | Scaffold |
| MetStab-high | Classification (metabolic stability) | 2,127 | ROC AUC | Random |
| MetStab-low | Classification (metabolic stability) | 2,127 | ROC AUC | Random |
Baselines include GCN, Weave, EAGCN, Random Forest (RF), and SVM. Each model receives the same hyperparameter search budget (150 or 500 evaluations). Results are averaged over 6 random train/validation/test splits.
Main results
MAT achieves the best average rank across all seven tasks:
| Model | Avg. Rank (500 budget) | Avg. Rank (150 budget) |
|---|---|---|
| MAT | 2.42 | 2.71 |
| RF | 3.14 | 3.14 |
| SVM | 3.57 | 3.28 |
| GCN | 3.57 | 3.71 |
| Weave | 3.71 | 3.57 |
| EAGCN | 4.14 | 4.14 |
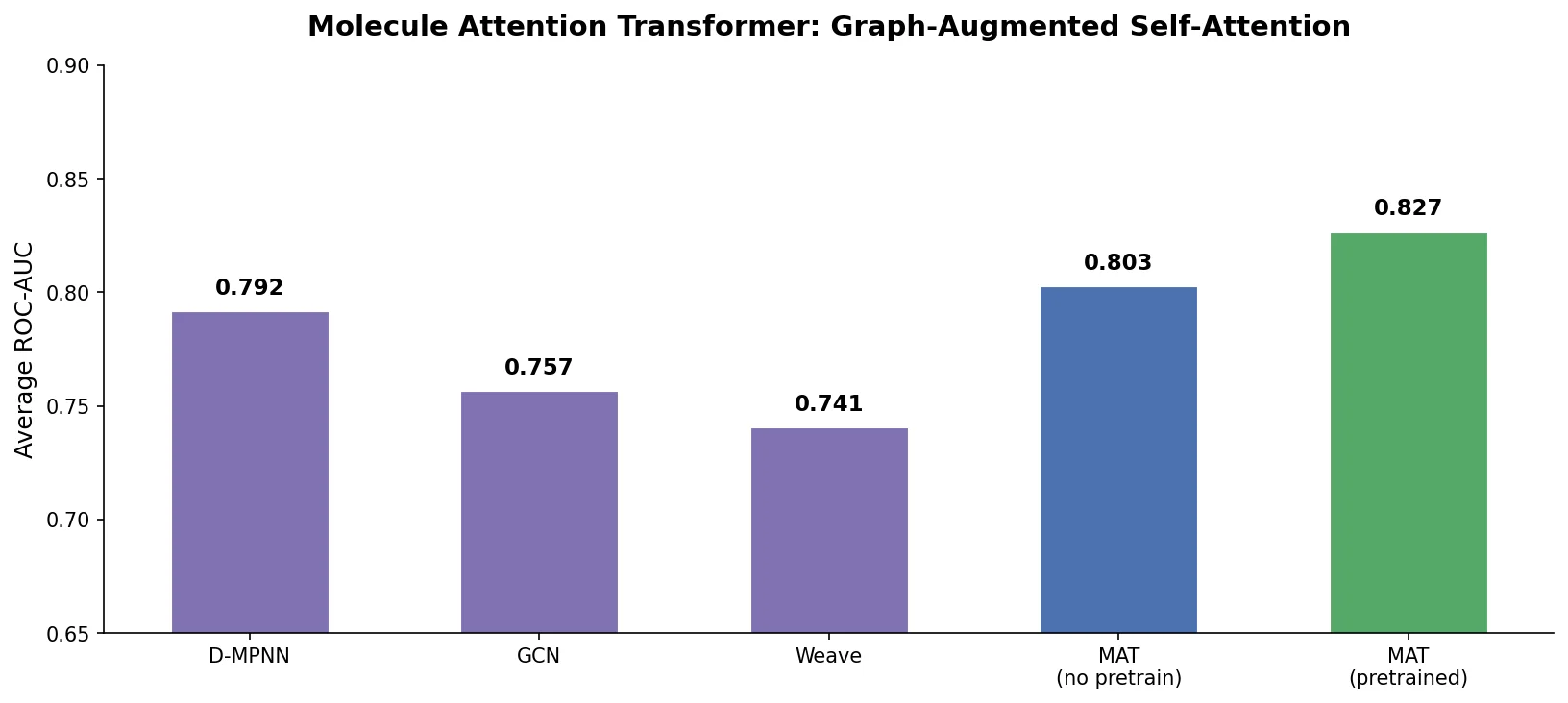
With self-supervised pretraining, Pretrained MAT achieves an average rank of 1.57, outperforming both Pretrained EAGCN (4.0) and SMILES Transformer (4.29). Pretrained MAT requires tuning only the learning rate (7 values tested), compared to 500 hyperparameter combinations for the non-pretrained models.
Ablation results
Ablation studies on BBBP, ESOL, and FreeSolv reveal:
| Variant | BBBP (AUC) | ESOL (RMSE) | FreeSolv (RMSE) |
|---|---|---|---|
| MAT (full) | .723 | .286 | .250 |
| - Graph | .716 | .316 | .276 |
| - Distance | .729 | .281 | .281 |
| - Attention | .692 | .306 | .329 |
| - Dummy node | .714 | .317 | .249 |
| + Edge features | .683 | .314 | .358 |
Removing any single component degrades performance on at least one task, supporting the value of combining all three information sources. Adding edge features does not help, suggesting the adjacency and distance matrices already capture sufficient bond-level information.
Interpretability analysis
Individual attention heads in the first layer learn chemically meaningful functions. Six heads were identified that focus on specific chemical patterns: 2-neighbored aromatic carbons, sulfur atoms, non-ring nitrogens, carbonyl oxygens, 3-neighbored aromatic atoms (substitution positions), and aromatic ring nitrogens. Statistical validation using Kruskal-Wallis tests confirmed that atoms matching these SMARTS patterns receive significantly higher attention weights ($p < 0.001$ for all patterns).
Findings, Limitations, and Future Directions
MAT demonstrates that augmenting Transformer self-attention with molecular graph structure and 3D distance information produces a model that performs consistently well across diverse property prediction tasks. The key practical finding is that self-supervised pretraining dramatically reduces the hyperparameter tuning burden: Pretrained MAT matches or exceeds the performance of extensively tuned models while requiring only learning rate selection.
Several limitations are acknowledged:
- Fingerprint-based models still win on some tasks: RF and SVM with extended-connectivity fingerprints outperform MAT on metabolic stability and Estrogen-beta tasks, suggesting that incorporating fingerprint representations could improve MAT further.
- Single conformer: Only one pre-computed 3D conformer is used per molecule. More sophisticated conformer sampling or ensemble strategies were not explored.
- Limited pretraining exploration: Only the masked atom prediction task from Hu et al. (2019) was used. The authors note that exploring additional pretraining objectives is a promising direction.
- Scalability: The pretrained model uses 1024-dimensional embeddings with 8 layers and 16 attention heads, fitting the largest model that fits in GPU memory.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pretraining | ZINC15 | 2M molecules | Sampled from ZINC database |
| Evaluation | FreeSolv | 642 | Hydration free energy regression |
| Evaluation | ESOL | 1,128 | Log solubility regression |
| Evaluation | BBBP | 2,039 | Blood-brain barrier classification |
| Evaluation | Estrogen-alpha/beta | 2,398 / 1,961 | Receptor activity classification |
| Evaluation | MetStab-high/low | 2,127 each | Metabolic stability classification |
Algorithms
- Optimizer: Adam with Noam learning rate scheduler (warmup then inverse square root decay)
- Pretraining: 8 epochs, learning rate 0.001, batch size 256, binary cross-entropy loss
- Fine-tuning: 100 epochs, batch size 32, learning rate selected from {1e-3, 5e-4, 1e-4, 5e-5, 1e-5, 5e-6, 1e-6}
- Distance kernel: exponential decay $g(d) = \exp(-d)$ for pretrained model
- Lambda weights: $\lambda_{a} = \lambda_{d} = 0.33$ for pretrained model
Models
- Pretrained MAT: 1024-dim embeddings, 8 layers, 16 attention heads, 1 feed-forward layer per block
- Dropout: 0.0, weight decay: 0.0 for pretrained model
- Atom featurization: 26-dimensional one-hot encoding (Table 1 in paper)
Evaluation
- Regression: RMSE (FreeSolv, ESOL)
- Classification: ROC AUC (BBBP, Estrogen-alpha/beta, MetStab-high/low)
- All experiments repeated 6 times with different train/validation/test splits
- Scaffold split for BBBP, Estrogen, random split for others
Hardware
The paper does not specify exact hardware details. The pretrained model is described as “the largest model that still fits the GPU memory.”
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| gmum/MAT | Code | MIT | Official implementation with pretrained weights |
Paper Information
Citation: Maziarka, Ł., Danel, T., Mucha, S., Rataj, K., Tabor, J., & Jastrzębski, S. (2020). Molecule Attention Transformer. arXiv preprint arXiv:2002.08264.
@article{maziarka2020molecule,
title={Molecule Attention Transformer},
author={Maziarka, {\L}ukasz and Danel, Tomasz and Mucha, S{\l}awomir and Rataj, Krzysztof and Tabor, Jacek and Jastrz{\k{e}}bski, Stanis{\l}aw},
journal={arXiv preprint arXiv:2002.08264},
year={2020}
}