Paper Information
Citation: Tripp, A., & Hernández-Lobato, J. M. (2023). Genetic algorithms are strong baselines for molecule generation. arXiv preprint arXiv:2310.09267. https://arxiv.org/abs/2310.09267
Publication: arXiv preprint, 2023
Additional Resources:
A Position Paper on Molecular Generation Baselines
This is a Position paper that argues genetic algorithms (GAs) are underused and underappreciated as baselines in the molecular generation community. The primary contribution is empirical evidence that a simple GA implementation (MOL_GA) matches or outperforms many sophisticated deep learning methods on standard benchmarks. The authors propose the “GA criterion” as a minimum bar for evaluating new molecular generation algorithms.
Why Molecular Generation May Be Easier Than Assumed
Drug discovery is fundamentally a molecular generation task, and many machine learning methods have been proposed for it (Du et al., 2022). The problem has many variants, from unconditional generation of novel molecules to directed optimization of specific molecular properties.
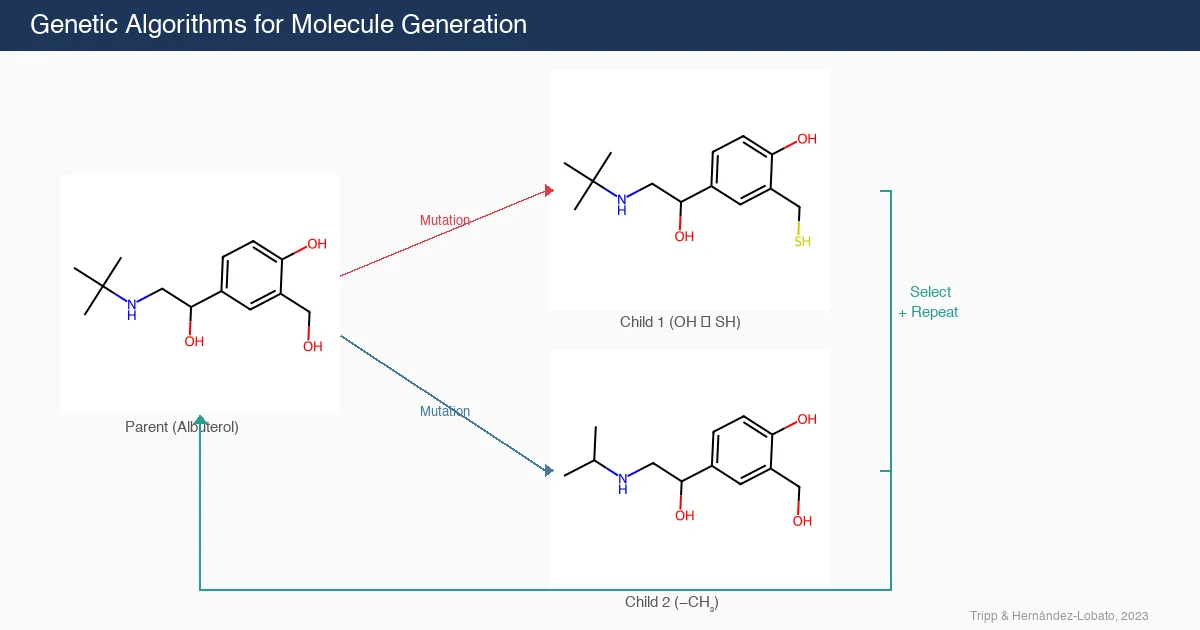
The authors observe that generating valid molecules is, in some respects, straightforward. The rules governing molecular validity are well-defined bond constraints that can be checked using standard cheminformatics software like RDKit. This means new molecules can be generated simply by adding, removing, or substituting fragments of known molecules. When applied iteratively, this is exactly what a genetic algorithm does. Despite this, many papers in the field propose complex deep learning methods without adequately comparing to simple GA baselines.
The GA Criterion for Evaluating New Methods
The core proposal is the GA criterion: new methods in molecular generation should offer some clear advantage over genetic algorithms. This advantage can be:
- Empirical: outperforming GAs on relevant benchmarks
- Conceptual: identifying and overcoming a specific limitation of randomly modifying known molecules
The authors argue that the current state of molecular generation research reflects poor empirical practices, where comprehensive baseline evaluation is treated as optional rather than essential.
Genetic Algorithm Framework and Benchmark Experiments
How Genetic Algorithms Work for Molecules
GAs operate through the following iterative procedure:
- Start with an initial population $P$ of molecules
- Sample a subset $S \subseteq P$ from the population (possibly biased toward better molecules)
- Generate new molecules $N$ from $S$ via mutation and crossover operations
- Select a new population $P’$ from $P \cup N$ (e.g., keep the highest-scoring molecules)
- Set $P \leftarrow P’$ and repeat from step 2
The MOL_GA implementation uses:
- Quantile-based sampling (step 2): molecules are sampled from the top quantiles of the population using a log-uniform distribution over quantile thresholds:
$$ u \sim \mathcal{U}[-3, 0], \quad \epsilon = 10^{u} $$
A molecule is drawn uniformly from the top $\epsilon$ fraction of the population.
- Mutation and crossover (step 3): graph-based operations from Jensen (2019), as implemented in the GuacaMol benchmark (Brown et al., 2019)
- Greedy population selection (step 4): molecules with the highest scores are retained
Unconditional Generation on ZINC 250K
The first experiment evaluates unconditional molecule generation, where the task is to produce novel, valid, and unique molecules distinct from a reference set (ZINC 250K). Success is measured by validity, novelty (at 10,000 generated molecules), and uniqueness.
| Method | Paper | Validity | Novelty@10k | Uniqueness |
|---|---|---|---|---|
| JT-VAE | Jin et al. (2018) | 99.8% | 100% | 100% |
| GCPN | You et al. (2018) | 100% | 100% | 99.97% |
| MolecularRNN | Popova et al. (2019) | 100% | 100% | 99.89% |
| Graph NVP | Madhawa et al. (2019) | 100% | 100% | 94.80% |
| Graph AF | Shi et al. (2020) | 100% | 100% | 99.10% |
| MoFlow | Zang and Wang (2020) | 100% | 100% | 99.99% |
| GraphCNF | Lippe and Gavves (2020) | 96.35% | 99.98% | 99.98% |
| Graph DF | Luo et al. (2021) | 100% | 100% | 99.16% |
| ModFlow | Verma et al. (2022) | 98.1% | 100% | 99.3% |
| GraphEBM | Liu et al. (2021) | 99.96% | 100% | 98.79% |
| AddCarbon | Renz et al. (2019) | 100% | 99.94% | 99.86% |
| MOL_GA | (this paper) | 99.76% | 99.94% | 98.60% |
All methods perform near 100% on all metrics, demonstrating that unconditional molecule generation is not a particularly discriminative benchmark. The authors note that generation speed (molecules per second) is an important missing dimension from these comparisons, where simple methods like GAs have a clear advantage.
Molecule Optimization on the PMO Benchmark
The second experiment evaluates directed molecule optimization on the Practical Molecular Optimization (PMO) benchmark (Gao et al., 2022), which measures the ability to find molecules optimizing a scalar objective function $f: \mathcal{M} \mapsto \mathbb{R}$ with a budget of 10,000 evaluations.
A key insight is that previous GA implementations in PMO used large generation sizes ($\approx 100$), which limits the number of improvement iterations. The authors set the generation size to 5, allowing approximately 2,000 iterations of improvement within the same evaluation budget.
| Task | REINVENT | Graph GA | MOL_GA |
|---|---|---|---|
| albuterol_similarity | 0.882 +/- 0.006 | 0.838 +/- 0.016 | 0.896 +/- 0.035 |
| amlodipine_mpo | 0.635 +/- 0.035 | 0.661 +/- 0.020 | 0.688 +/- 0.039 |
| celecoxib_rediscovery | 0.713 +/- 0.067 | 0.630 +/- 0.097 | 0.567 +/- 0.083 |
| drd2 | 0.945 +/- 0.007 | 0.964 +/- 0.012 | 0.936 +/- 0.016 |
| fexofenadine_mpo | 0.784 +/- 0.006 | 0.760 +/- 0.011 | 0.825 +/- 0.019 |
| isomers_c9h10n2o2pf2cl | 0.642 +/- 0.054 | 0.719 +/- 0.047 | 0.865 +/- 0.012 |
| sitagliptin_mpo | 0.021 +/- 0.003 | 0.433 +/- 0.075 | 0.582 +/- 0.040 |
| zaleplon_mpo | 0.358 +/- 0.062 | 0.346 +/- 0.032 | 0.519 +/- 0.029 |
| Sum (23 tasks) | 14.196 | 13.751 | 14.708 |
| Rank | 2 | 3 | 1 |
MOL_GA achieves the highest aggregate score across all 23 PMO tasks, outperforming both the previous best GA (Graph GA) and the previous best overall method (REINVENT). The authors attribute this partly to the tuning of the baselines in PMO rather than MOL_GA being an especially strong method, since MOL_GA is essentially the same algorithm as Graph GA with different hyperparameters.
Implications for Molecular Generation Research
The key findings and arguments are:
GAs match or outperform deep learning methods on standard molecular generation benchmarks, both for unconditional generation and directed optimization.
Hyperparameter choices matter significantly: MOL_GA’s strong performance on PMO comes partly from using a smaller generation size (5 vs. ~100), which allows more iterations of refinement within the same evaluation budget.
The GA criterion should be enforced in peer review: new molecular generation methods should demonstrate a clear advantage over GAs, whether empirical or conceptual.
Deep learning methods may implicitly do what GAs do explicitly: many generative models are trained on datasets of known molecules, so the novel molecules they produce may simply be variants of their training data. The authors consider this an important direction for future investigation.
Poor empirical practices are widespread: the paper argues that many experiments in molecule generation are conducted with an explicit desired outcome (that the novel algorithm is the best), leading to inadequate baseline comparisons.
The authors are careful to note that this result should not be interpreted as GAs being exceptional algorithms. Rather, it is an indication that more complex methods have made surprisingly little progress beyond what simple heuristic search can achieve.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Unconditional generation | ZINC 250K | 250,000 molecules | Reference set for novelty evaluation |
| Directed optimization | PMO benchmark | 23 tasks | 10,000 evaluation budget per task |
Algorithms
- GA implementation: MOL_GA package, using graph-based mutation and crossover from Jensen (2019) via the GuacaMol implementation
- Generation size: 5 molecules per iteration (allowing ~2,000 iterations with 10,000 evaluations)
- Population selection: Greedy (highest-scoring molecules retained)
- Sampling: Quantile-based with log-uniform distribution over quantile thresholds
Evaluation
| Metric | Benchmark | Notes |
|---|---|---|
| Validity, Novelty@10k, Uniqueness | ZINC 250K unconditional | Calculated using MOSES package |
| AUC top-10 scores | PMO benchmark | 23 optimization tasks with 10,000 evaluation budget |
Hardware
The paper does not specify hardware requirements. Given that GAs are computationally lightweight compared to deep learning methods, standard CPU hardware is likely sufficient.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| MOL_GA | Code | MIT | Python package for molecular genetic algorithms |
| MOL_GA on PyPI | Code | MIT | pip-installable package |
Citation
@article{tripp2023genetic,
title={Genetic algorithms are strong baselines for molecule generation},
author={Tripp, Austin and Hern{\'a}ndez-Lobato, Jos{\'e} Miguel},
journal={arXiv preprint arXiv:2310.09267},
year={2023}
}