A Fourier-Space Long-Range Correction for Molecular GNNs
This is a Method paper that introduces Ewald message passing (Ewald MP), a general framework for incorporating long-range interactions into message passing neural networks (MPNNs) for molecular potential energy surface prediction. The key contribution is a nonlocal Fourier-space message passing scheme, grounded in the classical Ewald summation technique from computational physics, that complements the short-range message passing of existing GNN architectures.
The Long-Range Interaction Problem in Molecular GNNs
Standard MPNNs for molecular property prediction rely on a spatial distance cutoff to define atomic neighborhoods. While this locality assumption enables favorable scaling with system size and provides a useful inductive bias, it fundamentally limits the model’s ability to capture long-range interactions such as electrostatic forces and van der Waals (London dispersion) interactions. These interactions decay slowly with distance (e.g., electrostatic energy follows a $1/r$ power law), and truncating them with a distance cutoff can introduce severe artifacts in thermochemical predictions.
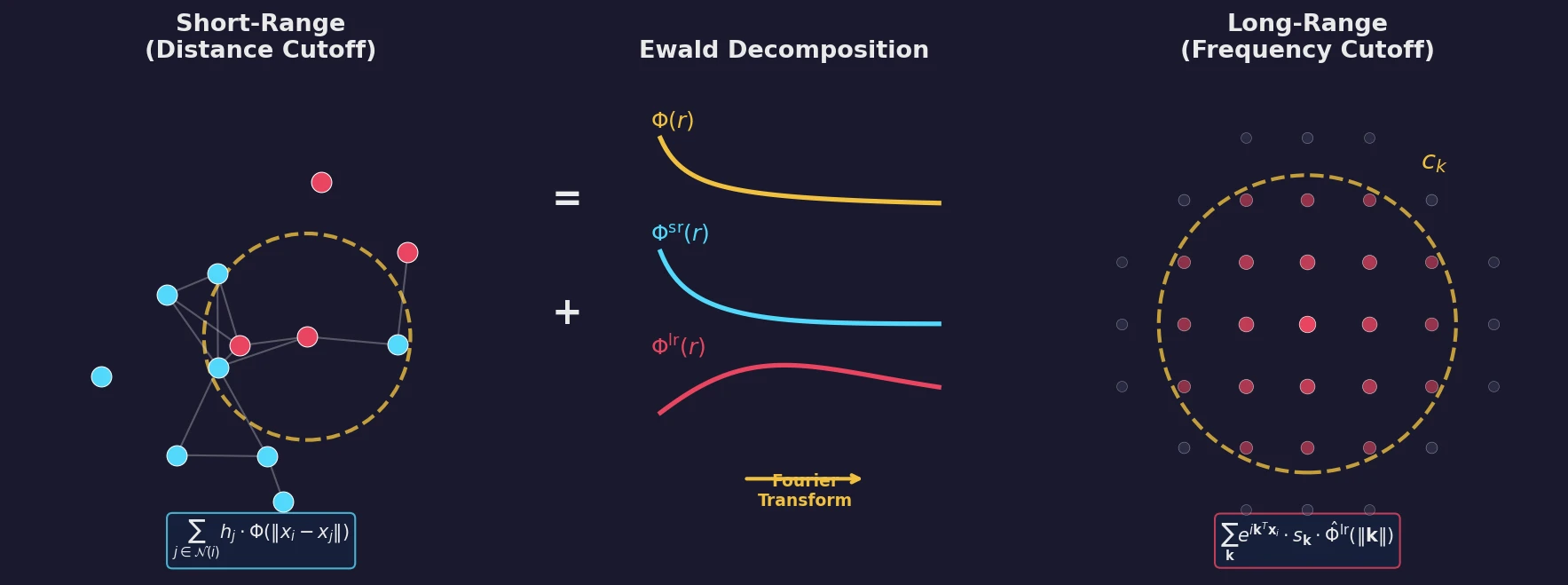
This problem is well-known in molecular dynamics, where empirical force fields explicitly separate bonded (short-range) and non-bonded (long-range) energy terms. The Ewald summation technique addresses this by decomposing interactions into a short-range part that converges quickly with a distance cutoff and a long-range part whose Fourier transform converges quickly with a frequency cutoff. The authors propose bringing this same strategy into the GNN paradigm.
From Ewald Summation to Learnable Fourier-Space Messages
The core insight is a formal analogy between the continuous-filter convolution used in MPNNs and the electrostatic potential computation in Ewald summation. In a standard continuous-filter convolution, the message sum for atom $i$ is:
$$ M_i^{(l+1)} = \sum_{j \in \mathcal{N}(i)} h_j^{(l)} \cdot \Phi^{(l)}(| \mathbf{x}_i - \mathbf{x}_j |) $$
where $h_j^{(l)}$ are atom embeddings and $\Phi^{(l)}$ is a learned radial filter. Comparing this to the electrostatic potential $V_i^{\text{es}}(\mathbf{x}_i) = \sum_{j \neq i} q_j \cdot \Phi^{\text{es}}(| \mathbf{x}_i - \mathbf{x}_j |)$ reveals a direct correspondence: atom embeddings play the role of partial charges, and learned filters replace the $1/r$ kernel.
Ewald MP decomposes the learned filter into short-range and long-range components. The short-range part is handled by any existing GNN architecture with a distance cutoff. The long-range part is computed as a sum over Fourier frequencies:
$$ M^{\text{lr}}(\mathbf{x}_i) = \sum_{\mathbf{k}} \exp(i \mathbf{k}^T \mathbf{x}_i) \cdot s_{\mathbf{k}} \cdot \hat{\Phi}^{\text{lr}}(| \mathbf{k} |) $$
where $s_{\mathbf{k}}$ are structure factor embeddings, computed as:
$$ s_{\mathbf{k}} = \sum_{j \in \mathcal{S}} h_j \exp(-i \mathbf{k}^T \mathbf{x}_j) $$
These structure factor embeddings are a Fourier-space representation of the atom embedding distribution, and truncating to low frequencies effectively coarse-grains the hidden model state while preserving long-range information. The frequency filters $\hat{\Phi}^{\text{lr}}$ are learned, making the entire scheme data-driven rather than tied to a fixed physical functional form.
The method handles both periodic systems (where the reciprocal lattice provides a natural frequency discretization) and aperiodic systems (where the Fourier domain is discretized using a cubic voxel grid with SVD-based rotation alignment to preserve rotation invariance). The combined embedding update becomes:
$$ h_i^{(l+1)} = \frac{1}{\sqrt{3}} \left[ h_i^{(l)} + f_{\text{upd}}^{\text{sr}}(M_i^{\text{sr}}) + f_{\text{upd}}^{\text{lr}}(M_i^{\text{lr}}) \right] $$
The computational complexity is $\mathcal{O}(N_{\text{at}} N_{\text{k}})$, and by fixing the number of frequency vectors $N_{\text{k}}$, linear scaling $\mathcal{O}(N_{\text{at}})$ is achievable.
Experiments Across Four GNN Architectures and Two Datasets
The authors test Ewald MP as an augmentation on four baseline architectures: SchNet, PaiNN, DimeNet++, and GemNet-T. Two datasets are used:
- OC20 (Chanussot et al., 2021): ~265M periodic structures of adsorbate-catalyst systems with DFT-computed energies and forces. The OC20-2M subsplit is used for training.
- OE62 (Stuke et al., 2020): ~62,000 large aperiodic organic molecules with DFT-computed energies that include a DFT-D3 dispersion correction for London dispersion interactions.
All baselines use a 6 Å distance cutoff and 50 maximum neighbors. The Ewald modification is minimal: the long-range message sum is added as an additional skip connection term in each interaction block. Comparison studies include: (1) increasing the distance cutoff to match the computational cost of Ewald MP, (2) replacing the Ewald block with a SchNet interaction block at increased cutoff, and (3) increasing atom embedding dimensions to match Ewald MP’s parameter count.
Key Energy MAE Results on OE62
| Model | Baseline (meV) | Ewald MP (meV) | Improvement |
|---|---|---|---|
| SchNet | 133.5 | 79.2 | 40.7% |
| PaiNN | 61.4 | 57.9 | 5.7% |
| DimeNet++ | 51.2 | 46.5 | 9.2% |
| GemNet-T | 51.5 | 47.4 | 8.0% |
Key Energy MAE Results on OC20 (Averaged Across Test Splits)
| Model | Baseline (meV) | Ewald MP (meV) | Improvement |
|---|---|---|---|
| SchNet | 895 | 830 | 7.3% |
| PaiNN | 448 | 393 | 12.3% |
| DimeNet++ | 496 | 445 | 10.4% |
| GemNet-T | 346 | 307 | 11.3% |
Robust Long-Range Improvements and Dispersion Recovery
Ewald MP achieves consistent improvements across all models and both datasets, averaging 16.1% on OE62 and 10.3% on OC20. Several findings stand out:
Robustness: Unlike the increased-cutoff and SchNet-LR alternatives, Ewald MP never produces detrimental effects in any tested configuration. The increased cutoff setting hurts SchNet and PaiNN on OE62, and the SchNet-LR block fails to improve DimeNet++ and GemNet-T.
Long-range specificity: A binning analysis on OE62 groups molecules by the magnitude of their DFT-D3 dispersion correction. Ewald MP shows an outsize improvement for structures with large long-range energy contributions. It recovers or surpasses a “cheating” baseline that receives the exact DFT-D3 ground truth as an additional input.
Efficiency on periodic systems: Ewald MP achieves similar relative improvements on OC20 at roughly half the relative computational cost compared to OE62, suggesting periodic structures as a particularly attractive application domain.
Force predictions: Improvements in force MAEs are consistent but small, which is expected since the frequency truncation removes high-frequency contributions to the potential energy surface.
Ablation studies: Results are robust across different frequency cutoffs, voxel resolutions, and filtering strategies, with the non-radial periodic filtering scheme outperforming radial alternatives on out-of-distribution generalization.
Limitations include the current focus on scalar (invariant) embeddings only (PaiNN’s equivariant vector embeddings are not augmented), and the potential for a “gap” of medium-range interactions when $N_{\text{k}}$ is fixed for linear scaling. The authors suggest adapting more efficient Ewald summation variants (e.g., particle mesh Ewald with $\mathcal{O}(N \log N)$ scaling) as future work.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training (periodic) | OC20-2M | ~2M structures | Subsplit of OC20; PBC; DFT energies and forces |
| Training (aperiodic) | OE62 | ~62,000 molecules | Large organic molecules; DFT energies with D3 correction |
| Evaluation | OC20-test (4 splits: ID, OOD-ads, OOD-cat, OOD-both) | Varies | Evaluated via submission to OC20 evaluation server |
| Evaluation | OE62-val, OE62-test | ~6,000 each | Direct evaluation |
Algorithms
- Ewald message passing is integrated as an additional skip connection term in each interaction block
- For periodic systems: non-radial filtering with fixed reciprocal lattice positions ($N_x, N_y, N_z$ hyperparameters)
- For aperiodic systems: radial Gaussian basis function filtering with frequency cutoff $c_k$ and voxel resolution $\Delta = 0.2$ Å$^{-1}$
- SVD-based coordinate alignment for rotation invariance in the aperiodic case
- Bottleneck dimension $N_\downarrow = 16$ (GemNet-T) or $N_\downarrow = 8$ (others)
- Update function: dense layer + $N_{\text{hidden}}$ residual layers ($N_{\text{hidden}} = 3$, except PaiNN with $N_{\text{hidden}} = 0$)
Models
| Model | Embedding Size (OE62) | Interaction Blocks | Ewald Params (OE62) |
|---|---|---|---|
| SchNet | 512 | 4 | 12.2M total |
| PaiNN | 512 | 4 | 15.7M total |
| DimeNet++ | 256 | 3 | 4.8M total |
| GemNet-T | 256 | 3 | 16.1M total |
Evaluation
- Primary metric: Energy mean absolute error (EMAE) in meV
- Secondary metric: Force MAE in meV/Å (OC20 only)
- Loss: Linear combination of energy and force MAEs (Eq. 15) with model-specific force multipliers
- Optimizer: Adam with weight decay ($\lambda = 0.01$)
Hardware
- All runtime measurements on NVIDIA A100 GPUs
- Runtimes measured after 50 warmup batches, averaged over 500 batches, minimum of 3 repetitions
- Code: EwaldMP (Hippocratic License 3.0)
Paper Information
Citation: Kosmala, A., Gasteiger, J., Gao, N., & Günnemann, S. (2023). Ewald-based Long-Range Message Passing for Molecular Graphs. In Proceedings of the 40th International Conference on Machine Learning (ICML 2023).
Publication: ICML 2023
@inproceedings{kosmala2023ewald,
title={Ewald-based Long-Range Message Passing for Molecular Graphs},
author={Kosmala, Arthur and Gasteiger, Johannes and Gao, Nicholas and G{\"u}nnemann, Stephan},
booktitle={Proceedings of the 40th International Conference on Machine Learning},
year={2023},
series={PMLR},
volume={202}
}