A Perspective on Code-Generating LLMs for Chemistry
This is a Position paper that argues large language models (LLMs) capable of generating code from natural language prompts, specifically OpenAI’s Codex and GPT-3, are poised to transform both chemistry research and chemistry education. Published in the inaugural volume of Digital Discovery (RSC), the paper combines a brief history of NLP developments with concrete demonstrations of code generation for computational chemistry tasks, then offers a forward-looking perspective on challenges and opportunities.
Bridging the Gap Between Natural Language and Scientific Software
The authors identify a core friction in modern computational chemistry: while the number of available software packages has grown dramatically, researchers spend a large fraction of their time learning interfaces to these packages rather than doing science. Tasks like searching documentation, following tutorials, and trial-and-error experimentation with APIs consume effort that could be directed at research itself.
At the same time, programming assignments in chemistry courses serve dual pedagogical purposes (reinforcing physical intuition and teaching marketable skills), but are constrained by students’ median programming experience. The emergence of code-generating NLP models opens the possibility of reducing both barriers simultaneously.
Code Generation as a Chemistry Interface
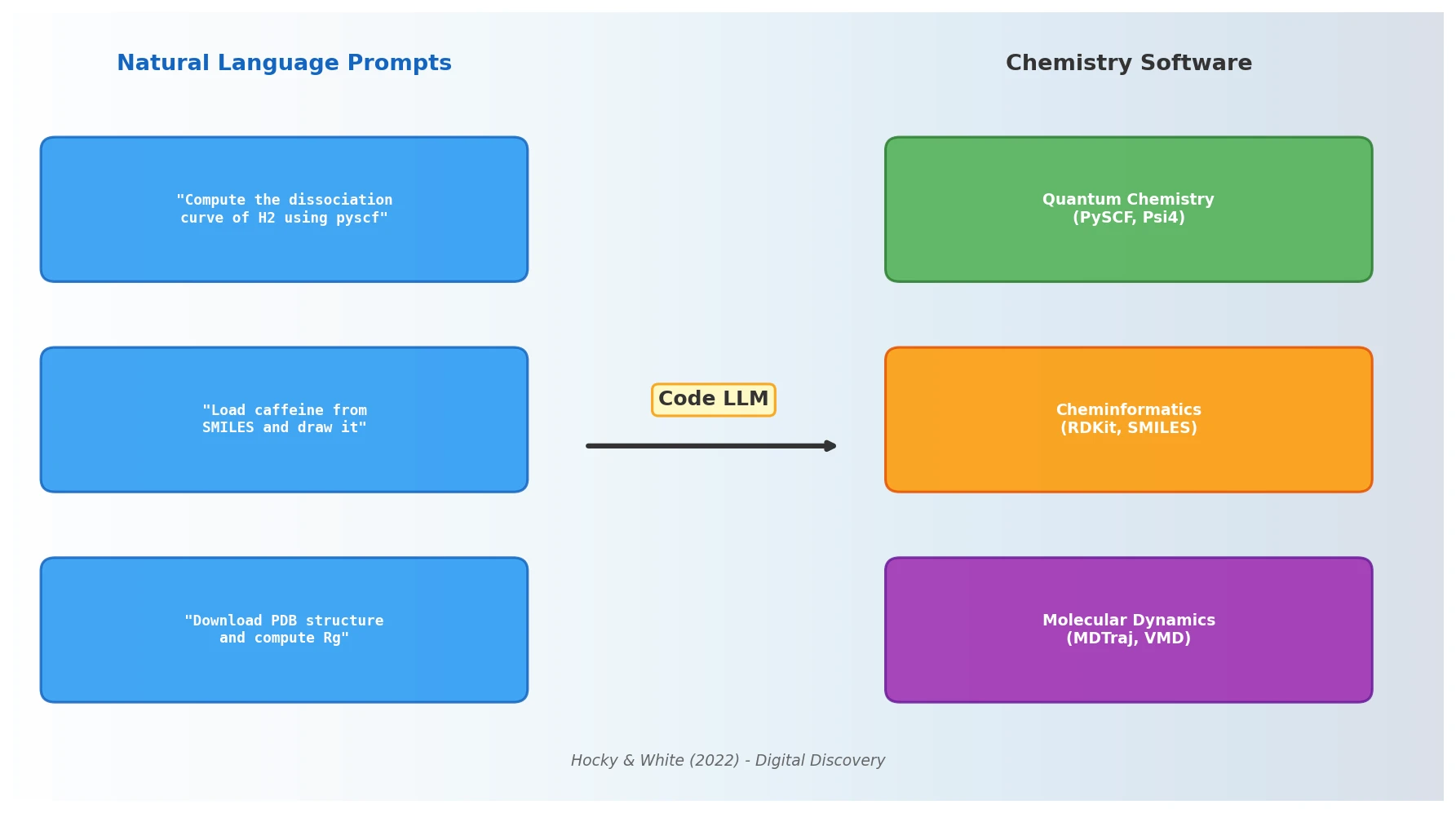
The paper’s core thesis is that NLP models trained on code can serve as a natural language interface to the entire ecosystem of scientific computing tools. The authors demonstrate this with several concrete examples using OpenAI Codex:
Quantum chemistry: Prompting Codex to “compute the dissociation curve of H2 using pyscf” produced correct, runnable code that selected Hartree-Fock with STO-3G. A follow-up prompt requesting “the most accurate method” caused it to switch to CCSD in a large basis set.
Chemical entity recognition: Using GPT-3 with only three training examples, the authors demonstrated extraction of chemical entity names from published text, a task that previously required thousands of labeled examples.
Molecular visualization: Drawing caffeine from its SMILES string, generating Gaussian input files from SMILES, implementing random walks, and downloading and analyzing PDB structures with MDTraj.
Voice-controlled molecular dynamics: The authors previously built MARVIS, a voice-controlled molecular dynamics analysis tool that uses GPT-3 to convert natural language into VMD commands. Only about a dozen examples were needed to teach GPT-3 to render proteins, change representations, and select atoms.
An important caveat: the authors emphasize that all chemistry “knowledge” (including the SMILES string for caffeine) is entirely contained in the model’s learned floating-point weights. The model has no access to databases or curated lists of chemical concepts.
Demonstrations and Practical Evaluation
Rather than a formal experimental evaluation with benchmarks and metrics, this perspective paper relies on qualitative demonstrations. The key examples, with full details provided in the ESI, include:
| Task | Input | Result |
|---|---|---|
| H2 dissociation curve | Natural language prompt | Correct PySCF code (HF/STO-3G) |
| Upgrade method accuracy | Follow-up prompt | Switched to CCSD with large basis |
| Chemical NER | 3 examples + new text | Extracted compound names (with some gaps) |
| Molecule drawing | “Load caffeine from SMILES, draw it” | Correct RDKit rendering |
| Gaussian input file | Function with docstring | Complete file writer with B3LYP/6-31G(d) |
| PDB analysis | Natural language description | Downloaded structure and computed radius of gyration |
The authors note that Codex generates correct code at about a 30% rate on a single attempt for standard problems, improving to above 50% when multiple solutions are tried. Mistakes tend to occur when complex algorithms are requested with little specificity, and the code rarely has syntax errors but may fail in obvious ways (missing imports, wrong data types).
Challenges: Access, Correctness, and Bias
The paper identifies three ongoing challenges:
Access and price. Advanced models from OpenAI were, at the time of writing, limited to early testers. Per-query costs (1-3 cents for GPT-3) would become prohibitive at the scale needed for parsing academic literature or supporting medium-sized courses. The authors advocate for open-source models and equitable deployment by researchers with computational resources.
Correctness. Code generation does not guarantee correctness. The authors raise a subtle point: Codex may produce code that executes successfully but does not follow best scientific practice for a particular computational task. Over-reliance on AI-generated code without verification could erode trust in scientific software. However, they argue that strategies for assessing code correctness apply equally to human-written and AI-generated code.
Fairness and bias. The authors flag several concerns: AI-generated code trained on its own outputs could narrow the range of packages, methods, or programming languages used in chemistry. They observed Codex’s preference for Python and for specific popular libraries (e.g., defaulting to Psi4 for single-point energy calculations). GPT-3 has also been shown to reflect racism, sexism, and other biases present in its training data.
Implications for Research and Education
The authors conclude with an optimistic but measured outlook:
- For research: NLP code generation will increase accessibility of software tools and expand what a single research group can accomplish. Better tools have historically not reduced the need for scientists but expanded the complexity of problems that can be tackled.
- For programming skills: Using Codex will make chemists better programmers, not worse. The process of crafting prompts, mentally checking outputs, testing on sample inputs, and iterating develops algorithmic thinking. The authors report discovering chemistry software libraries they would not have found otherwise through iterative prompt creation.
- For education: Instructors should rethink programming assignments. The authors suggest moving toward more difficult compound assignments, treating code exercises as laboratory explorations of scientific concepts rather than syntax drills, and aligning coursework with the tools students will have access to in their careers.
- For accessibility: NLP models can reduce barriers for non-native English speakers (though accuracy with non-English prompts was not fully explored) and for users who have difficulty with keyboard-and-mouse interfaces (via voice control).
The paper acknowledges that these capabilities were, in early 2022, just beginning, with Codex being the first capable code-generation model. Already at the time of writing, models surpassing GPT-3 in language tasks had appeared, and models matching GPT-3 with 1/20th the parameters had been demonstrated.
Reproducibility Details
This is a perspective paper with qualitative demonstrations rather than a reproducible experimental study. The authors provide all prompts and multiple responses in the ESI.
Data
All prompts and code outputs are provided in the Electronic Supplementary Information (ESI) available from the RSC.
Algorithms
The paper does not introduce new algorithms. It evaluates existing models (GPT-3, Codex) on chemistry-related code generation tasks.
Models
| Model | Provider | Access |
|---|---|---|
| GPT-3 | OpenAI | API access (commercial) |
| Codex | OpenAI | Early tester program (2021) |
| GPT-Neo | EleutherAI | Open source |
Evaluation
No formal metrics are reported for the chemistry demonstrations. The authors cite the Codex paper’s reported ~30% pass rate on single attempts and >50% with multiple attempts on standard programming problems.
Hardware
No hardware requirements are specified for the demonstrations (API-based inference).
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| MARVIS | Code | MIT | Voice-controlled MD analysis using GPT-3 |
Paper Information
Citation: Hocky, G. M., & White, A. D. (2022). Natural language processing models that automate programming will transform chemistry research and teaching. Digital Discovery, 1(2), 79-83. https://doi.org/10.1039/d1dd00009h
@article{hocky2022natural,
title={Natural language processing models that automate programming will transform chemistry research and teaching},
author={Hocky, Glen M. and White, Andrew D.},
journal={Digital Discovery},
volume={1},
number={2},
pages={79--83},
year={2022},
publisher={Royal Society of Chemistry},
doi={10.1039/d1dd00009h}
}