A Benchmark for Multimodal Scientific Reasoning
MaCBench is a Resource contribution that provides a comprehensive benchmark for evaluating vision language models (VLLMs) on real-world chemistry and materials science tasks. Rather than testing general-purpose visual reasoning or text-only scientific knowledge, MaCBench specifically targets the interplay between visual and textual modalities across the scientific workflow. The benchmark contains 779 multiple-choice questions and 374 numeric-answer questions organized into 11 topics across three pillars: data extraction, experimental execution, and data interpretation. Through systematic ablation studies, the authors identify fundamental limitations in spatial reasoning, cross-modal synthesis, and multi-step inference that current VLLMs exhibit.
Why Multimodal Evaluation Matters for Chemistry
Scientific research inherently requires integrating multiple information modalities: reading plots, interpreting spectra, evaluating laboratory setups, and connecting visual observations with domain knowledge. While text-only benchmarks like ChemBench have evaluated LLM capabilities in chemistry, and general multimodal benchmarks have tested visual reasoning, no prior work had systematically assessed how VLLMs handle the specific multimodal demands of the chemistry and materials science workflow.
Existing evaluations treated either the scientific reasoning dimension or the multimodal dimension in isolation. This left a critical gap: can VLLMs reliably assist with tasks that require both visual perception and scientific reasoning simultaneously? For example, identifying laboratory equipment is a perception task, but evaluating whether a laboratory setup is safe requires integrating visual understanding with domain-specific knowledge about hazards.
The authors designed MaCBench to fill this gap by constructing tasks that mirror actual scientific workflows and by including ablation studies that isolate specific failure modes.
Benchmark Design: Three Pillars of Scientific Work
The benchmark is structured around three pillars reflecting the scientific process:
Data Extraction covers parsing scientific literature, including extracting values from tables and plots, interpreting chemical structure diagrams, and identifying reaction components. Tasks range from simple value extraction to complex spatial reasoning about molecular relationships (e.g., identifying isomeric relationships between compounds).
Experimental Execution evaluates understanding of laboratory operations and crystallographic analysis. This includes equipment identification, safety assessment of laboratory setups, and interpretation of crystal structure renderings (space group assignment, atomic species counting, density calculations).
Data Interpretation tests analysis of experimental outputs: spectral analysis (XRD, NMR, mass spectrometry), electronic structure interpretation, adsorption isotherm analysis, and AFM image interpretation.
Each task uses a single prompt template containing multiple questions. All questions pair images with text-based prompts. The dataset was curated manually, with questions reviewed by multiple scientists before inclusion. A BigBench canary string is embedded in each file to prevent data contamination during future model training.
Evaluation of Frontier VLLMs and Ablation Studies
The authors evaluated four frontier VLLMs: Claude 3.5 Sonnet, GPT-4o, Gemini 1.5 Pro, and Llama 3.2 90B Vision. Performance is reported relative to random baselines to account for the varying number of answer choices across MCQ tasks:
$$ \text{acc}_{\text{rel}} = \text{acc} - \text{acc}_{\text{baseline}} $$
Each benchmark run was repeated five times to capture variability, with standard deviations reported as error bars.
Overall Performance Landscape
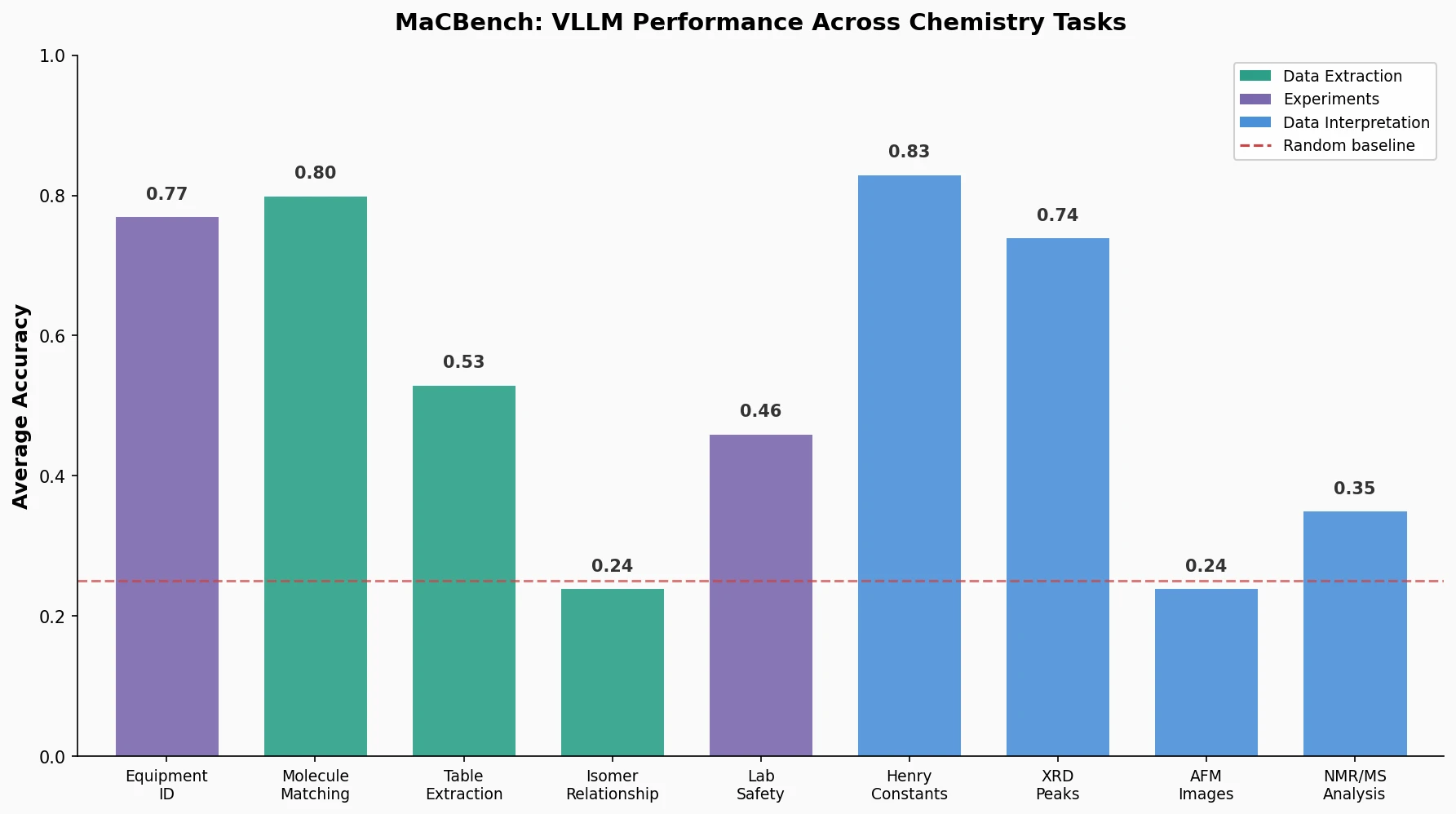
Claude 3.5 Sonnet was the leading model across all three task families, though no model dominated across all individual tasks. Key findings:
- Equipment identification: average accuracy of 0.77 (strong perception performance)
- Hand-drawn molecule to SMILES matching: average accuracy of 0.80
- Table composition extraction: average accuracy of 0.53 (Llama 3.2 indistinguishable from random guessing)
- Isomer relationship identification: average accuracy of 0.24 (barely above the 0.14 baseline)
- Laboratory safety assessment: average accuracy of 0.46
- AFM image interpretation: average accuracy of 0.24
- NMR and mass spectrometry analysis: average accuracy of 0.35
Ablation Studies: Four Dimensions of Failure
The authors designed ablations isolating four specific dimensions:
1. Modality (Image vs. Text): When identical information was presented as text instead of images, performance improved consistently across all tasks. For XRD peak identification, models showed a roughly 35% performance increase when peaks were provided as text rather than displayed visually. Even crystal structure volume calculations differed by four percentage points between visual and textual input of unit cell parameters.
2. Multi-Step Reasoning: Performance degraded consistently as tasks required more reasoning steps. For XRD analysis, identifying the highest peak achieved 0.74 average accuracy, while ranking relative peak intensities dropped to 0.28. Isotherm analysis showed the same pattern: finding the maximum value was easier than ordering multiple values.
3. Scientific Terminology: Removing domain-specific terminology (e.g., using IUPAC names instead of SMILES notation) improved performance on several tasks, suggesting models are sensitive to specific vocabularies rather than understanding underlying concepts. Gemini 1.5 Pro showed particular sensitivity to exact prompt wording, with large performance variations from minor changes like replacing “image” with “diagram” or “plot.”
4. Guidance: Adding step-by-step instructions improved performance for most models on spectral analysis and XRD pattern matching, with the notable exception of Claude 3.5 Sonnet, whose performance did not improve with guidance.
Internet Frequency Correlation
The authors measured the correlation between model performance and the number of Google search results for various crystal structures (as a proxy for training data frequency). For all tested cases, structures with correct model responses had higher Internet presence. This effect held even for pure perception tasks like counting atomic species, suggesting models may rely on memorized patterns rather than genuine visual reasoning.
Limitations of Current VLLMs for Scientific Assistance
The results reveal three fundamental limitations of current VLLMs:
Spatial reasoning failure: Models perform well on perception tasks (identifying equipment, matching hand-drawn molecules) but fail when spatial understanding is required (stereochemistry assignment at 0.24 accuracy, space group identification at 0.45). This limitation undermines one of the most intuitive potential use cases of vision models.
Incomplete cross-modal integration: The consistent performance gap between text and image presentations of identical information demonstrates that current models have not developed robust strategies for visual information processing. The models process text and images through fundamentally different pathways, with text consistently yielding better results.
Multi-step reasoning brittleness: The systematic degradation across reasoning steps indicates that chaining logical operations, a core requirement for scientific reasoning, remains a fundamental weakness.
The authors note that compared to text-only benchmarks (e.g., ChemBench), multimodal systems show much higher performance variability across tasks, suggesting greater fragility. They propose that advances in synthetic training data generation (particularly for spatial reasoning) and modality transformation training tasks could help address these limitations. They also acknowledge that future workflows with machine-actionable data formats may reduce the need for some multimodal parsing capabilities.
The benchmark does not encompass the full scope of scientific reasoning, and the evaluated models are not exhaustive of all available architectures. The authors call for continued research across wider task and model sets, along with interpretability studies to distinguish genuine reasoning from pattern matching.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Evaluation | MaCBench | 779 MCQs + 374 numeric questions | 11 topics across 3 pillars |
| Evaluation | MaCBench-Ablations | Subset with ablation variants | Modality, terminology, guidance, step complexity |
Both datasets are available on HuggingFace. Questions are stored in extended BigBench format with base-64-encoded images and BigBench canary strings.
Algorithms
The evaluation pipeline builds on the ChemBench framework (v0.3.0). Answer extraction uses regex-based parsing backed by an LLM extractor (Claude 3.5 Sonnet) for fallback cases. Refusal detection combines LLM Guard regex patterns with a fine-tuned DistilRoBERTa model, with up to five retries for refused responses.
Scoring:
- MCQs: correct if Hamming loss is zero (exact match)
- Numeric: correct if mean absolute error falls within specified tolerance (default 1%, up to 5% for specific tasks)
- Random baseline: random option selection for MCQs; mean of all target values in a topic for numeric questions
Models
Four frontier VLLMs evaluated:
- Claude 3.5 Sonnet (Anthropic)
- GPT-4o (OpenAI)
- Gemini 1.5 Pro (Google)
- Llama 3.2 90B Vision (Meta)
Default quality/resolution settings were used for each provider.
Evaluation
| Metric | Best Model | Value | Baseline | Notes |
|---|---|---|---|---|
| Equipment identification | Average | 0.77 | varies | Near-ceiling perception |
| Hand-drawn molecule matching | Average | 0.80 | ~0.20 | 4x above baseline |
| Isomer relationship | Average | 0.24 | 0.14 | Near random |
| Laboratory safety | Average | 0.46 | varies | Below practical utility |
| AFM interpretation | Average | 0.24 | varies | Near random |
| Henry constant comparison | Average | 0.83 | varies | Strongest interpretation task |
Hardware
The paper does not specify hardware requirements. All evaluations were run through commercial API endpoints.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| MaCBench Repository | Code | MIT | Benchmark data and evaluation card |
| ChemBench Framework | Code | MIT | Evaluation pipeline (v0.3.0) |
| MaCBench Dataset | Dataset | CC-BY-4.0 | 1,153 questions with images |
| MaCBench-Ablations | Dataset | CC-BY-4.0 | Ablation task variants |
| ChemBench v0.3.0 (Zenodo) | Code | MIT | Archived release |
Paper Information
Citation: Alampara, N., Schilling-Wilhelmi, M., Ríos-García, M., Mandal, I., Khetarpal, P., Grover, H. S., Krishnan, N. M. A., & Jablonka, K. M. (2025). Probing the limitations of multimodal language models for chemistry and materials research. Nature Computational Science, 5(10), 952-961. https://doi.org/10.1038/s43588-025-00836-3
@article{alampara2025macbench,
title={Probing the limitations of multimodal language models for chemistry and materials research},
author={Alampara, Nawaf and Schilling-Wilhelmi, Mara and R{\'\i}os-Garc{\'\i}a, Marti{\~n}o and Mandal, Indrajeet and Khetarpal, Pranav and Grover, Hargun Singh and Krishnan, N. M. Anoop and Jablonka, Kevin Maik},
journal={Nature Computational Science},
volume={5},
number={10},
pages={952--961},
year={2025},
publisher={Nature Publishing Group},
doi={10.1038/s43588-025-00836-3}
}