LLM-Generated Text as Molecular Representations
This is a Method paper that proposes using large language models (specifically ChatGPT) to generate natural language explanations for molecules represented as SMILES strings, and then using those explanations as input representations for downstream molecular property prediction. The approach is called Captions as new Representations (CaR). The authors also evaluate ChatGPT directly on zero-shot and few-shot molecular classification to gauge in-context learning ability on chemical data.
Bridging Molecular Data and Natural Language Understanding
Molecular property prediction is central to virtual screening, drug discovery, and materials design. Molecules are typically represented either as graphs (processed by GNNs) or as SMILES strings (processed by NLP-based methods). While both paradigms have shown success, they do not directly use the broad world knowledge embedded in large language models.
LLMs such as ChatGPT demonstrate strong capabilities in text understanding and can generate informative descriptions when given SMILES strings, including functional groups, chemical properties, and potential pharmaceutical applications. The question motivating this work is whether LLM-generated textual descriptions can serve as better molecular representations than raw SMILES or graph encodings for property prediction tasks.
Prior work had not systematically explored two directions: (1) whether LLMs can perform molecular classification via in-context learning, and (2) whether LLM-generated captions can serve as transferable representations for small downstream models.
Captions as Representations (CaR)
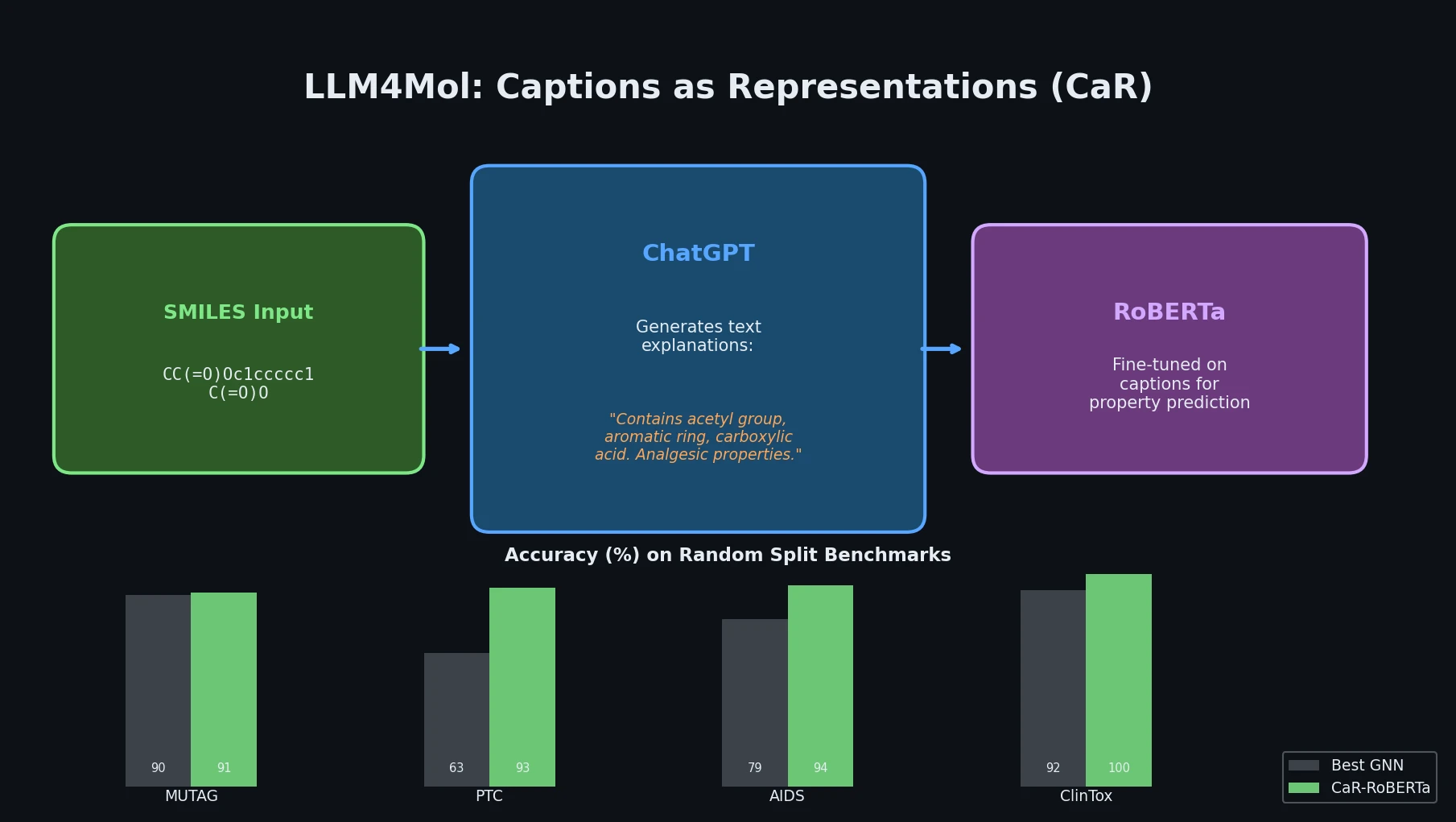
The core contribution is the CaR framework, which operates in two stages:
Caption generation: Given a molecule’s SMILES string, ChatGPT is prompted to produce a detailed textual explanation covering functional groups, chemical properties, and potential applications.
Fine-tuning a small LM: The generated text explanations replace the original SMILES as input to a pre-trained language model (e.g., RoBERTa). This small LM is then fine-tuned on downstream classification or regression tasks.
The insight is that ChatGPT’s world knowledge can enrich the molecular representation with semantically meaningful features that raw SMILES lack. For example, on the PTC (Predictive Toxicology Challenge) dataset, the authors performed keyword searches for terms like “toxicity”, “cancer”, and “harmful” in the ChatGPT-generated explanations and found that these keywords appeared predominantly in entries labeled as toxic, indicating that the generated captions carry predictive signal.
The authors also explore in-context molecular classification, where ChatGPT is directly prompted with zero or few examples to classify molecules. This serves as a preliminary evaluation of LLM reasoning capabilities on molecular data.
Experimental Setup and Benchmarks
Datasets
The evaluation spans 9 datasets across classification and regression:
- Classification (TUDataset): MUTAG, PTC, AIDS
- Classification (MoleculeNet): SIDER, ClinTox, BACE, BBBP
- Regression (MoleculeNet): ESOL, Lipophilicity
Baselines
Baselines include GNN-based methods (GCN, GIN, ChebyNet, D-MPNN, GraphMVP, InfoGraph, G-Motif, Mole-BERT) and SMILES-based methods (ECFP4-MLP, SMILES-Transformer, MolR, ChemBERTa, MolKD).
Splitting Strategies
- Random splitting: 8/1/1 train/validate/test with 10-fold cross-validation
- Scaffold splitting: 5 random seeds, reported as mean and standard deviation
Key Results: Random Splitting
Under random splitting, CaR-RoBERTa achieves the best results on almost all datasets:
| Method | MUTAG (ACC) | PTC (ACC) | AIDS (ACC) | SIDER (AUC) | ClinTox (AUC) | ESOL (RMSE) | Lipo (RMSE) |
|---|---|---|---|---|---|---|---|
| GCN | 90.00 | 62.57 | 78.68 | 64.24 | 91.88 | 0.77 | 0.80 |
| GIN | 89.47 | 58.29 | 78.01 | 66.19 | 92.08 | 0.67 | 0.79 |
| ECFP4-MLP | 96.84 | 85.71 | 94.64 | 90.19 | 95.81 | 0.60 | 0.60 |
| CaR-RoBERTa | 91.05 | 93.14 | 94.37 | 88.81 | 99.80 | 0.45 | 0.47 |
CaR-RoBERTa improves over the best GNN by up to 53% on PTC and reduces RMSE by 35-37% on regression tasks. However, ECFP4-MLP outperforms CaR on MUTAG (96.84 vs. 91.05).
Key Results: Scaffold Splitting
Under the more challenging scaffold splitting:
| Method | SIDER (AUC) | ClinTox (AUC) | BACE (AUC) | BBBP (AUC) | ESOL (RMSE) | Lipo (RMSE) |
|---|---|---|---|---|---|---|
| GraphMVP-C | 63.90 | 77.50 | 81.20 | 72.40 | 1.03 | 0.68 |
| Mole-BERT | 62.80 | 78.90 | 80.80 | 71.90 | 1.02 | 0.68 |
| MolKD | 61.30 | 83.80 | 80.10 | 74.80 | - | - |
| CaR-RoBERTa | 58.06 | 84.16 | 80.73 | 81.99 | 0.96 | 1.02 |
Results are more mixed under scaffold splitting. CaR achieves the best performance on ClinTox (+30% over GNNs) and BBBP (+15%), but underperforms on SIDER and Lipophilicity.
Few-Shot Classification with ChatGPT
Direct few-shot classification with ChatGPT shows mixed results. On MUTAG, ChatGPT underperforms classical methods across all shot counts. On PTC, ChatGPT outperforms GNNs in the few-shot regime. Performance improves with increasing number of shots, but results are inconsistent across different prompts.
Replacing the Small LM
The authors test CaR with different downstream models: RoBERTa, DeBERTa, and an adaptive language model for molecules. Pre-trained models all perform similarly, and all outperform a DeBERTa trained from scratch, validating that CaR’s effectiveness comes from the caption quality rather than the specific choice of downstream model.
Findings, Limitations, and Future Directions
Key Findings
- ChatGPT-generated text explanations serve as effective molecular representations, outperforming GNNs and SMILES-based methods on most benchmarks under random splitting.
- ChatGPT has some capacity for few-shot molecular classification, but performance is inconsistent and prompt-sensitive.
- The CaR approach is model-agnostic: different pre-trained small LMs achieve similar results when fine-tuned on the generated captions.
- Under scaffold splitting, CaR shows strong results on some datasets (ClinTox, BBBP) but underperforms on others (SIDER, Lipophilicity).
Limitations Acknowledged by the Authors
- Single LLM: Only ChatGPT was used. Other LLMs (GPT-4, domain-specific models like MolReGPT) were not evaluated.
- No graph structure integration: CaR treats molecular prediction purely as an NLP task and does not incorporate structural graph information, which is known to be important for molecular properties.
- Limited to small molecules: The approach works only for molecules representable as SMILES. Proteins, antibodies, and other large biomolecules with 3D structure are not addressed.
Additional Considerations
The random splitting results are notably strong, but random splits tend to overestimate performance compared to scaffold splits, which test generalization to structurally novel molecules. The high variance on some scaffold-split results (e.g., ClinTox with 17.63 standard deviation) suggests instability. The reliance on a proprietary API (ChatGPT) also limits reproducibility and introduces cost constraints for large-scale applications.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Classification | MUTAG (TUDataset) | 188 molecules | Mutagenicity prediction |
| Classification | PTC (TUDataset) | 344 molecules | Predictive toxicology |
| Classification | AIDS (TUDataset) | 2,000 molecules | HIV activity |
| Classification | SIDER (MoleculeNet) | 1,427 molecules | Side effect prediction |
| Classification | ClinTox (MoleculeNet) | 1,478 molecules | Clinical trial toxicity |
| Classification | BACE (MoleculeNet) | 1,513 molecules | Beta-secretase inhibition |
| Classification | BBBP (MoleculeNet) | 2,039 molecules | Blood-brain barrier penetration |
| Regression | ESOL (MoleculeNet) | 1,128 molecules | Aqueous solubility |
| Regression | Lipophilicity (MoleculeNet) | 4,200 molecules | Lipophilicity |
Algorithms
- ChatGPT (GPT-3.5) generates textual explanations for SMILES strings
- RoBERTa is fine-tuned on generated captions using HuggingFace Transformers with default parameters
- 10-fold cross-validation for random split; 5 random seeds for scaffold split
Models
- ChatGPT (GPT-3.5) for caption generation
- RoBERTa-base for downstream fine-tuning (default HuggingFace parameters)
- DeBERTa and adaptive-lm-molecules tested as alternatives
Evaluation
- Classification: accuracy (ACC) and ROC-AUC
- Regression: RMSE
- Mean and standard deviation reported across folds/seeds
Hardware
Not specified in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| LLM4Mol | Code | Not specified | Official implementation |
Paper Information
Citation: Qian, C., Tang, H., Yang, Z., Liang, H., & Liu, Y. (2023). Can Large Language Models Empower Molecular Property Prediction? arXiv preprint arXiv:2307.07443. https://arxiv.org/abs/2307.07443
@article{qian2023can,
title={Can Large Language Models Empower Molecular Property Prediction?},
author={Qian, Chen and Tang, Huayi and Yang, Zhirui and Liang, Hong and Liu, Yong},
journal={arXiv preprint arXiv:2307.07443},
year={2023},
doi={10.48550/arxiv.2307.07443}
}