Paper Information
Citation: White, A. D., Hocky, G. M., Gandhi, H. A., Ansari, M., Cox, S., Wellawatte, G. P., Sasmal, S., Yang, Z., Liu, K., Singh, Y., & Peña Ccoa, W. J. (2023). Assessment of chemistry knowledge in large language models that generate code. Digital Discovery, 2(2), 368-376. https://doi.org/10.1039/d2dd00087c
Publication: Digital Discovery 2023
Additional Resources:
- nlcc-data benchmark repository
- Evaluation completions website
- Zenodo evaluation data (DOI: 10.5281/zenodo.6800475)
Benchmarking Chemistry Knowledge in Code-Generating LLMs
This is an Empirical paper that evaluates code-generating large language models on chemistry tasks. The primary contribution is a categorized benchmark of 84 chemistry problems across 10 topics, along with a systematic evaluation of several LLMs (Codex cushman, Codex davinci, text-davinci-003, InCoder, CodeGen) on these tasks. The paper also provides practical guidance on prompt engineering strategies that improve accuracy.
Why Evaluate LLMs on Chemistry Coding Tasks
As of late 2022, LLMs trained on code (such as Codex and InCoder) had become widely available through tools like GitHub Copilot and Tabnine. An open question was whether these general-purpose code models contained sufficient domain knowledge to solve chemistry problems expressed as coding tasks. Chemistry has specialized language, equations, and conventions (e.g., SMILES notation, thermodynamic relationships, molecular simulation methods) that may not be well-represented in general code training data. Prior work had shown that knowledge of the periodic table requires very high parameter counts, but the broader extent of chemistry knowledge in code LLMs was unexplored.
The authors sought to answer a specific question: do code-generating LLMs “know” chemistry? This means evaluating whether LLMs can correlate natural language descriptions of chemistry problems with correct code implementations, including proper equations, units, and use of domain-specific libraries.
Benchmark Design and Prompt Engineering Strategies
The benchmark covers 10 topic categories:
| Topic | Abbreviation | N | Expert-only |
|---|---|---|---|
| Biochemistry | bio | 13 | 2 |
| Cheminformatics | cheminf | 10 | 0 |
| General chemistry | genchem | 11 | 0 |
| Molecular dynamics | md | 11 | 3 |
| Plotting | plot | 10 | 10 |
| Quantum mechanics | qm | 8 | 3 |
| Simulation methods | sim | 8 | 5 |
| Spectroscopy | spect | 11 | 1 |
| Statistics | stats | 11 | 1 |
| Thermodynamics | thermo | 10 | 0 |
Each task is formatted as a Python function with a docstring describing the expected behavior. The LLM must generate a completion that passes automated unit tests. Of the 84 total prompts, 25 require expert evaluation (e.g., plotting tasks) where automated testing is insufficient.
The key prompt engineering insight is the use of “contexts,” which are code prepended before prompts. The authors tested several context strategies:
- Custom context: Topic-specific imports (e.g., RDKit for cheminformatics) plus a one-line completion example to teach the model how to signal the end of output.
- Insert context: Uses model infilling capabilities instead of completion-based generation. Available for davinci and InCoder.
- Copyright context: Adding a copyright notice at the top of the file, which conditions the model toward higher-quality code patterns.
- Authority context: Adding “This is written by an expert Python programmer.”
The copyright notice improved accuracy at higher temperatures. The intuition is that copyrighted code in training data tends to be higher-quality, so the notice acts similarly to lowering temperature. The best model/temperature combination (davinci at T=0.05) was already operating at effectively low temperature, so the copyright trick did not further improve it.
Experimental Setup: Models, Sampling, and Expert Evaluation
Models evaluated
The study compared five models, all decoder-only architectures:
| Model | Abbreviation | Parameters | Source |
|---|---|---|---|
| code-cushman-001 | cushman | 12B | OpenAI (GPT-3 fine-tuned on code) |
| code-davinci-002 | davinci | ~175B (estimated) | OpenAI (GPT-3.5 class) |
| text-davinci-003 | davinci3 | ~175B (estimated) | OpenAI (RLHF-adapted from davinci) |
| InCoder | incoder | 6B | Fried et al. 2022 |
| CodeGen | codegen | 16B | Nijkamp et al. 2022 |
Sampling and evaluation
Completions were generated using top-k sampling (k=5) at three temperatures: T=0.05, 0.2, and 0.5. For InCoder-6B, GPU memory limited sampling to k=1. Error bars in all reported results are 95% confidence intervals from bootstrap resampling across top-k samples.
Accuracy was defined following the HumanEval approach: a completion is correct if the code runs and passes unit tests, regardless of whether it matches a reference implementation.
Expert evaluation
Nine co-authors (postdoctoral scholars and Ph.D. students) performed 650 evaluations of davinci completions through a web interface. Each completion was scored on a 5-point scale: Perfect (5), Correct but not perfect (4), Runs and is almost correct (3), Does not run but is almost correct (2), Far from correct (1). Expert-evaluated accuracy counted only “Perfect” and “Correct but not perfect” as correct.
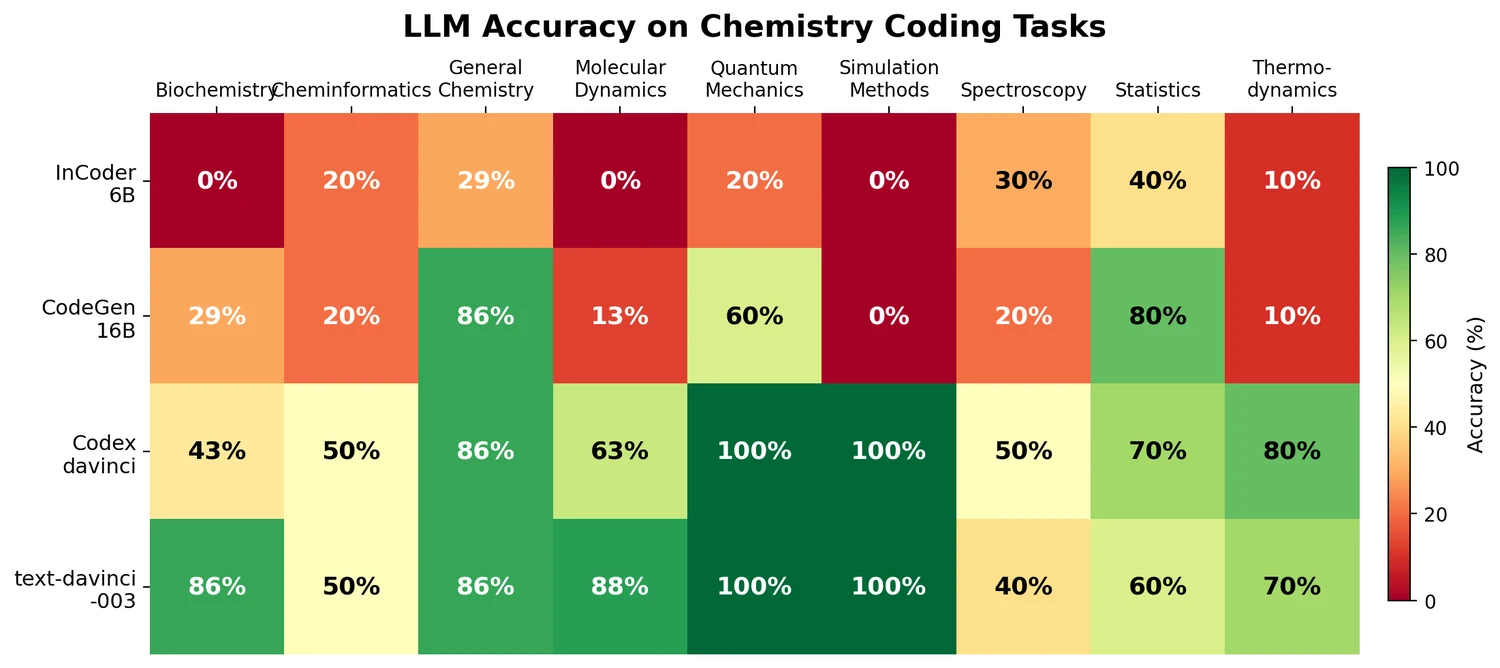
Key results by topic and model
| Topic | incoder | codegen | davinci | davinci3 |
|---|---|---|---|---|
| bio | 0% | 29% | 43% | 86% |
| cheminf | 20% | 20% | 50% | 50% |
| genchem | 29% | 86% | 86% | 86% |
| md | 0% | 13% | 63% | 88% |
| qm | 20% | 60% | 100% | 100% |
| sim | 0% | 0% | 100% | 100% |
| spect | 30% | 20% | 50% | 40% |
| stats | 40% | 80% | 70% | 60% |
| thermo | 10% | 10% | 80% | 70% |
| total | 17% | 35% | 72% | 75% |
All accuracies reported use the best context for each model (copyright for incoder-6B, authority for codegen-16B, insert for davinci) at T=0.2.
Findings: LLMs Know Chemistry, With Caveats
The central finding is that code-generating LLMs do contain substantial chemistry knowledge. The best model (davinci) achieved 72% overall accuracy, with prompt engineering contributing approximately 30 percentage points to this figure. The text-davinci-003 model, which was fine-tuned with RLHF, achieved 75% and showed reduced sensitivity to prompt engineering, suggesting that human feedback alignment partially subsumes the benefits of manual prompt design.
Strengths and successful domains
- Quantum mechanics and simulation: davinci achieved 100% on both categories, indicating strong knowledge of computational chemistry equations and simulation patterns.
- General chemistry: All models except InCoder performed well (86%), suggesting that general chemistry concepts are well-represented in code training data.
- Molecular structure generation: InstructGPT showed some ability to connect natural language descriptions with SMILES strings, generating valid (though not exact) molecular structures from prompts like “a phenol derivative.”
Limitations and failure modes
- Lack of reasoning: The authors emphasize that LLMs demonstrate knowledge correlation, not reasoning. Davinci frequently uses “relativistic Hartree-Fock” for any prompt requesting a “highly accurate” quantum calculation, because it has memorized the association between “relativistic” and “accurate” rather than understanding the underlying chemistry.
- Hallucinated functions: When given difficult prompts (e.g., “return the residual dipolar couplings given a SMILES string”), the model invents non-existent functions like
MolToRDC. - API version mismatches: Many errors in the molecular dynamics category stem from the model using outdated function signatures for packages like MDTraj, likely reflecting the training data cutoff.
- Expert-evaluated accuracy is lower: On topics requiring expert evaluation (generally harder tasks), accuracy drops, and it correlates negatively with perceived difficulty.
Practical recommendations
The paper offers several practical tips for using code LLMs in chemistry:
- Use correctly spelled, precise prompts. If a function should “return” a value, use the word “return” rather than “compute.”
- Be explicit about what variables represent (e.g., specify that k is a spring constant, not Boltzmann’s constant).
- Import only the packages you intend to use, as the model will attempt to use all imported libraries.
- Adding a copyright notice or “expert programmer” statement can improve accuracy, though RLHF-trained models are less sensitive to this.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Evaluation | nlcc-data benchmark | 84 prompts across 10 chemistry topics | Open source, community-extensible |
| Expert evaluation | Human evaluations CSV | 650 evaluations | Available in Supporting Information |
Algorithms
Evaluation uses automated unit testing for 59 of 84 prompts. Expert evaluation covers the remaining 25 prompts through a web-based scoring interface. Five completions per prompt were generated via top-k sampling at three temperatures.
Models
All models evaluated are external (OpenAI API for Codex/davinci, HuggingFace for InCoder/CodeGen). No new models were trained. Python version and packages were pinned to June 2021 to avoid library changes influencing results.
Evaluation
Accuracy is binary: a completion passes all unit tests (1.0) or fails (0.0), averaged across top-k samples and temperatures. Expert evaluation uses a 5-point scale collapsed to binary (Perfect or Correct = 1.0).
Hardware
GPU memory limitations are mentioned for InCoder-6B (limiting k=1 instead of k=5). No other hardware details are specified.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| nlcc-data benchmark | Dataset | Unknown | Open-source benchmark prompts and solutions |
| Evaluation website | Other | Unknown | Web interface showing completions |
| Zenodo evaluation data | Dataset | Unknown | Expert evaluation completions in HTML |
| Paper (open access) | Other | CC-BY-NC | Published article |
Citation
@article{white2023assessment,
title={Assessment of chemistry knowledge in large language models that generate code},
author={White, Andrew D. and Hocky, Glen M. and Gandhi, Heta A. and Ansari, Mehrad and Cox, Sam and Wellawatte, Geemi P. and Sasmal, Subarna and Yang, Ziyue and Liu, Kangxin and Singh, Yuvraj and Peña Ccoa, Willmor J.},
journal={Digital Discovery},
volume={2},
number={2},
pages={368--376},
year={2023},
publisher={Royal Society of Chemistry},
doi={10.1039/d2dd00087c}
}