A Scientific Language Model Trained on Curated Knowledge
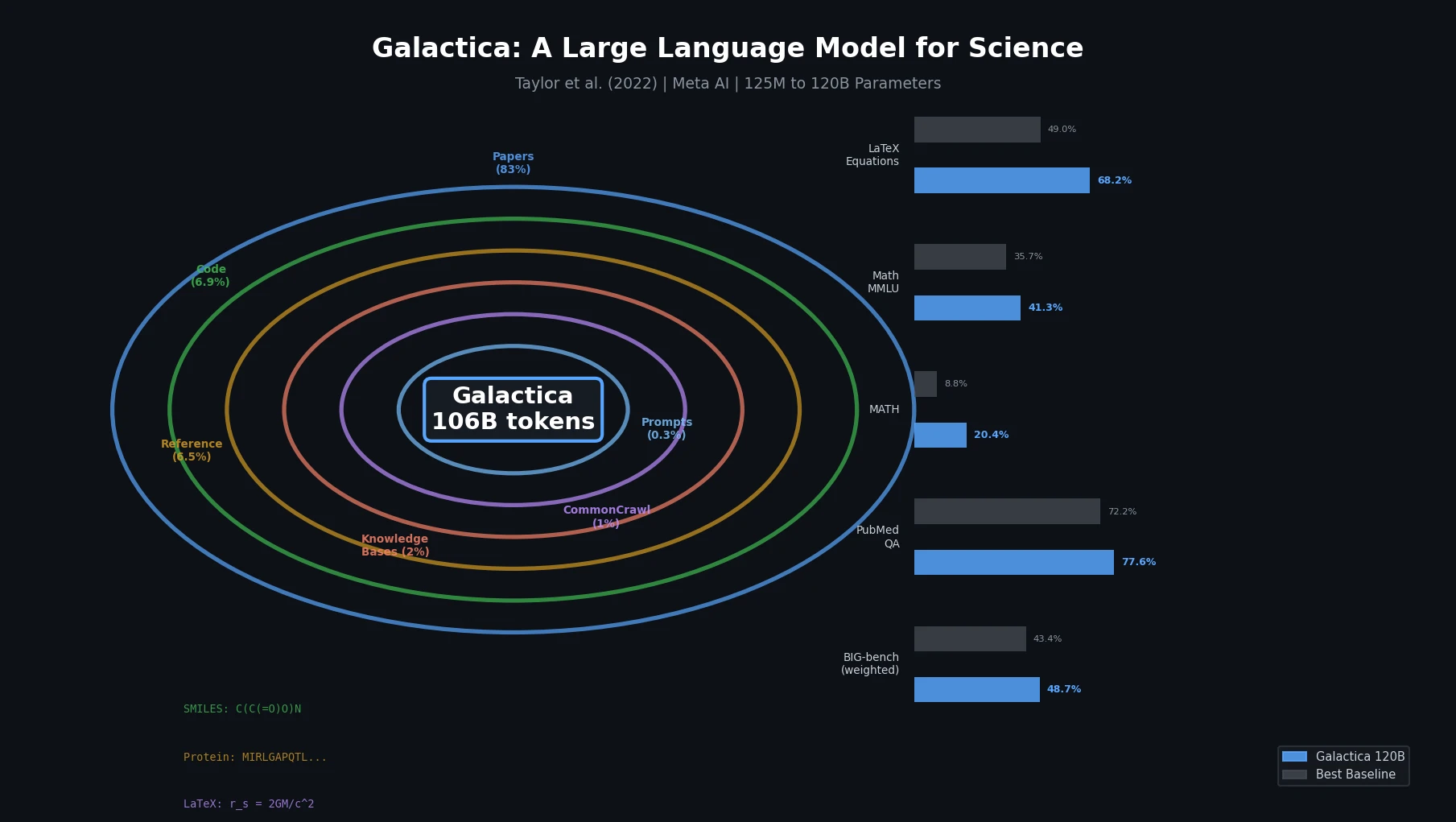
Galactica is a Resource contribution: a family of decoder-only Transformer language models (125M to 120B parameters) trained on a curated corpus of 106 billion tokens from scientific papers, reference material, knowledge bases, and other sources. The paper also introduces several specialized tokenization schemes for scientific modalities (SMILES, amino acid sequences, DNA sequences, LaTeX, citations) and a working memory token (<work>) for step-by-step reasoning. All model weights are open-sourced under the Apache 2.0 license.
Information Overload as the Motivating Problem
The volume of scientific literature has grown beyond any individual’s capacity to process. An average of 516 papers per day were submitted to arXiv as of May 2022, and databases like NCBI GenBank contained $1.49 \times 10^{12}$ nucleotide bases as of August 2022. Current search engines point to secondary knowledge layers (Wikipedia, UniProt, PubChem) that require costly human curation, creating a throughput bottleneck.
The authors argue that large language models can serve as a new interface for science by storing, combining, and reasoning about scientific knowledge in weight memory, rather than relying on the traditional store-and-retrieve paradigm. Prior scientific language models (SciBERT, BioLM) were small in scale, and general LLMs (GPT-3, PaLM) trained on uncurated web data that is inefficient for scientific tasks.
Curated Corpus and Specialized Tokenization
The core innovation has two components: a normative approach to dataset curation and a set of specialized tokens for different scientific modalities.
The Galactica Corpus
The training corpus consists of 106 billion tokens with a deliberate focus on quality over quantity:
| Data Source | Documents | Tokens | Token % |
|---|---|---|---|
| Papers | 48 million | 88 billion | 83.0% |
| Code | 2 million | 7 billion | 6.9% |
| Reference Material | 8 million | 7 billion | 6.5% |
| Knowledge Bases | 2 million | 2 billion | 2.0% |
| Filtered CommonCrawl | 0.9 million | 1 billion | 1.0% |
| Prompts | 1.3 million | 0.4 billion | 0.3% |
| Other | 0.02 million | 0.2 billion | 0.2% |
Papers come from arXiv (35B tokens), PMC (23B), Semantic Scholar (18B), and PubMed abstracts (5B), among others. Reference material includes Wikipedia (5B tokens), StackExchange (1B), textbooks, and lecture notes. Knowledge bases include PubChem Compound (2M compounds, 1B tokens), UniProt (552K reviewed Swiss-Prot proteins, 0.6B tokens), and the RefSeq Genome.
All data is processed into a common markdown format. Mathematical LaTeX is preserved where available, and papers are citation-processed with title-based identifiers.
Specialized Tokenization
Galactica introduces several modality-specific tokenization strategies:
Citations: Wrapped with
[START_REF]and[END_REF]tokens using paper titles as identifiers, enabling the model to predict citations in context.Working Memory (
<work>): Step-by-step reasoning is wrapped in<work>and</work>tokens that mimic an internal working memory, allowing the model to perform multi-step computation. This differs from chain-of-thought prompting in that it is learned during pre-training rather than elicited through prompt engineering.SMILES: Wrapped with
[START_SMILES]/[END_SMILES]tokens and character-level tokenization.Amino Acid Sequences: Wrapped with
[START_AMINO]/[END_AMINO]tokens with character-level tokenization (one token per residue).DNA Sequences: Wrapped with
[START_DNA]/[END_DNA]tokens with character-level tokenization (one token per nucleotide base).Mathematics: ASCII operations split into individual characters; digits split into individual tokens.
Prompt Pre-Training
Rather than using instruction tuning as a separate fine-tuning stage, Galactica includes task-specific prompts (358 million tokens total) directly in pre-training alongside the general corpus. This includes question answering, entity extraction, summarization, dialog, and chemical property prediction prompts. The authors frame this as occupying a middle ground between pure self-supervised pre-training and instruction tuning, providing task signal without degrading general capability.
Architecture, Training, and Evaluation Setup
Architecture
Galactica uses a standard decoder-only Transformer with several modifications:
- GeLU activations
- 2048-token context window
- No biases in dense kernels or layer norms
- Learned positional embeddings
- 50K BPE vocabulary
Five model sizes were trained:
| Model | Parameters | Layers | $d_{\text{model}}$ | Heads | Batch Size | Max LR |
|---|---|---|---|---|---|---|
| GAL 125M | 125M | 12 | 768 | 12 | 0.5M | $6 \times 10^{-4}$ |
| GAL 1.3B | 1.3B | 24 | 2,048 | 32 | 1.0M | $2 \times 10^{-4}$ |
| GAL 6.7B | 6.7B | 32 | 4,096 | 32 | 2.0M | $1.2 \times 10^{-4}$ |
| GAL 30B | 30.0B | 48 | 7,168 | 56 | 2.0M | $1 \times 10^{-4}$ |
| GAL 120B | 120.0B | 96 | 10,240 | 80 | 2.0M | $0.7 \times 10^{-5}$ |
Training used AdamW with $\beta_1 = 0.9$, $\beta_2 = 0.95$, weight decay of 0.1, gradient clipping at 1.0, and linear learning rate decay to 10% of peak value. Dropout and attention dropout were set to $p = 0.1$.
Training on Repeated Tokens
Models were trained for 450 billion tokens, approximately 4.25 epochs of the corpus. Validation loss continued to fall through four epochs for all model sizes, with the 120B model only beginning to overfit at the start of the fifth epoch. This is notable because it challenges the prevailing view that repeated tokens are harmful for LLM training. Performance on out-of-domain BIG-bench tasks also continued to improve through training, suggesting no overfitting on downstream generalization.
Key Evaluation Results
Knowledge Probes: On LaTeX equation prediction across 434 equations from chemistry, physics, mathematics, statistics, and economics, GAL 120B achieved 68.2% accuracy versus GPT-3’s 49.0% (zero-shot). On chemical reactions, GAL 120B scored 43.1% versus GPT-3’s 35.1%.
Mathematical Reasoning: With the <work> token, GAL 120B achieved 41.3% on mathematical MMLU (average across abstract algebra, elementary, high school, college math, and formal logic), compared to Chinchilla’s 35.7% (5-shot). On the MATH benchmark, GAL 120B scored 20.4% (5-shot chain-of-thought) versus PaLM 540B’s 8.8%.
Scientific QA: Galactica set state-of-the-art results on PubMedQA (77.6%) and MedMCQA dev (52.9%), outperforming prior fine-tuned models (72.2% and 41.0% respectively).
Citation Prediction: GAL 120B achieved 51.9% accuracy on PWC Citations and 69.1% on Extended Citations, outperforming both sparse (ElasticSearch) and dense (Contriever) retrieval baselines.
BIG-bench (57 tasks): Despite training only on scientific data, GAL 120B (48.7% weighted accuracy) outperformed OPT 175B (43.4%) and BLOOM 176B (42.6%) on primarily non-scientific tasks.
MoleculeNet Classification: Using SMILES in natural language prompts with weak supervision, GAL 120B achieved an average ROC-AUC of 0.690 across six MoleculeNet classification benchmarks (BACE, BBBP, ClinTox, HIV, SIDER, Tox21). This lagged the specialist Uni-Mol model (0.770), which uses 3D molecular information and 10x more molecules.
IUPAC Name Prediction: GAL 120B achieved 39.2% accuracy on predicting IUPAC names from SMILES in a self-supervised setting, with attention visualization showing the model attends to chemically relevant functional groups (e.g., attending to the $\text{-NH}_2$ group when predicting “amino”).
Protein Function Prediction: GAL 120B achieved a ROUGE-L of 0.252 on generating free-form protein function descriptions from amino acid sequences, and an $F_1$ of 48.7% on protein keyword prediction from the UniProt general validation set.
Bias and Toxicity: On CrowS-Pairs, GAL 120B scored 60.5% (closer to ideal 50%) versus OPT 175B’s 69.5%. On StereoSet, GAL 120B achieved an ICAT score of 65.6 versus OPT’s 60.0 and GPT-3’s 60.8. Toxicity rates on RealToxicityPrompts were substantially lower than comparison models.
Findings, Limitations, and Future Directions
Key Findings
Curated data enables repeated training: The curated scientific corpus allows training for multiple epochs without overfitting, contrary to prevailing assumptions about repeated token degradation.
Scientific LLMs generalize beyond science: Despite training only on scientific text, Galactica outperforms general LLMs on non-scientific BIG-bench tasks, suggesting data quality matters more than data breadth.
Weight memory can outperform retrieval: For citation prediction, Galactica’s weight memory outperforms traditional sparse and dense retrieval methods, demonstrating the context-associative power of language models.
Multi-modal learning via text: SMILES and protein sequences can be learned alongside natural language in a single model, and the model attends to chemically interpretable features.
Limitations
The authors acknowledge several limitations:
- Corpus constraints: Restricted to open-access papers; much scientific knowledge in closed-access papers and textbooks is excluded. Only 2M of 110M PubChem compounds and 0.5M of 227M UniProt sequences were included.
- Corpus vs. prompt effects: The paper does not disentangle whether performance gains come from the scientific corpus or from the prompt pre-training strategy.
- Citation bias: The model still shows bias toward predicting more popular papers, though this decreases with scale.
- No geometry: SMILES-based representations lack 3D geometric information, limiting chemical understanding.
- Hallucination: Title-based citation identifiers are more prone to hallucination at smaller scales, though accuracy improves with scale.
- No instruction tuning comparison: The paper does not compare prompt pre-training against instruction tuning as a follow-up step.
Future Directions
The paper identifies retrieval augmentation, extending to images, larger context windows, mixture-of-denoising training objectives, and more diverse <work> reasoning examples as promising directions.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training | Galactica Corpus | 106B tokens | Papers (83%), code (6.9%), reference material (6.5%), knowledge bases (2%), CommonCrawl (1%), prompts (0.3%) |
| Training (Molecules) | PubChem Compound subset | 2M compounds (of 110M available) | Character-level SMILES tokenization |
| Training (Proteins) | Swiss-Prot (UniProt) | 552K reviewed sequences (of 227M available) | Character-level amino acid tokenization |
| Evaluation | LaTeX Equations | 434 equations | Chemistry, physics, math, stats, economics |
| Evaluation | MMLU, MATH | Standard benchmarks | Out-of-domain evaluation |
| Evaluation | PubMedQA, MedMCQA, BioASQ | Standard biomedical QA | In-domain (training prompts included) |
| Evaluation | MoleculeNet (6 tasks) | Standard molecular benchmarks | BACE, BBBP, ClinTox, HIV, SIDER, Tox21 |
| Evaluation | BIG-bench (57 tasks) | Standard NLP benchmark | Out-of-domain, non-scientific |
Algorithms
- Decoder-only Transformer with GeLU activations, no biases
- AdamW optimizer: $\beta_1 = 0.9$, $\beta_2 = 0.95$, weight decay 0.1
- Gradient clipping at global norm 1.0
- Linear LR decay to 10% of peak
- Dropout: $p = 0.1$ (attention and residual)
- BPE vocabulary: 50K tokens from 2% corpus sample
- Training: 450B tokens (~4.25 epochs)
Models
| Artifact | Type | License | Notes |
|---|---|---|---|
| Galactica models (galai) | Code + Model | Apache-2.0 | Official implementation with 125M, 1.3B, 6.7B, 30B, 120B checkpoints |
Evaluation
| Metric | GAL 120B | Best Baseline | Notes |
|---|---|---|---|
| LaTeX Equations (zero-shot) | 68.2% | GPT-3: 49.0% | 434 equations across 5 domains |
Math MMLU (<work>) | 41.3% | Chinchilla (5-shot): 35.7% | Average over 5 math subjects |
| MATH (5-shot CoT) | 20.4% | PaLM 540B: 8.8% | Minerva 540B (fine-tuned): 33.6% |
| PubMedQA | 77.6% | Prior SOTA: 72.2% | In-domain |
| MedMCQA dev | 52.9% | Prior SOTA: 41.0% | In-domain |
| BIG-bench (weighted) | 48.7% | OPT 175B: 43.4% | 57 non-scientific tasks |
| MoleculeNet ROC-AUC (avg) | 0.690 | Uni-Mol (3D): 0.770 | Weak supervision vs. direct fine-tuning |
| CrowS-Pairs (lower = less biased) | 60.5% | OPT 175B: 69.5% | Ideal: 50% |
Hardware
- 120B model training: 128 NVIDIA A100 80GB nodes
- 120B model inference: single NVIDIA A100 node
- Training library: metaseq (Meta AI)
Paper Information
Citation: Taylor, R., Kardas, M., Cucurull, G., Scialom, T., Hartshorn, A., Saravia, E., Poulton, A., Kerkez, V., & Stojnic, R. (2022). Galactica: A Large Language Model for Science. arXiv preprint arXiv:2211.09085.
@article{taylor2022galactica,
title={Galactica: A Large Language Model for Science},
author={Taylor, Ross and Kardas, Marcin and Cucurull, Guillem and Scialom, Thomas and Hartshorn, Anthony and Saravia, Elvis and Poulton, Andrew and Kerkez, Viktor and Stojnic, Robert},
journal={arXiv preprint arXiv:2211.09085},
year={2022},
doi={10.48550/arxiv.2211.09085}
}