GPT-3 as a Molecular Property Classifier
This is an Empirical paper that evaluates the effectiveness of fine-tuning OpenAI’s GPT-3 language model (specifically the “ada” base model) for predicting electronic and functional properties of organic molecules. Rather than proposing a new architecture, the work systematically tests whether a general-purpose LLM can learn chemically meaningful patterns from SMILES strings when fine-tuned on classification tasks. The primary contribution is the empirical characterization of GPT-3’s performance, robustness, and limitations for molecular property prediction, including extensive ablation studies.
Why Fine-Tune a General-Purpose LLM for Chemistry?
Machine learning for molecular property prediction typically relies on specialized representations: molecular graphs processed by graph neural networks (GNNs), engineered molecular descriptors, or domain-specific chemical language models trained from scratch on SMILES or SELFIES. These approaches require varying levels of domain expertise to design the inputs and architecture.
GPT-3, pre-trained on vast amounts of general text, already has an internal representation of language structure. SMILES notation, as a text-based molecular representation, can be treated as a “language” with its own syntax. The authors hypothesize that GPT-3’s language understanding capabilities, combined with the human-readable nature of SMILES, may enable the model to recognize significant patterns within chemical structures and capture structure-property dependencies. The key question is whether fine-tuning alone is sufficient, or whether specialized architectures provide fundamental advantages.
Prior work by Jablonka et al. showed that fine-tuned GPT-3 could perform surprisingly well on low-data chemistry tasks, sometimes surpassing dedicated models. This paper extends that investigation with a focus on electronic properties (HOMO and LUMO energies) of organic semiconductors, with deeper analysis of robustness and failure modes.
SMILES-to-Classification via Prompt-Completion Fine-Tuning
The core approach is straightforward. Each training example is a prompt-completion pair in JSONL format:
{"prompt": "SMILES_string", "completion": "class_label"}
The SMILES string serves as the prompt, and the fine-tuned model learns to complete it with a class label (0/1 for binary, 0/1/2 for ternary, 0/1/2/3 for quaternary classification). Class thresholds are determined by equally segmenting the property value range. The authors use GPT-3’s default tokenizer, which breaks SMILES strings into subword tokens that do not correspond to chemically meaningful units (e.g., “c1ccccc1” for benzene gets tokenized into arbitrary fragments).
This design choice has important implications. The model must learn chemical semantics from token patterns that are not aligned with atoms or bonds. The authors note this as a limitation and hypothesize that a chemistry-aware tokenizer could improve performance.
Experimental Setup and Baseline Comparisons
Datasets
The primary dataset is a collection of 48,182 organic semiconductor (OSC) molecules extracted from the Cambridge Structural Database (CSD). Each molecule has a SMILES representation and quantum-chemically computed electronic properties (HOMO and LUMO energies). A secondary dataset of 572 aromatic molecular photocatalysts (AMPs) with experimentally measured hydrogen evolution rates (HER) provides an additional test case.
Baselines
Three baselines are compared:
- Directed message-passing neural network (D-MPNN) via Chemprop, using default molecular graph representations
- RDKit molecular descriptors + SVM, using the top 20 descriptors selected by SelectKBest
- Prior ML results from the original AMP dataset paper (using engineered domain-specific features)
Main Results
| Dataset | Task | Classes | GPT-3 Accuracy | GNN Accuracy | Descriptors Accuracy |
|---|---|---|---|---|---|
| OSCs (48,182) | HOMO | 3 | 0.92 | 0.94 | 0.87 |
| OSCs (48,182) | HOMO | 4 | 0.68 | 0.75 | 0.47 |
| OSCs (48,182) | HOMO | 5 | 0.60 | 0.68 | 0.40 |
| OSCs (48,182) | LUMO | 3 | 0.94 | 0.94 | 0.91 |
| AMPs (572) | HER | 2 | 0.88 | 0.86 | 0.87 |
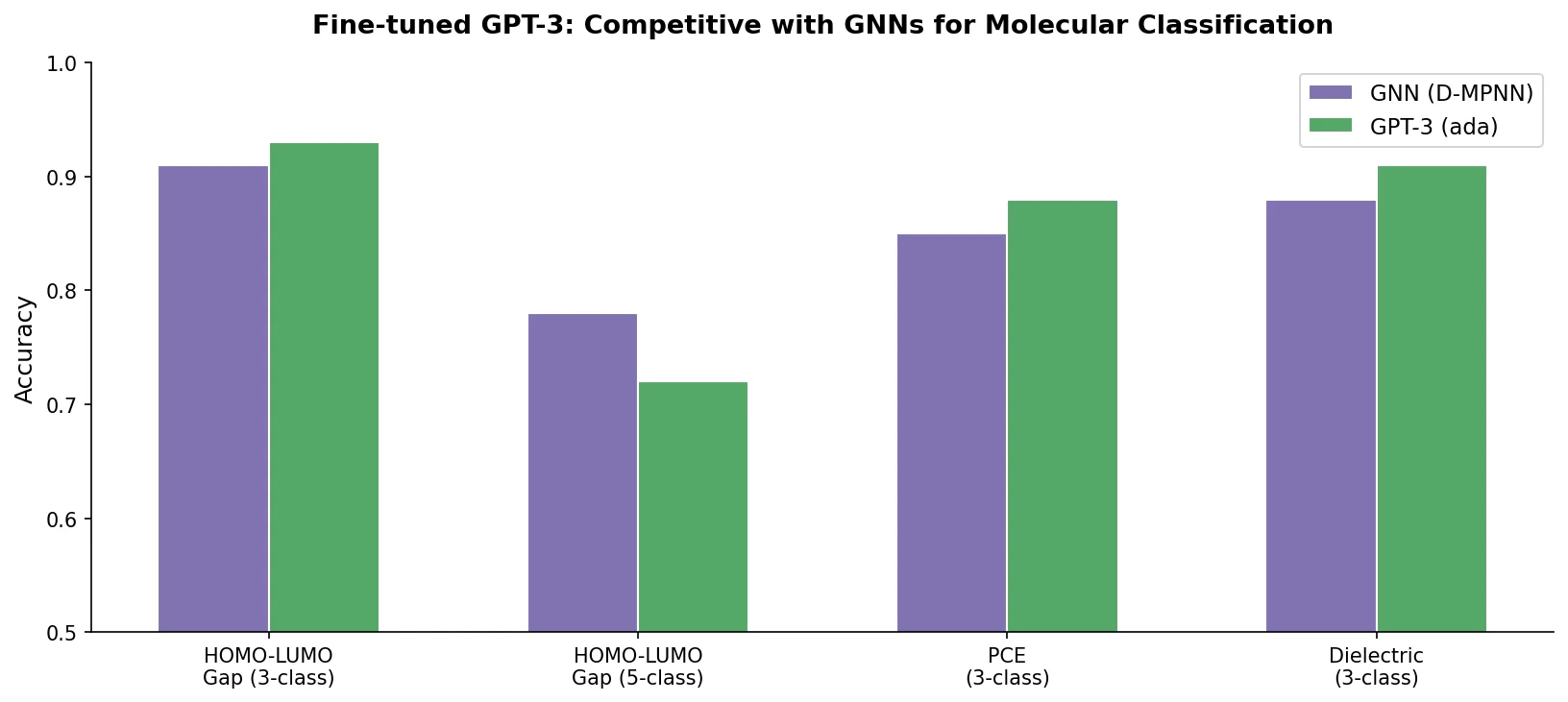
For ternary classification, GPT-3 performs on par with GNNs (0.92 vs. 0.94 for HOMO; 0.94 vs. 0.94 for LUMO). Performance degrades more steeply than GNNs as the number of classes increases: at 5-class HOMO, GPT-3 achieves only 0.60 vs. GNN’s 0.68. On the small AMP dataset (572 molecules), GPT-3 slightly outperforms the GNN (0.88 vs. 0.86).
Learning Curves
The data efficiency analysis reveals that GPT-3 needs at least 20% of the OSC dataset (approximately 9,600 molecules) to reach accuracy above 0.9. Below 1,000 training points, accuracy drops below 0.6. GNNs outperform GPT-3 in this low-data regime, which the authors attribute to (1) the molecular graph being chemically more expressive than SMILES for these tasks, and (2) fine-tuning requiring sufficient data to capture relevant SMILES patterns.
Ablation Study 1: Single-Atom Removal
The authors tested robustness by removing individual non-hydrogen, non-carbon atoms from SMILES strings and replacing them with a <missing> token. Out of 45,763 ablation tests on 7,714 correctly predicted molecules, 95.2% retained the same classification. This suggests the model captures redundant structural information rather than relying on any single atom.
Ablation Study 2: Single-Group Removal
Fifteen chemical groups (nitrile, nitro, enamine, ketone, etc.) were individually ablated. The fine-tuned model attributed the most importance to acetylene (81% agreement for HOMO), enamine (85%), nitro (86%), and ketone (87%) groups, as these altered HOMO predictions in more than 10% of tests. Interestingly, groups that participate in electronic pi-conjugation tended to be more “important” to the model’s HOMO predictions.
When ablated atoms were replaced with random elements instead of the <missing> token, the model failed in 80% of cases for a representative molecule. This suggests the model may “fill in” the missing information when seeing the <missing> token but gets confused by incorrect atomic identities.
Predicting Unknown Molecular Families
The authors held out entire families of polycyclic aromatic hydrocarbons (naphthalene, anthracene, tetracene, pyrene, perylene), quinones, and imides during training, then tested predictions on these unseen families. Results for the first five PAH families:
| Fragment Family | Molecules | GPT-3 HOMO | GNN HOMO | GPT-3 LUMO | GNN LUMO |
|---|---|---|---|---|---|
| Naphthalene | 475 | 0.94 | 0.95 | 0.88 | 0.91 |
| Anthracene | 577 | 0.99 | 1.00 | 0.93 | 0.97 |
| Tetracene | 72 | 0.96 | 1.00 | 0.90 | 0.99 |
| Pyrene | 237 | 0.98 | 1.00 | 0.97 | 0.99 |
| Perylene | 41 | 0.98 | 1.00 | 0.98 | 0.95 |
GPT-3 generalizes well to unknown PAH families, though GNNs have a slight edge on HOMO prediction. Performance degrades somewhat for quinones and imides.
Canonical vs. Non-Canonical SMILES
A model fine-tuned only on canonical SMILES performed poorly on non-canonical variants: only 1,622 of 8,578 molecules achieved consistent predictions across all 11 SMILES variants (1 canonical + 10 non-canonical). Augmenting the training data with 5 non-canonical SMILES per molecule dramatically improved consistency to 7,243 of 8,578 molecules and nearly eliminated erroneous (non-class-label) responses. This finding highlights that GPT-3’s pattern matching is highly sensitive to surface-level string representation and benefits substantially from SMILES enumeration data augmentation.
Key Findings and Limitations
The main findings are:
- Fine-tuned GPT-3 (ada) achieves competitive accuracy with GNNs for coarse-grained (ternary) HOMO/LUMO classification, but performance drops more steeply with finer granularity.
- The model shows robustness to single-atom and single-group ablation, suggesting it captures chemically redundant patterns.
- Generalization to held-out molecular families is strong, though GNNs maintain a slight advantage.
- SMILES augmentation with non-canonical variants is essential for consistent predictions.
The authors acknowledge several limitations:
- Black-box nature: GPT-3 provides no physical insight or interpretability, unlike GNN models where molecular graph features can be augmented with domain knowledge.
- Tokenization: The generic tokenizer does not respect chemical structure. A chemistry-aware tokenizer could improve data efficiency and accuracy.
- SELFIES underperformance: Initial tests with SELFIES did not improve over SMILES, likely because generic tokenization stripped away the extra chemical information SELFIES encodes.
- Cost: Fine-tuning via OpenAI’s API cost approximately $500 for the experiments, and the model is closed-source, preventing systematic interpretation of learned representations.
- Classification only: The approach performs coarse-grained classification rather than regression, limiting utility for applications requiring precise numerical predictions.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training/Evaluation | OSC molecules from CSD | 48,182 | SMILES + DFT-computed HOMO/LUMO energies |
| Training/Evaluation | Aromatic molecular photocatalysts (AMPs) | 572 | Experimental hydrogen evolution rates |
Algorithms
- Fine-tuning uses OpenAI’s GPT-3 “ada” base model via the API
- Prompt-completion pairs in JSONL format
- Default GPT-3 tokenizer
- 80/20 train/test split for OSC; stratified 10-fold CV for AMPs
- Non-canonical SMILES generated using RDKit (10 per molecule for testing, 5 per molecule for augmented training)
Models
- GPT-3 “ada” (fine-tuned, closed-source, accessed via OpenAI API)
- Chemprop D-MPNN baseline (open-source)
- RDKit descriptors + scikit-learn SVM baseline
Evaluation
| Metric | Best GPT-3 Value | Best GNN Value | Task |
|---|---|---|---|
| Accuracy | 0.92 | 0.94 | 3-class HOMO (OSCs) |
| Accuracy | 0.94 | 0.94 | 3-class LUMO (OSCs) |
| Accuracy | 0.88 | 0.86 | 2-class HER (AMPs) |
Hardware
The paper does not specify local hardware requirements. All GPT-3 fine-tuning was conducted via OpenAI’s cloud API at a total cost of approximately $500.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| Chem-GPT-Finetune | Code | Not specified | Python code and datasets for fine-tuning and evaluation |
Paper Information
Citation: Xie, Z., Evangelopoulos, X., Omar, O. H., Troisi, A., Cooper, A. I., & Chen, L. (2024). Fine-tuning GPT-3 for machine learning electronic and functional properties of organic molecules. Chemical Science, 15(2), 500-510.
@article{xie2024finetuning,
title={Fine-tuning {GPT-3} for machine learning electronic and functional properties of organic molecules},
author={Xie, Zikai and Evangelopoulos, Xenophon and Omar, {\"O}mer H. and Troisi, Alessandro and Cooper, Andrew I. and Chen, Linjiang},
journal={Chemical Science},
volume={15},
number={2},
pages={500--510},
year={2024},
publisher={Royal Society of Chemistry},
doi={10.1039/D3SC04610A}
}