A Benchmark Resource for LLM Chemistry Evaluation
This is a Resource paper that introduces ChemLLMBench, a comprehensive benchmark for evaluating large language models on practical chemistry tasks. The primary contribution is the systematic design of eight chemistry tasks organized around three fundamental capabilities (understanding, reasoning, and explaining) along with a standardized evaluation framework that includes prompt templates, in-context learning strategies, and comparison against domain-specific baselines. The benchmark provides the first broad-scope assessment of general-purpose LLMs on chemistry problems, establishing baseline performance levels across multiple models and task types.
Why Benchmark LLMs for Chemistry?
At the time of this work, large language models had demonstrated broad reasoning capabilities across many domains, but their application to practical chemistry tasks remained underexplored. Prior studies (e.g., Nascimento and Pimentel, 2023; Jablonka et al., 2023; White et al., 2023) had examined LLMs on specific chemistry case studies, but no comprehensive or systematic evaluation existed. Two challenges motivated this benchmark:
- Chemistry encompasses diverse task types that require different capabilities. Some tasks can be formulated as problems that LLMs can address (classification, text generation), while others demand deep understanding of molecular representations that LLMs may lack.
- Reliable evaluation requires careful standardization of prompts, demonstration examples, and evaluation procedures. The stochastic nature of LLM outputs and the cost of API calls further constrain experimental design.
The authors, a joint team of AI researchers and chemists at Notre Dame (including the NSF Center for Computer Assisted Synthesis, C-CAS), designed this benchmark to clarify where LLMs are useful for chemistry practitioners and where they fall short.
Eight Tasks Across Three Chemistry Capabilities
The benchmark organizes eight tasks into three capability categories:
Understanding tasks test whether LLMs can interpret molecular representations:
- Name prediction: Translation between SMILES, IUPAC names, and molecular formulas (four subtasks)
- Property prediction: Binary classification on five MoleculeNet datasets (BBBP, HIV, BACE, Tox21, ClinTox)
Reasoning tasks require knowledge of chemical reactions and transformations:
- Yield prediction: Binary classification of high/low yield on Buchwald-Hartwig and Suzuki-Miyaura HTE datasets
- Reaction prediction: Generating product SMILES from reactants/reagents (USPTO-Mixed)
- Reagents selection: Ranking candidate reactants, solvents, or ligands (Suzuki HTE dataset)
- Retrosynthesis: Predicting reactant SMILES from a target product (USPTO-50k)
Explaining tasks leverage LLMs’ natural language capabilities:
- Text-based molecule design: Generating SMILES from a textual molecular description (ChEBI-20)
- Molecule captioning: Generating textual descriptions of molecules from SMILES (ChEBI-20)
Each task uses 100 test instances randomly sampled from established datasets, with evaluations repeated five times to account for LLM output variability.
Evaluation Framework and In-Context Learning Design
Models evaluated
Five LLMs were tested: GPT-4, GPT-3.5 (ChatGPT), Davinci-003, Llama2-13B-chat, and Galactica-30B.
Prompt design
The authors developed a standardized zero-shot prompt template instructing the LLM to act as “an expert chemist” with task-specific input/output descriptions. For in-context learning (ICL), they designed a four-part template: {General Template}{Task-Specific Template}{ICL}{Question}. The task-specific template includes input explanations, output explanations, and output restrictions to reduce hallucinations.
ICL strategies
Two retrieval strategies were explored for selecting demonstration examples:
- Random: Randomly selecting k examples from the candidate pool
- Scaffold: Finding the top-k most similar examples using Tanimoto similarity on Morgan fingerprints (for SMILES inputs) or sequence matching (for text inputs)
The number of examples k was varied per task (typically k in {4, 5, 8, 10, 20}). A validation set of 30 instances was used to select the best five configurations, which were then applied to the test set.
Results summary
The authors classify LLM performance into three categories:
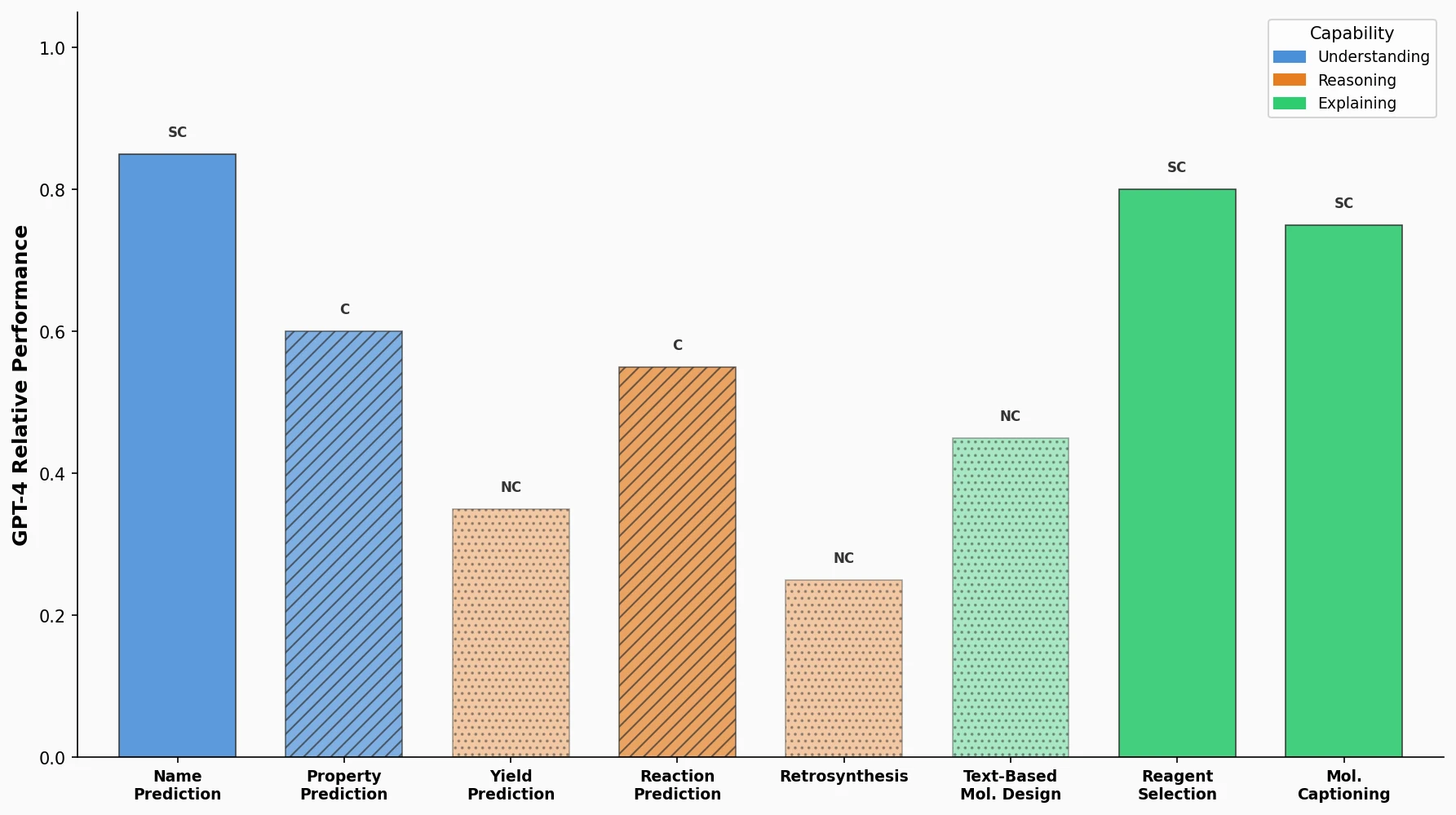
| Category | Tasks | Key Observation |
|---|---|---|
| Not Competitive (NC) | Name prediction, Reaction prediction, Retrosynthesis | LLMs lack deep understanding of SMILES strings; 70% lower accuracy than Chemformer on reaction prediction |
| Competitive (C) | Yield prediction, Reagents selection | Classification/ranking formulations are more tractable; GPT-4 reaches 80% accuracy on Buchwald-Hartwig yield prediction vs. 96.5% for UAGNN |
| Selectively Competitive (SC) | Property prediction, Molecule design, Molecule captioning | Performance depends heavily on prompt design; GPT-4 outperforms RF/XGBoost on HIV and ClinTox when property label semantics are included in prompts |
GPT-4 ranked first on 6 of 8 tasks by average performance, with an overall average rank of 1.25 across all tasks.
Key findings on ICL
Three consistent observations emerged across tasks:
- ICL prompting outperforms zero-shot prompting on all tasks
- Scaffold-based retrieval of similar examples generally outperforms random sampling
- Using more ICL examples (larger k) typically improves performance
SMILES vs. SELFIES comparison
The authors tested SELFIES representations as an alternative to SMILES on four tasks. SMILES outperformed SELFIES on all tasks, likely because LLM pretraining data contains more SMILES-related content. However, SELFIES produced fewer invalid molecular strings, consistent with its design guarantee of chemical validity.
Key Findings and Limitations
Performance patterns
The benchmark reveals a clear performance hierarchy: GPT-4 outperforms all others, followed by Davinci-003 and GPT-3.5 (roughly comparable), with Llama2-13B-chat and Galactica-30B trailing well behind. The ranking is consistent across most tasks.
LLMs perform best when chemistry tasks can be cast as classification or ranking problems rather than generation tasks requiring precise SMILES output. Text-related tasks (molecule captioning, property prediction with label semantics) also play to LLM strengths.
Fundamental limitation: SMILES understanding
The paper identifies a core limitation: LLMs treat SMILES strings as character sequences via byte-pair encoding tokenization, which fragments molecular structure information. Specific issues include:
- Inability to infer implicit hydrogen atoms
- Failure to recognize equivalent SMILES representations of the same molecule
- Tokenization that breaks SMILES into subwords not aligned with chemical substructures
- Generation of chemically invalid SMILES (up to 27.8% invalid for Llama2-13B-chat on reaction prediction)
Hallucination in chemistry
Two types of hallucinations were identified:
- Input hallucinations: Misinterpreting SMILES input (e.g., failing to count atoms or recognize functional groups)
- Output hallucinations: Generating chemically unreasonable molecules when SMILES output is required
Evaluation metric limitations
The authors note that standard NLP metrics (BLEU, ROUGE) do not fully capture chemical correctness. For molecule design, exact match is a more meaningful metric than BLEU, yet GPT-4 achieves only 17.4% exact match despite a BLEU score of 0.816. This highlights the need for chemistry-specific evaluation metrics.
Future directions
The authors suggest several promising directions: advanced prompting techniques (chain-of-thought, decomposed prompting), coupling LLMs with chemistry-specific tools (e.g., RDKit), and developing chemistry-aware ICL methods for higher-quality demonstration examples.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Understanding | PubChem | 630 molecules | Name prediction (500 ICL, 100 test) |
| Understanding | BBBP, HIV, BACE, Tox21, ClinTox (MoleculeNet) | 2,053-41,127 ICL candidates | Property prediction, MIT license |
| Reasoning | Buchwald-Hartwig, Suzuki-Miyaura (HTE) | 3,957 / 5,650 | Yield prediction, MIT license |
| Reasoning | USPTO-Mixed | 409,035 ICL candidates | Reaction prediction, MIT license |
| Reasoning | Suzuki HTE | 5,760 | Reagents selection, MIT license |
| Reasoning | USPTO-50k | 40,029 ICL candidates | Retrosynthesis, MIT license |
| Explaining | ChEBI-20 | 26,407 ICL candidates | Molecule design and captioning, CC BY 4.0 |
Algorithms
- Zero-shot and few-shot ICL prompting with standardized templates
- Scaffold-based retrieval using Tanimoto similarity on 2048-bit Morgan fingerprints (radius=2)
- Text similarity via Python’s difflib.SequenceMatcher
- Grid search over k and retrieval strategies on a 30-instance validation set
- Five repeated evaluations per task configuration to account for LLM stochasticity
Models
Five LLMs evaluated: GPT-4, GPT-3.5-turbo, text-davinci-003, Llama2-13B-chat, and Galactica-30B. Baselines include Chemformer (reaction prediction, retrosynthesis), UAGNN (yield prediction), MolT5-Large (molecule design, captioning), STOUT (name prediction), and RF/XGBoost from MoleculeNet (property prediction).
Evaluation
- Accuracy and F1 score for classification tasks (property prediction, yield prediction)
- Top-1 accuracy and invalid SMILES rate for generation tasks (reaction prediction, retrosynthesis)
- BLEU, exact match, Levenshtein distance, validity, fingerprint Tanimoto similarity (MACCS, RDK, Morgan), and FCD for molecule design
- BLEU-2, BLEU-4, ROUGE-1/2/L, and METEOR for molecule captioning
- All evaluations repeated 5 times; mean and standard deviation reported
Hardware
Not specified in the paper. Evaluation was conducted via API calls for GPT models; local inference details for Llama and Galactica are not provided.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| ChemLLMBench | Code | Not specified | Official benchmark code and prompts (Jupyter notebooks) |
Paper Information
Citation: Guo, T., Guo, K., Nan, B., Liang, Z., Guo, Z., Chawla, N. V., Wiest, O., & Zhang, X. (2023). What can Large Language Models do in chemistry? A comprehensive benchmark on eight tasks. Advances in Neural Information Processing Systems 36 (NeurIPS 2023), 59662-59688.
@inproceedings{guo2023chemllmbench,
title={What can Large Language Models do in chemistry? A comprehensive benchmark on eight tasks},
author={Guo, Taicheng and Guo, Kehan and Nan, Bozhao and Liang, Zhenwen and Guo, Zhichun and Chawla, Nitesh V. and Wiest, Olaf and Zhang, Xiangliang},
booktitle={Advances in Neural Information Processing Systems 36 (NeurIPS 2023)},
pages={59662--59688},
year={2023}
}