A Hierarchical Benchmark for Chemistry LLMs
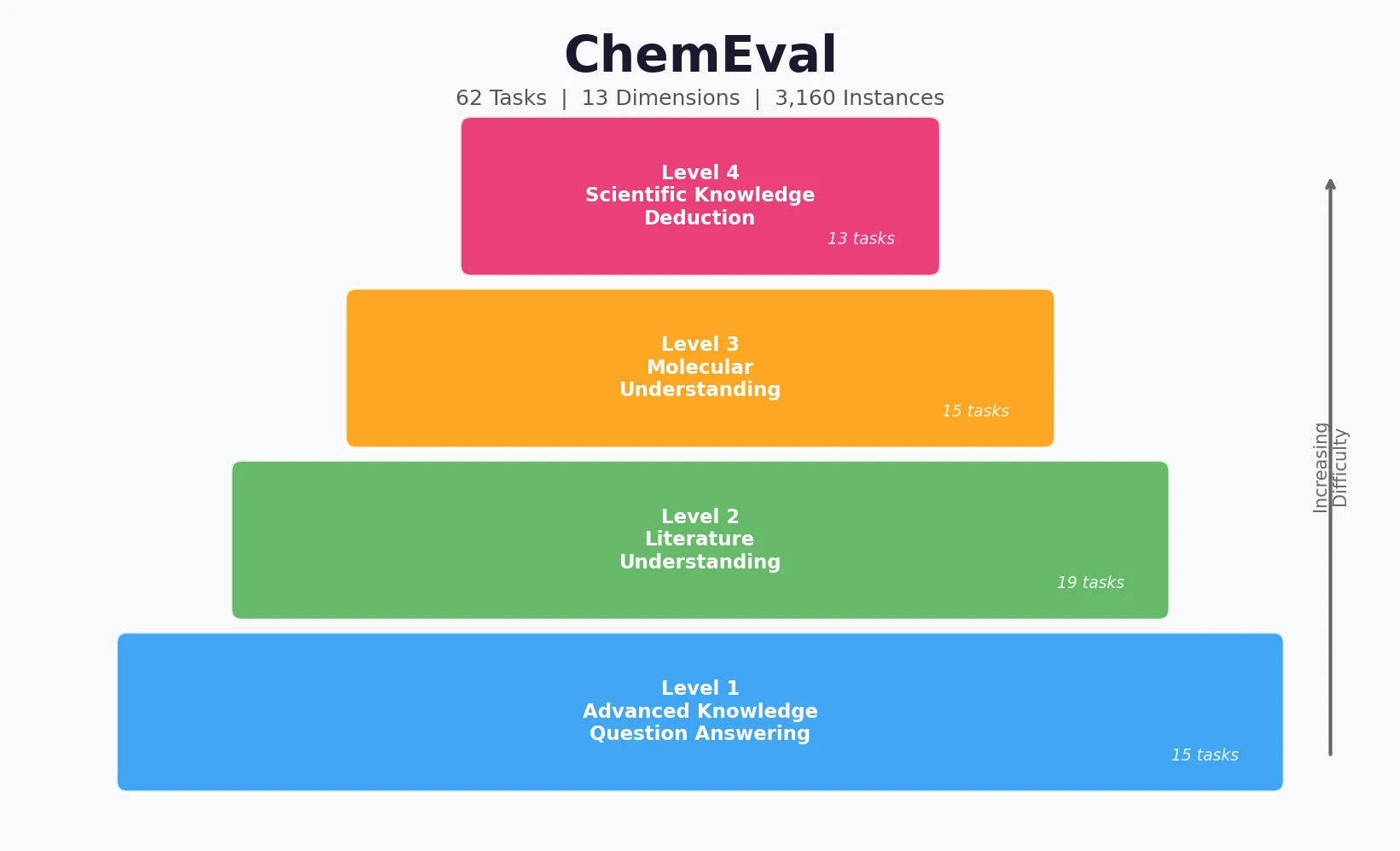
ChemEval is a Resource paper that introduces a comprehensive, hierarchical benchmark for evaluating large language models on chemical tasks. The benchmark spans four progressive levels of difficulty (Advanced Knowledge Question Answering, Literature Understanding, Molecular Understanding, and Scientific Knowledge Deduction), encompasses 13 capability dimensions, and contains 62 distinct tasks with 3,160 evaluation instances. It covers both text-only and multimodal settings, making it one of the most extensive chemistry-specific LLM evaluation frameworks to date.
Gaps in Existing Chemistry Benchmarks
Prior benchmarks for chemistry LLMs had several shortcomings:
- General benchmarks (MMLU, XieZhi, C-Eval) include some chemistry questions but lack the depth needed for meaningful evaluation of domain expertise.
- SciEVAL covers scientific tasks broadly but treats chemistry superficially with overly simplistic questions.
- ChemLLMBench (Guo et al., 2023) includes only 8 task categories derived from existing public datasets, offering insufficient breadth.
- ChemBench (Mirza et al., 2024) provides 7,000 samples but relies exclusively on multiple-choice questions and lacks open-ended evaluation for tasks like synthesis pathway recommendation.
- MaCBench (Alampara et al., 2025) introduces multimodal evaluation but remains limited in task diversity.
None of these benchmarks address LLMs’ ability to extract chemical information from text and tables, and none provide a graduated, multi-level assessment of chemical competence from basic knowledge through to advanced scientific reasoning.
A Four-Level Hierarchical Evaluation Framework
ChemEval’s core innovation is its hierarchical structure that mirrors how chemical expertise develops, from foundational knowledge through applied scientific reasoning.
Level 1: Advanced Knowledge Question Answering
This level assesses fundamental chemical knowledge through 15 tasks across two dimensions:
- Objective Questions (ObjQA): multiple choice, fill-in-the-blank, and true/false tasks spanning seven core chemistry disciplines (organic, inorganic, materials, analytical, biochemistry, physical, and polymer chemistry).
- Subjective Questions (SubjQA): short answer and calculation tasks requiring detailed reasoning and explanation.
Level 2: Literature Understanding
This level evaluates the ability to interpret chemical literature through 19 tasks across three dimensions:
- Information Extraction (InfoE): 11 tasks covering named entity recognition, relationship classification, substrate extraction, additive/solvent/temperature/time extraction, product extraction, characterization method extraction, catalysis type extraction, and yield extraction.
- Inductive Generation (InducGen): abstract generation, research outline generation, topic classification, and reaction type recognition.
- Molecular Name Recognition (MNR): molecular formula recognition, chemical reaction equation recognition, 2D molecular structure recognition, and synthetic pathway analysis (multimodal tasks).
Level 3: Molecular Understanding
This level tests molecular-level comprehension through 15 tasks across four dimensions:
- Molecular Name Generation (MNGen): generating SMILES from text descriptions.
- Molecular Name Translation (MNTrans): IUPAC to molecular formula, SMILES to molecular formula, IUPAC to SMILES, SMILES to IUPAC, and SMILES/SELFIES interconversion.
- Molecular Property Prediction (MPP): classification (ClinTox, HIV inhibition, polarity) and regression (lipophilicity, boiling point).
- Molecular Description (MolDesc): physicochemical property prediction from molecular structures and various spectral inputs (IR, Raman, UV-Vis, diffraction, mass spectrum, NMR).
Level 4: Scientific Knowledge Deduction
The most advanced level covers 13 tasks across four dimensions:
- Retrosynthetic Analysis (ReSyn): substrate recommendation, synthetic pathway recommendation, and synthetic difficulty evaluation.
- Reaction Condition Recommendation (RCRec): ligand, reagent, solvent, catalyst, temperature, and time recommendation.
- Reaction Outcome Prediction (ROP): product prediction, yield prediction, and reaction rate prediction.
- Reaction Mechanism Analysis (RMA): intermediate derivation.
Data Construction
The benchmark combines open-source datasets (ChemRxnExtractor, Mol-Instructions, ChemLLMBench, SMolInstruct) with domain-expert data curated from approximately 500 university-level chemistry textbooks and 9,000 real-world experimental records. Expert-crafted questions were written from scratch to prevent data leakage. A three-tier quality assurance pipeline (annotation by undergraduate students, review by graduate students, final audit by chemistry faculty) ensures correctness.
The text subset contains 1,960 instances (18 open-source tasks, 24 in-house tasks), while the multimodal subset contains 1,200 instances (12 open-source tasks, 30 in-house tasks).
Experimental Setup and Model Comparison
Models Evaluated
ChemEval evaluates a broad set of models under both zero-shot and 3-shot settings:
General LLMs: OpenAI-o1, OpenAI-o3-mini, GPT-4o, Claude-3.7-Sonnet (thinking and non-thinking modes), Gemini-2.5-Pro, Grok3, DeepSeek-V3, DeepSeek-R1, Qwen2.5 (7B/14B/32B/72B), LLaMA3.3-8B.
Chemistry-specific LLMs: ChemDFM, LlaSMol, ChemLLM, ChemSpark.
Multimodal LLMs (for multimodal tasks): GPT-4o, Claude-3.7-Sonnet, Qwen-VL Max, Phi-Vision-3.5, Gemini-2.5-Pro, GLM-4V.
Evaluation Metrics
The benchmark employs task-appropriate metrics: F1 score, Accuracy, BLEU, Exact Match, Normalized RMSE, Tanimoto similarity (with valid output ratio), LLM Score (judged by GPT-4o), L2 Score for molecular formula similarity, and Overlap for range prediction.
Key Results (Zero-Shot Text Tasks)
| Level | Top General LLM | Score | Top Chemistry LLM | Score |
|---|---|---|---|---|
| Knowledge QA (MCTask) | Gemini-2.5-Pro | 87.60% | ChemCrow | 58.00% |
| Literature (CNER) | Gemini-2.5-Pro | 68.30 F1 | ChemSpark | 71.44 F1 |
| Molecular (MolNG) | Gemini-2.5-Pro | 71.11 Tan. | ChemSpark | 74.81 Tan. |
| Molecular (IUPAC2SMILES) | Gemini-2.5-Pro | 61.33 Tan. | ChemSpark | 87.54 Tan. |
| Scientific (SubRec) | OpenAI-o3-mini | 4.67 F1 | ChemSpark | 12.37 F1 |
| Scientific (CatRec) | All models | 0.00 F1 | ChemSpark | 0.20 F1 |
Key Findings and Performance Patterns
General vs. Chemistry-Specific LLMs
General-purpose LLMs excel at Advanced Knowledge QA and Literature Understanding, benefiting from strong document comprehension and instruction-following abilities. Chemistry-specialized models (particularly ChemSpark) outperform in tasks demanding domain-specific molecular knowledge, such as molecular name translation and reaction condition recommendation. However, specialized models show notably weaker instruction-following capability and suffer from catastrophic forgetting of general language abilities during fine-tuning. For example, ChemLLM scores 0.00 on multiple information extraction tasks where general LLMs achieve 60-95%.
Impact of Few-Shot Learning
General LLMs tend to benefit from few-shot prompting, particularly for subjective QA and literature understanding tasks. OpenAI-o1 improved on 9 of 10 evaluated tasks. In contrast, chemistry-specialized models often show performance degradation with few-shot examples, likely due to loss of in-context learning capabilities during task-specific fine-tuning. ChemSpark decreased on 7 of 10 tasks in the 3-shot setting.
Impact of Model Scaling
Experiments with Qwen2.5 at 7B, 14B, 32B, and 72B parameters show that scaling improves performance on knowledge QA and literature understanding tasks. However, molecular understanding and scientific knowledge deduction tasks show minimal improvement, and some tasks (e.g., molecular property classification) even decline at the largest scale. Tasks requiring specialized chemical knowledge, like IUPAC-to-SMILES conversion and catalyst recommendation, remain near zero regardless of model size.
Thinking Models
Comparing OpenAI-o1 vs. GPT-4o and DeepSeek-R1 vs. DeepSeek-V3, thinking models show comparable overall performance to their non-thinking counterparts. They occasionally excel on specific tasks (e.g., reaction product prediction) but do not consistently outperform across chemical tasks. The authors conclude that the primary bottleneck is insufficient domain-specific knowledge, not reasoning depth.
Multimodal Tasks
Multimodal LLMs handle basic tasks like molecular formula recognition well (GLM-4V and Qwen-VL Max: 100% accuracy) but struggle with advanced challenges. Synthetic pathway analysis yielded 0% F1 across all models. 2D molecular structure recognition produced Tanimoto scores below 21% for all models tested. The performance gap between basic recognition and advanced chemical reasoning is substantial.
Limitations
The authors acknowledge several limitations:
- Limited instances per task: with 62 task types and 3,160 total instances, individual tasks may have as few as 20 samples.
- Static, single-turn evaluation: the benchmark does not assess dynamic interaction, tool use, or agentic workflows.
- No chemistry-specific multimodal models tested: only general-purpose VLMs were evaluated on multimodal tasks.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Evaluation (text) | ChemEval text subset | 1,960 instances | 18 open-source + 24 in-house tasks |
| Evaluation (multimodal) | ChemEval multimodal subset | 1,200 instances | 12 open-source + 30 in-house tasks |
| Source (open-source) | ChemRxnExtractor, Mol-Instructions, ChemLLMBench, SMolInstruct | Various | Adapted for ChemEval format |
| Source (expert) | ~500 textbooks, ~9,000 experimental records | Various | Novel questions crafted by domain experts |
Algorithms
- Evaluation prompts: task-specific instructions designed for formatted output, with 0-shot and 3-shot variants.
- Decoding: greedy decoding for all LLM inference.
- LLM-as-judge: GPT-4o used for LLM Score metric on subjective tasks.
Evaluation
Key metrics by task type:
| Metric | Task Types | Notes |
|---|---|---|
| Accuracy | MCTask, TFTask, MolPC, SubE, etc. | Standard classification accuracy |
| F1 Score | CNER, CERC, extraction tasks, reaction prediction | Precision-recall harmonic mean |
| BLEU | SMILES2IUPAC | N-gram overlap with brevity penalty |
| Exact Match | SMILES2IUPAC | Strict string match |
| Tanimoto Similarity | Molecular generation/translation tasks | Fingerprint-based molecular similarity |
| NRMSE | Regression tasks (property, temperature, time) | Normalized prediction error |
| LLM Score | Subjective QA, abstract generation, pathway rec. | GPT-4o evaluation (0-100) |
| L2 Score | Molecular formula tasks | $1 / (1 + \text{L2 distance})$ between formulas |
| Overlap | Rate prediction | Intersection/union of predicted vs. reference ranges |
Hardware
- Chemistry-specific models run on two NVIDIA A40 48GB GPUs.
- General models accessed via official APIs.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| ChemEval Benchmark | Code + Data | Other (custom) | Evaluation framework and task data |
Paper Information
Citation: Huang, Y., Zhang, R., He, X., Zhi, X., Wang, H., Chen, N., Liu, Z., Li, X., Xu, F., Liu, D., Liang, H., Li, Y., Cui, J., Xu, Y., Wang, S., Liu, Q., Lian, D., Liu, G., & Chen, E. (2024). ChemEval: A Comprehensive Multi-Level Chemical Evaluation for Large Language Models. arXiv preprint arXiv:2409.13989.
@article{huang2024chemeval,
title={ChemEval: A Comprehensive Multi-Level Chemical Evaluation for Large Language Models},
author={Huang, Yuqing and Zhang, Rongyang and He, Xuesong and Zhi, Xuyang and Wang, Hao and Chen, Nuo and Liu, Zongbo and Li, Xin and Xu, Feiyang and Liu, Deguang and Liang, Huadong and Li, Yi and Cui, Jian and Xu, Yin and Wang, Shijin and Liu, Qi and Lian, Defu and Liu, Guiquan and Chen, Enhong},
journal={arXiv preprint arXiv:2409.13989},
year={2024},
doi={10.48550/arXiv.2409.13989}
}